Abstract
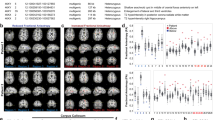
Williams syndrome (WS), caused by a heterozygous microdeletion on chromosome 7q11.23, is a neurodevelopmental disorder characterized by hypersociability and neurocognitive abnormalities. Of the deleted genes, general transcription factor IIi (Gtf2i) has been linked to hypersociability in WS, although the underlying mechanisms are poorly understood. We show that selective deletion of Gtf2i in the excitatory neurons of the forebrain caused neuroanatomical defects, fine motor deficits, increased sociability and anxiety. Unexpectedly, 70% of the genes with significantly decreased messenger RNA levels in the mutant mouse cortex are involved in myelination, and mutant mice had reduced mature oligodendrocyte cell numbers, reduced myelin thickness and impaired axonal conductivity. Restoring myelination properties with clemastine or increasing axonal conductivity rescued the behavioral deficits. The frontal cortex from patients with WS similarly showed reduced myelin thickness, mature oligodendrocyte cell numbers and mRNA levels of myelination-related genes. Our study provides molecular and cellular evidence for myelination deficits in WS linked to neuronal deletion of Gtf2i.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data are available in the main text or the supplementary materials. All RNA-Seq datasets generated and analyzed during the current study are available in the supplementary tables and from the corresponding author upon reasonable request. RNA data are available in the Gene Expression Omnibus (GEO) with accession code GSE128841. All data generated in this study are deposited in the GEO under accession code TBD.
References
Barak, B. & Feng, G. Neurobiology of social behavior abnormalities in autism and Williams syndrome. Nat. Neurosci. 19, 647–655 (2016).
Pober, B. R. Williams–Beuren syndrome. N. Engl. J. Med. 362, 239–252 (2010).
Karmiloff-Smith, A. et al. Social cognition in Williams syndrome: genotype/phenotype insights from partial deletion patients. Front. Psychol. 3, 168 (2012).
Dai, L. et al. Is it Williams syndrome? GTF2IRD1 implicated in visual-spatial construction and GTF2I in sociability revealed by high resolution arrays. Am. J. Med. Genet. A 149A, 302–314 (2009).
Morris, C. A. et al. GTF2I hemizygosity implicated in mental retardation in Williams syndrome: genotype–phenotype analysis of five families with deletions in the Williams syndrome region. Am. J. Med. Genet. A 123A, 45–59 (2003).
Bayés, M., Magano, L. F., Rivera, N., Flores, R. & Pérez Jurado, L. A. Mutational mechanisms of Williams–Beuren syndrome deletions. Am. J. Hum. Genet. 73, 131–151 (2003).
Roy, A. L. Biochemistry and biology of the inducible multifunctional transcription factor TFII-I: 10 years later. Gene 492, 32–41 (2012).
Osborne, L. R. Animal models of Williams syndrome. Am. J. Med. Genet. C 154C, 209–219 (2010).
Segura-Puimedon, M. et al. Heterozygous deletion of the Williams–Beuren syndrome critical interval in mice recapitulates most features of the human disorder. Hum. Mol. Genet. 23, 6481–6494 (2014).
Li, H. H. et al. Induced chromosome deletions cause hypersociability and other features of Williams–Beuren syndrome in mice. EMBO Mol. Med. 1, 50–65 (2009).
Borralleras, C., Sahun, I., Pérez-Jurado, L. A. & Campuzano, V. Intracisternal Gtf2i gene therapy ameliorates deficits in cognition and synaptic plasticity of a mouse model of Williams–Beuren syndrome. Mol. Ther. 23, 1691–1699 (2015).
Lucena, J. et al. Essential role of the N-terminal region of TFII-I in viability and behavior. BMC Med. Genet. 11, 61 (2010).
Sakurai, T. et al. Haploinsufficiency of Gtf2i, a gene deleted in Williams Syndrome, leads to increases in social interactions. Autism Res. 4, 28–39 (2011).
Enkhmandakh, B. et al. Essential functions of the Williams–Beuren syndrome-associated TFII-I genes in embryonic development. Proc. Natl Acad. Sci. USA 106, 181–186 (2009).
Enkhmandakh, B. et al. Generation of a mouse model for a conditional inactivation of Gtf2i allele. Genesis 54, 407–412 (2016).
Goebbels, S. et al. Genetic targeting of principal neurons in neocortex and hippocampus of NEX-Cre mice. Genesis 44, 611–621 (2006).
Chailangkarn, T. et al. A human neurodevelopmental model for Williams syndrome. Nature 536, 338–343 (2016).
Reiss, A. L. et al. An experiment of nature: brain anatomy parallels cognition and behavior in Williams syndrome. J. Neurosci. 24, 5009–5015 (2004).
Thompson, P. M. et al. Abnormal cortical complexity and thickness profiles mapped in Williams syndrome. J. Neurosci. 25, 4146–4158 (2005).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Chang, K. J., Redmond, S. A. & Chan, J. R. Remodeling myelination: implications for mechanisms of neural plasticity. Nat. Neurosci. 19, 190–197 (2016).
Rosenberg, S. S. & Chan, J. R. Modulating myelination: knowing when to say Wnt. Genes Dev. 23, 1487–1493 (2009).
Rosenberg, S. S., Powell, B. L. & Chan, J. R. Receiving mixed signals: uncoupling oligodendrocyte differentiation and myelination. Cell. Mol. Life Sci. 64, 3059–3068 (2007).
Nave, K. A. & Werner, H. B. Myelination of the nervous system: mechanisms and functions. Annu. Rev. Cell Dev. Biol. 30, 503–533 (2014).
Liu, J. et al. Impaired adult myelination in the prefrontal cortex of socially isolated mice. Nat. Neurosci. 15, 1621–1623 (2012).
Liu, J. et al. Clemastine enhances myelination in the prefrontal cortex and rescues behavioral changes in socially isolated mice. J. Neurosci. 36, 957–962 (2016).
Liu, J. et al. Chromatin landscape defined by repressive histone methylation during oligodendrocyte differentiation. J. Neurosci. 35, 352–365 (2015).
Chapman, C. A., du Plessis, A. & Pober, B. R. Neurologic findings in children and adults with Williams syndrome. J. Child Neurol. 11, 63–65 (1996).
Trauner, D. A., Bellugi, U. & Chase, C. Neurologic features of Williams and Down syndromes. Pediatr. Neurol. 5, 166–168 (1989).
Metz, G. A. & Whishaw, I. Q. Cortical and subcortical lesions impair skilled walking in the ladder rung walking test: a new task to evaluate fore- and hindlimb stepping, placing, and co-ordination. J. Neurosci. Methods 115, 169–179 (2002).
Liu, Y. et al. A sensitized IGF1 treatment restores corticospinal axon-dependent functions. Neuron 95, 817–833.e4 (2017).
Hayes, K. C. The use of 4-aminopyridine (fampridine) in demyelinating disorders. CNS Drug Rev. 10, 295–316 (2004).
Green, A. J. et al. Clemastine fumarate as a remyelinating therapy for multiple sclerosis (ReBUILD): a randomised, controlled, double-blind, crossover trial. Lancet 390, 2481–2489 (2017).
Cree, B. A. C. et al. Clemastine rescues myelination defects and promotes functional recovery in hypoxic brain injury. Brain 141, 85–98 (2018).
Wang, F. et al. Enhancing oligodendrocyte myelination rescues synaptic loss and improves functional recovery after chronic hypoxia. Neuron 99, 689–701.e5 (2018).
Mei, F. et al. Micropillar arrays as a high-throughput screening platform for therapeutics in multiple sclerosis. Nat. Med. 20, 954–960 (2014).
Arlinghaus, L. R., Thornton-Wells, T. A., Dykens, E. M. & Anderson, A. W. Alterations in diffusion properties of white matter in Williams syndrome. Magn. Reson. Imaging 29, 1165–1174 (2011).
Avery, S. N., Thornton-Wells, T. A., Anderson, A. W. & Blackford, J. U. White matter integrity deficits in prefrontal-amygdala pathways in Williams syndrome. Neuroimage 59, 887–894 (2012).
Faria, A. V. et al. Quantitative analysis of gray and white matter in Williams syndrome. Neuroreport 23, 283–289 (2012).
Hoeft, F. et al. More is not always better: increased fractional anisotropy of superior longitudinal fasciculus associated with poor visuospatial abilities in Williams syndrome. J. Neurosci. 27, 11960–11965 (2007).
Jabbi, M. et al. The Williams syndrome chromosome 7q11.23 hemideletion confers hypersocial, anxious personality coupled with altered insula structure and function. Proc. Natl Acad. Sci. USA 109, E860–E866 (2012).
Marenco, S. et al. Genetic contributions to white matter architecture revealed by diffusion tensor imaging in Williams syndrome. Proc. Natl Acad. Sci. USA 104, 15117–15122 (2007).
Meyer-Lindenberg, A. et al. Neural correlates of genetically abnormal social cognition in Williams syndrome. Nat. Neurosci. 8, 991–993 (2005).
Makinodan, M., Rosen, K. M., Ito, S. & Corfas, G. A critical period for social experience-dependent oligodendrocyte maturation and myelination. Science 337, 1357–1360 (2012).
Miller, V. M. et al. Novel inter-hemispheric white matter connectivity in the BTBR mouse model of autism. Brain Res. 1513, 26–33 (2013).
Hibbits, N., Pannu, R., Wu, T. J. & Armstrong, R. C. Cuprizone demyelination of the corpus callosum in mice correlates with altered social interaction and impaired bilateral sensorimotor coordination. ASN Neuro 1, e00013 (2009).
Xu, H., Yang, H. J., McConomy, B., Browning, R. & Li, X. M. Behavioral and neurobiological changes in C57BL/6 mouse exposed to cuprizone: effects of antipsychotics. Front. Behav. Neurosci. 4, 8 (2010).
Fairless, A. H. et al. Low sociability is associated with reduced size of the corpus callosum in the BALB/cJ inbred mouse strain. Brain Res. 1230, 211–217 (2008).
Seidl, A. H. Regulation of conduction time along axons. Neuroscience 276, 126–134 (2014).
Paul, L. K. et al. Agenesis of the corpus callosum: genetic, developmental and functional aspects of connectivity. Nat. Rev. Neurosci. 8, 287–299 (2007).
Blanco, J. E., Anderson, K. D. & Steward, O. Recovery of forepaw gripping ability and reorganization of cortical motor control following cervical spinal cord injuries in mice. Exp. Neurol. 203, 333–348 (2007).
Barak, B. et al. Neuron-specific expression of tomosyn1 in the mouse hippocampal dentate gyrus impairs spatial learning and memory. Neuromolecular Med. 15, 351–363 (2013).
Crawford, D. K. et al. Oestrogen receptor β ligand: a novel treatment to enhance endogenous functional remyelination. Brain 133, 2999–3016 (2010).
Crawford, D. K., Mangiardi, M. & Tiwari-Woodruff, S. K. Assaying the functional effects of demyelination and remyelination: revisiting field potential recordings. J. Neurosci. Methods 182, 25–33 (2009).
Asante, C. O. & Martin, J. H. Differential joint-specific corticospinal tract projections within the cervical enlargement. PLoS One 8, e74454 (2013).
Wang, X. et al. Deconstruction of corticospinal circuits for goal-directed motor skills. Cell 171, 440–455.e14 (2017).
Acknowledgements
The authors gratefully acknowledge M.E. Greenberg for advice and guidance and E. Okun, P. Monteiro, A. Krol, J. Wilde, A. Marco and S. Alon for insightful comments on the manuscript. H. Zaniewski and J. Wang provided technical help. Human tissue was obtained from the NIH NeuroBioBank at the University of Maryland. We thank the donors of the brain tissue and their families for their invaluable donations for the advancement of scientific understanding. This work is supported by a grant from the Simons Foundation (grant no. SFARI 240005 to G.F.), the Tang-Yang Center for Autism Research at MIT, the Poitras Center for Psychiatric Disorders Research at MIT, the Stanley Center for Psychiatric Research at Broad Institute of MIT and Harvard and the Simons Center for the Social Brain at MIT. B.B. was supported by postdoctoral fellowships from the Simons Center for the Social Brain at MIT and the Autism Science Foundation.
Author information
Authors and Affiliations
Contributions
B.B., Z.Z., Y.Liu., Z.H. and G.F. designed the experiments and wrote the manuscript. B.B. collected, analyzed and interpreted the results from studies related to behavior, biochemistry, genomics, immunofluorescence and pharmacology. M.E. collected and analyzed the results of immunofluorescence studies. Z.Z. collected, analyzed and interpreted the results from electrophysiological studies from the CST and corpus callosum. K.Q. collected, analyzed and interpreted the results from electrophysiological studies to characterize the properties of the basal membrane. K.M.L. collected and analyzed the results from the human immunostaining studies. Y.Liu collected, analyzed and interpreted the results from the fine motor skills and CST immunostaining studies. S.S.T. and A.N. collected, analyzed and interpreted the results from the clemastine study and the human immunofluorescence studies. D.W. administered the clemastine. Y.Li collected and analyzed the CST immunostaining studies. D.B. donated the Gtf2i loxP mice. B.B., G.L.B., Z.Z., Y.Liu, Z.H. and G.F. interpreted the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Table 1
Mouse cortex bulk RNAseq results
Supplementary Table 2
Human subjects information
Rights and permissions
About this article
Cite this article
Barak, B., Zhang, Z., Liu, Y. et al. Neuronal deletion of Gtf2i, associated with Williams syndrome, causes behavioral and myelin alterations rescuable by a remyelinating drug. Nat Neurosci 22, 700–708 (2019). https://doi.org/10.1038/s41593-019-0380-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0380-9
This article is cited by
-
Transposons in the Williams–Beuren Syndrome Critical Region are Associated with Social Behavior in Assistance Dogs
Behavior Genetics (2024)
-
Functional myelin in cognition and neurodevelopmental disorders
Cellular and Molecular Life Sciences (2024)
-
Dorsal visual stream and LIMK1: hemideletion, haplotype, and enduring effects in children with Williams syndrome
Journal of Neurodevelopmental Disorders (2023)
-
Neuronal Gtf2i deletion alters mitochondrial and autophagic properties
Communications Biology (2023)
-
In individuals with Williams syndrome, dysregulation of methylation in non-coding regions of neuronal and oligodendrocyte DNA is associated with pathology and cortical development
Molecular Psychiatry (2023)