Abstract
Microglia rapidly respond to changes in neural activity and inflammation to regulate synaptic connectivity. The extracellular signals, particularly neuron-derived molecules, that drive these microglial functions at synapses remain a key open question. Here we show that whisker lesioning, known to dampen cortical activity, induces microglia-mediated synapse elimination. This synapse elimination is dependent on signaling by CX3CR1, the receptor for microglial fractalkine (also known as CXCL1), but not complement receptor 3. Furthermore, mice deficient in CX3CL1 have profound defects in synapse elimination. Single-cell RNA sequencing revealed that Cx3cl1 is derived from cortical neurons, and ADAM10, a metalloprotease that cleaves CX3CL1 into a secreted form, is upregulated specifically in layer IV neurons and in microglia following whisker lesioning. Finally, inhibition of ADAM10 phenocopies Cx3cr1−/− and Cx3cl1−/− synapse elimination defects. Together, these results identify neuron-to-microglia signaling necessary for cortical synaptic remodeling and reveal that context-dependent immune mechanisms are utilized to remodel synapses in the mammalian brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data discussed in this publication have been deposited in the NCBI’s Gene Expression Omnibus64 and are accessible through GEO series accession number GSE129150. All tools, reagents, and data that support the findings will be shared on an unrestricted basis. All requests should be directed to the corresponding author.
References
Wu, Y., Dissing-Olesen, L., MacVicar, B. A. & Stevens, B. Microglia: dynamic mediators of synapse development and plasticity. Trends Immunol. 36, 605–613 (2015).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Hong, S., Dissing-Olesen, L. & Stevens, B. New insights on the role of microglia in synaptic pruning in health and disease. Curr. Opin. Neurobiol. 36, 128–134 (2016).
Stevens, B. et al. The classical complement cascade mediates CNS synapse elimination. Cell 131, 1164–1178 (2007).
Vasek, M. J. et al. A complement–microglial axis drives synapse loss during virus-induced memory impairment. Nature 534, 538–543 (2016).
Lui, H. et al. Progranulin deficiency promotes circuit-specific synaptic pruning by microglia via complement activation. Cell 165, 921–935 (2016).
Hong, S. et al. Complement and microglia mediate early synapse loss in Alzheimer mouse models. Science 352, 712–716 (2016).
Wolf, Y., Yona, S., Kim, K. W. & Jung, S. Microglia, seen from the CX3CR1 angle. Front. Cell. Neurosci. 7, 26 (2013).
Jung, S. et al. Analysis of fractalkine receptor CX3CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Nishiyori, A. et al. Localization of fractalkine and CXCR1 mRNAs in rat brain: does fractalkine play a role in signaling from neuron to microglia? FEBS Lett. 429, 167–172 (1998).
Schecter, R. W. et al. Experience-dependent synaptic plasticity in V1 occurs without microglial CX3CR1. J. Neurosci. 37, 10541–10553 (2017).
Lowery, R. L., Tremblay, M. E., Hopkins, B. E. & Majewska, A. K. The microglial fractalkine receptor is not required for activity-dependent plasticity in the mouse visual system. Glia 65, 1744–1761 (2017).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Hoshiko, M., Arnoux, I., Avignone, E., Yamamoto, N. & Audinat, E. Deficiency of the microglial receptor CX3CR1 impairs postnatal functional development of thalamocortical synapses in the barrel cortex. J. Neurosci. 32, 15106–15111 (2012).
Zhan, Y. et al. Deficient neuron–microglia signaling results in impaired functional brain connectivity and social behavior. Nat. Neurosci. 17, 400–406 (2014).
Woolsey, T. A. & Van der Loos, H. The structural organization of layer IV in the somatosensory region (SI) of mouse cerebral cortex. The description of a cortical field composed of discrete cytoarchitectonic units. Brain Res. 17, 205–242 (1970).
Van der Loos, H. & Woolsey, T. A. Somatosensory cortex: structural alterations following early injury to sense organs. Science 179, 395–398 (1973).
Fox, K. A critical period for experience-dependent synaptic plasticity in rat barrel cortex. J. Neurosci. 12, 1826–1838 (1992).
Glazewski, S., McKenna, M., Jacquin, M. & Fox, K. Experience-dependent depression of vibrissae responses in adolescent rat barrel cortex. Eur. J. Neurosci. 10, 2107–2116 (1998).
Oberlaender, M., Ramirez, A. & Bruno, R. M. Sensory experience restructures thalamocortical axons during adulthood. Neuron 74, 648–655 (2012).
Wimmer, V. C., Broser, P. J., Kuner, T. & Bruno, R. M. Experience-induced plasticity of thalamocortical axons in both juveniles and adults. J. Comp. Neurol. 518, 4629–4648 (2010).
Erzurumlu, R. S. & Kind, P. C. Neural activity: sculptor of ‘barrels’ in the neocortex. Trends Neurosci. 24, 589–595 (2001).
Sadaka, Y., Weinfeld, E., Lev, D. L. & White, E. L. Changes in mouse barrel synapses consequent to sensory deprivation from birth. J. Comp. Neurol. 457, 75–86 (2003).
Erzurumlu, R. S. & Gaspar, P. Development and critical period plasticity of the barrel cortex. Eur. J. Neurosci. 35, 1540–1553 (2012).
Tremblay, M. E., Lowery, R. L. & Majewska, A. K. Microglial interactions with synapses are modulated by visual experience. PLoS Biol. 8, e1000527 (2010).
Sipe, G. O. et al. Microglial P2Y12 is necessary for synaptic plasticity in mouse visual cortex. Nat. Commun. 7, 10905 (2016).
Weinhard, L. et al. Microglia remodel synapses by presynaptic trogocytosis and spine head filopodia induction. Nat. Commun. 9, 1228 (2018).
Kim, K. W. et al. In vivo structure/function and expression analysis of the CX3C chemokine fractalkine. Blood 118, e156–e167 (2011).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Hrvatin, S. et al. Single-cell analysis of experience-dependent transcriptomic states in the mouse visual cortex. Nat. Neurosci. 21, 120–129 (2018).
Hundhausen, C. et al. The disintegrin-like metalloproteinase ADAM10 is involved in constitutive cleavage of CX3CL1 (fractalkine) and regulates CX3CL1-mediated cell–cell adhesion. Blood 102, 1186–1195 (2003).
Suzuki, K. et al. Activity-dependent proteolytic cleavage of neuroligin-1. Neuron 76, 410–422 (2012).
Madoux, F. et al. Discovery of an enzyme and substrate selective inhibitor of ADAM10 using an exosite-binding glycosylated substrate. Sci. Rep. 6, 11 (2016).
Shackleton, B., Crawford, F. & Bachmeier, C. Inhibition of ADAM10 promotes the clearance of Aβ across the BBB by reducing LRP1 ectodomain shedding. Fluids Barriers CNS 13, 14 (2016).
Venkatesh, H. S. et al. Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549, 533–537 (2017).
Hickman, S. E. et al. The microglial sensome revealed by direct RNA sequencing. Nat. Neurosci. 16, 1896–1905 (2013).
Eyo, U. B. et al. Regulation of physical microglia–neuron interactions by fractalkine signaling after status epilepticus. eNeuro 3, ENEURO.0209-16.2016 (2017).
Datwani, A. et al. Classical MHCI molecules regulate retinogeniculate refinement and limit ocular dominance plasticity. Neuron 64, 463–470 (2009).
Huh, G. S. et al. Functional requirement for class I MHC in CNS development and plasticity. Science 290, 2155–2159 (2000).
Vainchtein, I. D. et al. Astrocyte-derived interleukin-33 promotes microglial synapse engulfment and neural circuit development. Science 359, 1269–1273 (2018).
Woodward, N. D., Giraldo-Chica, M., Rogers, B. & Cascio, C. J. Thalamocortical dysconnectivity in autism spectrum disorder: an analysis of the Autism Brain Imaging Data Exchange. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 2, 76–84 (2017).
Woodward, N. D., Karbasforoushan, H. & Heckers, S. Thalamocortical dysconnectivity in schizophrenia. Am. J. Psychiatry 169, 1092–1099 (2012).
Chen, R., Cohen, L. G. & Hallett, M. Nervous system reorganization following injury. Neuroscience 111, 761–773 (2002).
Lauro, C., Catalano, M., Trettel, F. & Limatola, C. Fractalkine in the nervous system: neuroprotective or neurotoxic molecule? Ann. NY Acad. Sci. 1351, 141–148 (2015).
Pruessmeyer, J. & Ludwig, A. The good, the bad and the ugly substrates for ADAM10 and ADAM17 in brain pathology, inflammation and cancer. Semin. Cell Dev. Biol. 20, 164–174 (2009).
Ransohoff, R. M. & Benveniste, E. N. Cytokines and the CNS 2nd edn (Taylor and Francis, 2006).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Aβ, tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Iaccarino, H. F. et al. Gamma frequency entrainment attenuates amyloid load and modifies microglia. Nature 540, 230–235 (2016).
Martorell, A. J. et al. Multi-sensory gamma stimulation ameliorates Alzheimer’s-associated pathology and improves cognition. Cell 177, 256–271.e222 (2019).
Chozinski, T. J. et al. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat. Methods 13, 485 (2016).
Yang, C. et al. NRBF2 is involved in the autophagic degradation process of APP-CTFs in Alzheimer disease models. Autophagy 13, 2028–2040 (2017).
Manabe, Y. et al. Glial cell line-derived neurotrophic factor protein prevents motor neuron loss of transgenic model mice for amyotrophic lateral sclerosis. Neurol. Res. 25, 195–200 (2003).
Gey, M. et al. Atf3 mutant mice show reduced axon regeneration and impaired regeneration-associated gene induction after peripheral nerve injury. Open Biol. 6, 160091 (2016).
Hao, F. et al. Long-term protective effects of AAV9-mesencephalic astrocyte-derived neurotrophic factor gene transfer in parkinsonian rats. Exp. Neurol. 291, 120–133 (2017).
Droguett, A. et al. Tubular overexpression of gremlin induces renal damage susceptibility in mice. PLoS One 9, e101879 (2014).
Viganò, F. et al. GPR17 expressing NG2-glia: oligodendrocyte progenitors serving as a reserve pool after injury. Glia 64, 287–299 (2016).
Kluge, M. G. et al. Age-dependent disturbances of neuronal and glial protein expression profiles in areas of secondary neurodegeneration post-stroke. Neuroscience 393, 185–195 (2018).
Funk, K. E. & Klein, R. S. CSF1R antagonism limits local restimulation of antiviral CD8+ T cells during viral encephalitis. J. Neuroinflamm. 16, 22 (2019).
Schafer, D. P., Lehrman, E. K., Heller, C. T. & Stevens, B. An engulfment assay: a protocol to assess interactions between CNS phagocytes and neurons. J. Vis. Exp. 88, e51482 (2014).
Zilionis, R. et al. Single-cell barcoding and sequencing using droplet microfluidics. Nat. Protoc. 12, 44–73 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Acknowledgements
The authors thank M. Freeman (OHSU), V. Budnik (UMMS), E. Baehrecke (UMMS), M. Francis (UMMS), P. Greer (UMMS), and R. Bruno (Columbia University) for their critical reading of the manuscript. They also thank the following individuals: M. Ansorage (Columbia University) and S. Nelson (Brandeis University) for providing the SERT-Cre mice; Cahill (UMMS) and A. Lotun (UMMS) for assistance with assessing microglia within the barrel cortex; S. Becker (UMMS) and J. Jung (UMMS) for assistance with tissue preparation and whisker-trimming experiments; H. Learnard (UMMS), A. Song (UMMS), Z. Zhang (Boston Children’s Hospital), and C. Woolf (Boston Children’s Hospital) for assistance with experiments to assess ATF3; and D. Bergles (John’s Hopkins) for advice and discussions related to identification of oligodendrocye precursor cells by Matn4 expression in the single-cell RNA-seq dataset. This work was funded by NIMH-R00MH102351 (D.P.S.), NIMH-R01MH113743 (D.P.S.), NIMH-R21MH115353 (D.P.S. and A.S.), NIH-T32A1095212 (E.M.), the Charles H. Hood Foundation (D.P.S.), the Brain & Behavior Research Foundation (D.P.S.), the Worcester Foundation (D.P.S.), and the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (D.P.S.).
Author information
Authors and Affiliations
Contributions
G.G. and D.P.S. designed the study, performed most experiments, analyzed most data, and wrote the manuscript. K.M.J. assisted in the design of initial experiments, and performed experiments to identify initial synapse remodeling and engulfment phenotypes. L.C., M.A.N., and M.E.G. performed the single-cell sequencing experiments. E.M. performed the in situ hybridization experiments. P.A., A.B., and A.S. performed the bulk RNA-seq experiments of whole barrel cortices. L.L. and A.R.T. performed the electrophysiology experiments. K.-W.K., S.M.B., and B.T.L. performed experiments related to Cx3cl1−/− mice, and S.A.L. provided the Cx3cl1−/− mice. R.M.R. provided critical input into the study design and feedback on the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Neuroscience thanks Marie-Ève Tremblay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 TC input elimination is observed following whisker trimming and with genetic labeling of TC inputs.
a, Daily whisker trimming from P4 results in decreased barrel fluorescence intensity by day 16 (bottom panels), but no by day 6 (top panels). Scale bar, 150 µm. b, Quantification for barrel fluorescence intensity in a (Two-way ANOVA with Sidak’s post hoc, 6-day control vs trimmed, n = 5 animals, P = 0.5093, t = 1.078, df = 14; 16-day control vs trimmed, n = 4 animals, **P = 0.0071, t =3.496, df = 14). Data are normalized to the control barrel cortex within the same animal for each timepoint. c, TC inputs labeled by transgenic expression of tdTomato are eliminated in the deprived barrel cortex in a CX3CR1-dependent manner following whisker lesioning by cauterization. Representative tangential sections of Sert-Cre tdTomato labeled TC inputs within layer IV of the control and deprived barrel cortices of Cx3cr1+/- (top row) and Cx3cr1-/- (bottom row) mice 7 d post-sensory deprivation. Scale bar, 150 µm. d, There is a significant decrease in TC inputs as measured by fluorescence intensity of tdTomato signal in the deprived (gray bars) vs. the contralateral control (black bars) barrel cortex in Cx3cr1+/- mice 7 d-post deprivation. No significant decrease in fluorescence was observed in Cx3cr1-/- mice. Data are normalized to the control barrel cortex for each genotype. (Two-Way ANOVA with Sidak’s post hoc, n=4 animals per genotype, Cx3cr1+/- control vs deprived, *P = 0.0405, t = 2.668, df = 12; Cx3cr1-/- control vs deprived, P = 0.9808, t = 0.1782, df = 12.) All data presented as mean ± SEM.
Supplementary Figure 2 Effects of whisker lesioning at P4 on cell death, axon degeneration, and cell stress in the barrel cortex circuit.
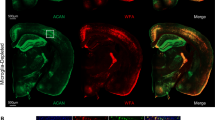
a, Immunostaining of trigeminal ganglia, which contain the neurons that innervate the whisker follicles. There is a significant increase in ATF3 (red, marker of cell stress) in NeuN-positive neurons (cyan) at 24h post whisker lesioning (deprived, bottom row). Scale bar, 50 µm. b, Quantification for ATF3 signal co-localized to NeuN in the control and deprived trigeminal ganglia. (Two-tailed Student’s t-Test, n = 4 animals; **P = 0.0012, t = 5.715, df = 6). c, Representative images from 3 animals show ATF3 signal (green) is not detected in the ventral posterior medial nucleus (VPM) of the thalamus (bottom panels; dotted line borders the VPM) nor the primary somatosensory cortex (top panels) 24 hours after whisker lesioning. Scale bars, 150 µm. d, Quantification for cell death marker cleaved caspase 3 shows cell death does not occur 24h nor 5d after whisker lesioning in the trigeminal ganglia. (Two-way ANOVA with Sidak’s post hoc, 24h control vs deprived, n = 3 animals, P >0.9999, t = 0, df = 8; 6d control vs deprived, n = 3 animals, P = 0.3520, t = 1.414, df = 8). Data presented as mean ± SEM. e-f, Representative images from 3 animals for the deprived somatosensory cortex and VPM 7 days after whisker lesioning in control mice shows no increased cell death (e; caspase 3, green) nor axon degeneration (f; APP, green) in either brain region. Scale bars, 150 µm.
Supplementary Figure 3 TC inputs are internalized within microglia following sensory deprivation.
a, Structured illumination microscopy (SIM) of a microglia (CX3CR1EGFP/+, green) which has engulfed VGluT2-positive TC presynaptic inputs (red) within its lysosomes (anti-CD68, cyan) in the deprived barrel cortex 24 h after whisker removal. Scale bar, 5 µm. b-c, Quantification of the % of microglia out of the total microglial population phagocytosing VGluT2-positive inputs 24h (b) and 5d (c) after whisker lesioning in the control (black bars) and deprived (grey bars) cortices. Microglia in the deprived cortex both 24h and 5d after lesioning have a higher proportion of cells with a high phagocytic index (b, >1.0%; c, >2.0%) compared to the control cortex (24h: n = 129 cells; Two-Sided Fisher’s Exact Test; for 0.6–0.8%; P = 0.1306; for 0.8–1.0%; P = 0.1941; for >0.1%, **P = 0.0051; 5d: n = 159 cells; Two-Sided Fisher’s Exact Test; for 0.4–0.8%; P >0.9999; for 0.8–1.2%; P >0.9999; for 1.2–1.6%; P = 0.6227; for 1.6–2.0%; P >0.9999; for >2.0% *P = 0.0298). Data represented as whole number percentage of the total cell population.
Supplementary Figure 4 CR3 is not required for whisker lesion-induced TC input engulfment and elimination.
a,b, Representative images of VGluT2 immunoreactivity in the non-deprived (a) and deprived (b) barrel cortices of CR3-KO mice show TC inputs are properly eliminated 6 d after whisker lesioning. Scale bar, 150 µm. c, Quantification of VGluT2 area within barrel centers 7 d after whisker removal reveals that TC inputs are still eliminated in CR3-KO mice. (Data normalized to the control, non-deprived VGluT2 area within each animal; Two-tailed Student’s t-test, n = 3 animals, **P = 0.0015, t =7.765 df =4). d,e, Top Panels, Fluorescent images of microglia within the control (d) and deprived (e) barrels of CR3-KO mice. Scale bar, 5 µm. Middle panels depict VGluT2 signal (red) within microglia (green) and within lysosomes (blue). Bottom panels are 3D surface-rendered insets (dotted boxes in middle panels) of control (d) and deprived (e) CR3-KO microglia. Arrows depict increased VGluT2 internalization within microglia in the deprived hemisphere. Scale bar, 2 µm. f, Quantification of VGluT2 engulfment within CR3-KO microglia 24 h after whisker removal reveals that CR3 deficiency fails to block microglial engulfment of TC inputs. (Data normalized to engulfment in microglia in the control hemisphere within each animal; Two-tailed Student’s t-test, n = 3 animals, *P = 0.0260, t =4.215 df =2). All data presented as mean ± SEM.
Supplementary Figure 5 Whisker lesioning in Cx3cr1-/- animals results in similar cell stress response and wound healing at the whisker follicles compared to Cx3cr1+/+ animals.
a-b, Whisker lesioning increases ATF3 signal in the ipsilateral trigeminal nerve ganglion of Cx3cr1-/- animals (deprived, bottom panels, compare to Supplementary Fig. 2a). Nuclei positive for ATF3 (red) are co-localized to NeuN signal (cyan). Scale bar, 50 µm. (Two-tailed Student’s t-Test, *P = 0.0152, t = 3.362, df = 6; n = 5 animals). c-e, Whisker lesioning in Cx3cr1-/- mice (bottom panels) results in similar wound healing response as measured by recruitment of CX3CR1-EGFP-positive (d; Two-tailed Student’s t-Test, P = 0.7115, t = 0.3852, df = 7; n = 4 Cx3cr1+/-, 5 Cx3cr1-/- animals) and CD45 macrophages/monocytes (e; Two-tailed Student’s t-Test, P = 0.6656, t = 0.4511, df = 7; n = 4 Cx3cr1+/-, 5 Cx3cr1-/- animals) compared to Cx3cr1+/+ animals (top panels) 24 hours after injury. Scale bars, 100 µm. Data presented as mean ± SEM. f-g, Whisker trimming in Cx3cr1-/- animals from P4 to P20 (16 days deprivation) does not result in a decrease in barrel fluorescence intensity (compare to Supplementary Fig. 1a, b). Top panel, control barrel field fluorescence; bottom panel, deprived barrel field fluorescence at P20. Representative images taken from 4 animals. Quantification for fluorescence intensity (g) does not show a significant decrease in fluorescence (Two-tailed Student’s t-test, n = 4 animals, P = 0.3455, t = 1.024, df = 6). Data presented as mean ± SEM. h, Cx3cr1 expression is specific to peripheral macrophages and monocytes in the whisker follicles and surrounding tissue before (left panels) and after (right panels) whisker lesioning in a Cx3cr1+/- animal. Cx3cr1EGFP/+ expression (EGFP, green) is co-localized with macrophage marker F4/80 (red) in both control and cauterized follicles. Scale bars, 100 µm. Representative images taken from 9 animals (n = 4 Cx3cr1+/-, 5 Cx3cr1-/- animals). i-l, Cx3cr1 expression is specific to microglia both before (control, top panels) and 24 hours after (deprived, bottom panels) whisker lesioning in the primary somatosensory cortex. Cx3cr1EGFP/+ expression is specific to microglia (i, P2RY12, low magnification scale bars 80 µm, high magnification scale bars 25 µm) but not neuronal (j, NeuN, scale bars 25 µm), astrocyte (k, ALDH1L1, scale bars 25 µm), or oligodendrocyte precursor (l, NG2, scale bars 25 µm) specific markers. Representative images taken from 3 Cx3cr1+/- animals.
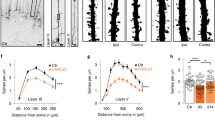
Supplementary Figure 6 Total number of microglia across the barrel field increases with postnatal age and is similar in Cx3cr1+/- and Cx3cr1-/- mice.
a-f, Uncropped images of microglia recruitment to barrel centers from Fig. 5. Scale bar, 150 µm. Representative images taken from 11 animals. g, For each genotype, the number of microglia within the septa (yellow highlighted area) and barrels (outlined by white dotted lines) were quantified in deprived and control, non-deprived layer IV barrel cortices. A ratio was then calculated: # of microglia within the barrel divided by the # of microglia within the septa. Scale bar, 150 µm. h, For each postnatal age analyzed, the total microglial cell density over the entire layer IV primary somatosensory cortex is not significantly changed between Cx3cr1+/- and Cx3cr1-/- mice. (Two-Way ANOVA with Dunnett’s post hoc test, n=4 animals per genotype; for P5: control +/- vs deprived +/-, P = 0.9997, q = 0.06821, df = 60, control +/- vs control -/-, P = 0.9371, q = 0.4673, df = 60, control +/- vs deprived -/-, P = 0.9949, q = 0.1929, df = 60; for P6: control +/- vs deprived +/-, P = 0.9549, q = 0.4136, df = 60, control +/- vs control -/-, P = 0.3484, q = 1.436, df = 60, control +/- vs deprived -/-, P = 0.7974, q = 0.7459, df = 60; for P7: control +/- vs deprived +/-, P = 0.9468, q = 0.4392, df = 60, control +/- vs control -/-, P = 0.8091, q = 0.7266, df = 60, control +/- vs deprived -/-, P = 0.9979, q = 0.1442, df = 60; for P8: control +/- vs deprived +/-, P = 0.5973, q = 1.043, df = 60, control +/- vs control -/-, P = 0.2481, q = 1.639, df = 60, control +/- vs deprived -/-, P = 3.194, q = 1.490, df = 60; for P9: control +/- vs deprived +/-, P = 0.3258, q = 1.479, df = 60, control +/- vs control -/-, P = 0.6219, q = 1.008, df = 60, control +/- vs deprived -/-, P = 0.0694, q = 2.269, df = 60). All data presented as mean ± SEM.
Supplementary Figure 7 Biological replicate contribution and sequencing depth of cells captured by inDrops.
a, Clustering cells from each animal condition across n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals reveals even distribution of cells from each animal across all identified cell populations. Each tSNE plot represents cells from each animal genotype and condition (Cx3cr1+/- control, Cx3cr1+/- deprived, Cx3cr1-/- control, Cx3cr1-/- deprived). b, The number of unique molecular identifiers (UMIs) per cell processed by inDrops single-cell RNAseq (logarithmic scale) across n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals. c, The number of genes positively identified increases with increasing UMIs. d, Table containing average number of reads per cell across individual animals.
Supplementary Figure 8 Cell populations clustered through principal component analysis were identified according to specific cell-type markers.
a-b, Microglia cluster identified by P2ry12 and C1qa. c-d, Astrocyte clusters identified by Aldoc and Aqp4 expression. e,f, Oligodendrocyte cluster identified by Plp1 and Mpb expression. e-f, Oligodendrocyte precursor cell clusters identified by Pdgfra and Matn4 expression. i-j, Neuron clusters identified by Tubb3 and Snap25 expression. k-l, Inhibitory neurons identified by Gad1 and Gad2 expression. All cells represented across n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals.
Supplementary Figure 9 Genes significantly changed for each cell population in the deprived cortex of Cx3cr1+/− animals.
Genes with significant changes in expression (FDR <0.10) and Log2 Fold Change greater than 0.5 or less than −0.5 in the deprived cortex for n = 4 Cx3cr1+/- are plotted for every identified cell population (Monocle2). Note ADAM10 (bold text and asterisks) is only significantly increased in layer IV excitatory neurons and microglia (outlined graphs).
Supplementary Figure 10 Sensory-lesion dependent gene expression changes in all major glial cell populations in the deprived cortices of Cx3cr1+/- and Cx3cr1-/- animals.
a, Gene expression changes for Cx3cr1+/- microglia (left panel, from n = 4 Cx3cr1+/- animals) and Cx3cr1-/- microglia (right panel, from n = 4 Cx3cr1-/- animals). Dotted lines indicates –log10FDR <0.10 and Log2 Fold Change > 0.5 and < −0.5 (Monocle2). b-f, Gene expression changes for all other identified glial cell types (Monocle2). g, Table containing the number of cells sequenced for each cell type for each animal genotype and condition. h, Heatmap for the Log2Fold Change for microglial genes changed across n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals hierarchically clustered by variable gene expression for Cx3cr1+/- samples (Monocle2).
Supplementary Figure 11 qPCR in Cx3cr1-/- primary somatosensory cortex reveals no significant increase in Cx3cl1 nor Adam10 expression after whisker lesioning.
a, Quantification for Cx3cl1 expression 6h and 24h post lesioning. (Two-way ANOVA with Sidak’s post hoc, 6h control vs deprived P = 0.6640, t = 0.8648, df = 6; 24h control vs deprived P = 0.2808, t = 1.641, df = 6; n = 3 animals per timepoint). b, Quantification for Adam10 expression 24h post lesioning. (Two-tailed Student’s t-Test, P = 0.4414, t = 0.9099, df = 4; n = 3 animals). Data presented as mean ± SEM.
Supplementary Figure 12 Baseline expression in Cx3cr1-/- microglia reveals significant changes in genes related to phagocytic signaling.
a, Volcano plot of genes significantly downregulated (fold change < −0.5, FDR <0.10, p<0.0005, Monocle2) in Cx3cr1-/- microglia within the non-deprived, control barrel cortex compared to Cx3cr1+/- littermates reveals several genes are dysregulated basally in Cx3cr1-/- microglia. (Microglia from n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals). b, Gene ontology clustering of genes in Cx3cr1-/- microglia (Dotted line, p<0.05) reveals several genes related to phagocytic signaling are significantly changed (single cell performed on n = 4 Cx3cr1+/- and n = 4 Cx3cr1-/- animals yielding 828 microglia, see also Supplementary Fig. 9g). Data analyzed through the use of IPA (QIAGEN Inc., https://www.qiagenbioinformatics.com/products/ingenuitypathway-analysis).
Supplementary information
Rights and permissions
About this article
Cite this article
Gunner, G., Cheadle, L., Johnson, K.M. et al. Sensory lesioning induces microglial synapse elimination via ADAM10 and fractalkine signaling. Nat Neurosci 22, 1075–1088 (2019). https://doi.org/10.1038/s41593-019-0419-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0419-y
This article is cited by
-
Cranial irradiation disrupts homeostatic microglial dynamic behavior
Journal of Neuroinflammation (2024)
-
Targeting synapse function and loss for treatment of neurodegenerative diseases
Nature Reviews Drug Discovery (2024)
-
Lifelong absence of microglia alters hippocampal glutamatergic networks but not synapse and spine density
EMBO Reports (2024)
-
Noteworthy perspectives on microglia in neuropsychiatric disorders
Journal of Neuroinflammation (2023)
-
Prolonged anesthesia induces neuroinflammation and complement-mediated microglial synaptic elimination involved in neurocognitive dysfunction and anxiety-like behaviors
BMC Medicine (2023)