Abstract
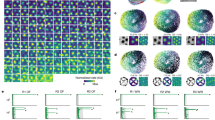
The network of grid cells in the medial entorhinal cortex (MEC) forms a fixed reference frame for mapping physical space. The mechanistic origin of the grid representation is unknown, but continuous attractor network models explain multiple fundamental features of grid cell activity. An untested prediction of these models is that the grid cell network should exhibit an activity correlation structure that transcends behavioral states. By recording from MEC cell ensembles during navigation and sleep, we found that spatial phase offsets of grid cells predict arousal-state-independent spike rate correlations. Similarly, state-invariant correlations between conjunctive grid–head direction and pure head direction cells were predicted by their head direction tuning offsets during awake behavior. Grid cells were only weakly correlated across grid modules, and module scale relationships disintegrated during slow-wave sleep, suggesting that grid modules function as independent attractor networks. Collectively, our observations imply that network states in MEC are expressed universally across brain and behavior states.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
Code for reproducing the analyses in this article is available from the corresponding authors upon reasonable request.
Data availability
The datasets generated during the current study are available from the corresponding authors upon reasonable request.
References
Hafting, T., Fyhn, M., Molden, S., Moser, M.-B. & Moser, E. I. Microstructure of a spatial map in the entorhinal cortex. Nature 436, 801–806 (2005).
O’Keefe, J. & Dostrovsky, J. The hippocampus as a spatial map. Preliminary evidence from unit activity in the freely-moving rat. Brain Res. 34, 171–175 (1971).
Taube, J. S., Muller, R. U. & Ranck, J. B. Head-direction cells recorded from the postsubiculum in freely moving rats. II Effects of environmental manipulations. J. Neurosci. 10, 436–447 (1990).
Solstad, T., Boccara, C. N., Kropff, E., Moser, M.-B. & Moser, E. I. Representation of geometric borders in the entorhinal cortex. Science 322, 1865–1868 (2008).
Lever, C., Burton, S., Jeewajee, A., O’Keefe, J. & Burgess, N. Boundary vector cells in the subiculum of the hippocampal formation. J. Neurosci. 29, 9771–9777 (2009).
Moser, E. I., Moser, M.-B. & McNaughton, B. L. Spatial representation in the hippocampal formation: a history. Nat. Neurosci. 20, 1448–1464 (2017).
Muller, R. U. & Kubie, J. L. The effects of changes in the environment on the spatial firing of hippocampal complex-spike cells. J. Neurosci. 7, 1951–1968 (1987).
Alme, C. B. et al. Place cells in the hippocampus: eleven maps for eleven rooms. Proc. Natl Acad. Sci. USA 111, 18428–18435 (2014).
Fyhn, M., Hafting, T., Treves, A., Moser, M.-B. & Moser, E. I. Hippocampal remapping and grid realignment in entorhinal cortex. Nature 446, 190–194 (2007).
Yoon, K. et al. Specific evidence of low-dimensional continuous attractor dynamics in grid cells. Nat. Neurosci. 16, 1077–1084 (2013).
Fuhs, M. C. & Touretzky, D. S. A spin glass model of path integration in rat medial entorhinal cortex. J. Neurosci. 26, 4266–4276 (2006).
McNaughton, B. L., Battaglia, F. P., Jensen, O., Moser, E. I. & Moser, M.-B. Path integration and the neural basis of the ‘cognitive map’. Nat. Rev. Neurosci. 7, 663–678 (2006).
Burgess, N., Barry, C. & O’Keefe, J. An oscillatory interference model of grid cell firing. Hippocampus 17, 801–812 (2007).
Amari, S. Dynamics of pattern formation in lateral-inhibition type neural fields. Biol. Cybern. 27, 77–87 (1977).
Burak, Y. & Fiete, I. R. Accurate path integration in continuous attractor network models of grid cells. PLoS Comput. Biol. 5, e1000291 (2009).
Tsodyks, M. & Sejnowski, T. J. Associative memory and hippocampal place cells. Int. J. Neural Syst. 6, 81–86 (1995).
McNaughton, B. L. et al. Deciphering the hippocampal polyglot: the hippocampus as a path integration system. J. Exp. Biol. 199, 173–185 (1996).
Samsonovich, A. & McNaughton, B. L. Path integration and cognitive mapping in a continuous attractor neural network model. J. Neurosci. 17, 5900–5920 (1997).
Moser, E. I. et al. Grid cells and cortical representation. Nat. Rev. Neurosci. 15, 466–481 (2014).
Welinder, P. E., Burak, Y. & Fiete, I. R. Grid cells: the position code, neural network models of activity, and the problem of learning. Hippocampus 18, 1283–1300 (2008).
Barry, C., Hayman, R., Burgess, N. & Jeffery, K. J. Experience-dependent rescaling of entorhinal grids. Nat. Neurosci. 10, 682–684 (2007).
Stensola, H. et al. The entorhinal grid map is discretized. Nature 492, 72–78 (2012).
Okun, M. et al. Diverse coupling of neurons to populations in sensory cortex. Nature 521, 511–515 (2015).
Tocker, G., Barak, O. & Derdikman, D. Grid cells correlation structure suggests organized feedforward projections into superficial layers of the medial entorhinal cortex. Hippocampus 25, 1599–1613 (2015).
Couey, J. J. et al. Recurrent inhibitory circuitry as a mechanism for grid formation. Nat. Neurosci. 16, 318–324 (2013).
Si, B. & Treves, A. A model for the differentiation between grid and conjunctive units in medial entorhinal cortex. Hippocampus 23, 1410–1424 (2013).
Peyrache, A., Lacroix, M. M., Petersen, P. C. & Buzsáki, G. Internally organized mechanisms of the head direction sense. Nat. Neurosci. https://doi.org/10.1038/nn.3968(2015).
Sargolini, F. et al. Conjunctive representation of position, direction, and velocity in entorhinal cortex. Science 312, 758–762 (2006).
Colgin, L. L., Moser, E. I. & Moser, M.-B. Understanding memory through hippocampal remapping. Trends Neurosci. 31, 469–477 (2008).
Knierim, J. J., Kudrimoti, H. S. & McNaughton, B. L. Place cells, head direction cells, and the learning of landmark stability. J. Neurosci. 15, 1648–1659 (1995).
Knierim, J. J., Kudrimoti, H. S. & McNaughton, B. L. Interactions between idiothetic cues and external landmarks in the control of place cells and head direction cells. J. Neurophysiol. 80, 425–446 (1998).
Hoydal, O. A., Skytoen, E. R., Moser, M.-B. & Moser, E. I. Object-vector coding in the medial entorhinal cortex. Preprint at bioRxiv https://doi.org/10.1101/286286 (2018).
Grosmark, A. D. & Buzsáki, G. Diversity in neural firing dynamics supports both rigid and learned hippocampal sequences. Science 351, 1440–1443 (2016).
Cohen, M. R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Ólafsdóttir, H. F., Carpenter, F. & Barry, C. Coordinated grid and place cell replay during rest. Nat. Neurosci. 19, 792–794 (2016).
Roth, F. C., Beyer, K. M., Both, M., Draguhn, A. & Egorov, A. V. Downstream effects of hippocampal sharp wave ripple oscillations on medial entorhinal cortex layer V neurons in vitro. Hippocampus 26, 1493–1508 (2016).
Dordek, Y., Soudry, D., Meir, R. & Derdikman, D. Extracting grid cell characteristics from place cell inputs using non-negative principal component analysis. eLife 5, e10094 (2016).
Stachenfeld, K. L., Botvinick, M. M. & Gershman, S. J. The hippocampus as a predictive map. Nat. Neurosci. 20, 1643–1653 (2017).
Franzius, M., Sprekeler, H. & Wiskott, L. Slowness and sparseness lead to place, head-direction, and spatial-view cells. PLoS Comput. Biol. 3, e166 (2007).
Chrobak, J. J. & Buzsáki, G. High-frequency oscillations in the output networks of the hippocampal-entorhinal axis of the freely behaving rat. J. Neurosci. 16, 3056–3066 (1996).
Csicsvari, J., Hirase, H., Mamiya, A. & Buzsáki, G. Ensemble patterns of hippocampal CA3-CA1 neurons during sharp wave-associated population events. Neuron 28, 585–594 (2000).
Patel, J., Schomburg, E. W., Berényi, A., Fujisawa, S. & Buzsáki, G. Local generation and propagation of ripples along the septotemporal axis of the hippocampus. J. Neurosci. 33, 17029–17041 (2013).
Dunn, B., Mørreaunet, M. & Roudi, Y. Correlations and functional connections in a population of grid cells. PLoS Comput. Biol. 11, e1004052 (2015).
Ólafsdóttir, H. F., Carpenter, F. & Barry, C. Task demands predict a dynamic switch in the content of awake hippocampal replay. Neuron 96, 925–935.e6 (2017).
O’Neill, J., Boccara, C. N., Stella, F., Schoenenberger, P. & Csicsvari, J. Superficial layers of the medial entorhinal cortex replay independently of the hippocampus. Science 355, 184–188 (2017).
Gardner, R. J. & Moser, M.-B. Multiple mechanisms for memory replay? Science 355, 131–132 (2017).
Trimper, T. B., Trettel, S. G., Hwaun, E. & Colgin, L. L. Methodological caveats in the detection of coordinated replay between place cells and grid cells. Front. Syst. Neurosci. https://doi.org/10.3389/fnsys.2017.00057 (2017).
Si, B., Romani, S. & Tsodyks, M. Continuous attractor network model for conjunctive position-by-velocity tuning of grid cells. PLoS Comput. Biol. 10, e1003558 (2014).
Kropff, E., Carmichael, J. E., Moser, M.-B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Hinman, J. R., Brandon, M. P., Climer, J. R., Chapman, G. W. & Hasselmo, M. E. Multiple running speed signals in medial entorhinal cortex. Neuron 91, 666–679 (2016).
Navratilova, Z., Giocomo, L. M., Fellous, J.-M., Hasselmo, M. E. & McNaughton, B. L. Phase precession and variable spatial scaling in a periodic attractor map model of medial entorhinal grid cells with realistic after-spike dynamics. Hippocampus 22, 772–789 (2012).
Si, B., Kropff, E. & Treves, A. Grid alignment in entorhinal cortex. Biol. Cybern. 106, 483–506 (2012).
Jun, J. J. et al. Fully integrated silicon probes for high-density recording of neural activity. Nature 551, 232–236 (2017).
Pachitariu, M., Steinmetz, N., Kadir, S., Carandini, M. & Harris, K. D. Kilosort: realtime spike-sorting for extracellular electrophysiology with hundreds of channels. Preprint at bioRxiv https://doi.org/10.1101/061481(2016).
Gray, C. M., Maldonado, P. E., Wilson, M. & McNaughton, B. Tetrodes markedly improve the reliability and yield of multiple single-unit isolation from multi-unit recordings in cat striate cortex. J. Neurosci. Methods 63, 43–54 (1995).
Henze, D. A. et al. Intracellular features predicted by extracellular recordings in the hippocampus in vivo. J. Neurophysiol. 84, 390–400 (2000).
Acharya, L., Aghajan, Z. M., Vuong, C., Moore, J. J. & Mehta, M. R. Causal influence of visual cues on hippocampal directional selectivity. Cell 164, 197–207 (2016).
Park, I. M., Meister, M. L. R., Huk, A. C. & Pillow, J. W. Encoding and decoding in parietal cortex during sensorimotor decision-making. Nat. Neurosci. 17, 1395–1403 (2014).
Phillips, K. G. et al. Decoupling of sleep-dependent cortical and hippocampal interactions in a neurodevelopmental model of schizophrenia. Neuron 76, 526–533 (2012).
Acknowledgements
We thank A. Z. Vollan for assistance with recordings, B. A. Dunn for insightful scientific discussions, and A. M. Amundsgård, K. Haugen, E. Kråkvik, and H. Waade for various technical assistance. The work was supported by an Advanced Investigator Grant from the European Research Council (GRIDCODE—grant no. 338865), the European Commission’s FP7 FET Proactive Programme on Neuro-Bio-Inspired Systems (GRIDMAP—Grant Agreement 600725), a NEVRONOR grant from the Research Council of Norway (grant no. 226003), the Centre of Excellence scheme and the National Infrastructure Scheme of the Research Council of Norway (Centre for Neural Computation, grant number 223262; NORBRAIN1, grant number 197467), the Louis Jeantet Prize, the Körber Prize, and the Kavli Foundation.
Author information
Authors and Affiliations
Contributions
R.J.G., M.-B.M., and E.I.M. designed the experiment; R.J.G. collected most data but used additional recordings from L.L. and T.W.; R.J.G. analyzed the data; R.J.G. wrote the paper, assisted by M.-B.M. and E.I.M. All authors read and commented on the paper. M.-B.M. and E.I.M. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information Nature Neuroscience thanks Adrien Peyrache, Francesca Sargolini, and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Basic LFP and single-unit spiking characteristics.
a: Example traces of MEC activity during RUN, SWS and REM. From top to bottom: combined spike rate of all recorded units; spike rasters for 8 grid cells; wideband LFP signal. b: MEC LFP power spectra generated from the recording shown in (a) (n = 1 recording, durations in minutes: RUN 34.0, SWS 222, REM 30.4). The power spectra were calculated in 5-second windows using the multitaper method, using all available time periods for each state. The lines and shaded areas represent mean ± S.E.M. c: Distributions of mean spike rates of all units during RUN, SWS and REM (n = 138 grid cells, 95 HD cells, 39 grid/HD cells). Box indicates median and quartiles; whiskers indicate 1.5 × interquartile range below or above the first and third quartiles respectively. d: Comparison of mean spike rates of grid, HD and grid/HD cells between RUN, SWS and REM. Cell types are displayed in the same vertical order as in (c). e: Distribution of functional unit types among parahippocampal subregions. “PaS” refers to parasubiculum; “POR-RSC-PaS” refers to the transitional area between postrhinal cortex, retrosplenial cortex and PaS.
Supplementary Figure 2 Effect of fluctuating population rate on Pearson and GLM coupling measures.
a: Demonstration of effect of global population rate modulation on coupling. a1 shows spike trains from two simulated neurons producing inhomogeneous Poisson spike trains (black rasters, top) with rates oppositely modulated by a signal of interest (grey line). The Pearson and GLM spike rate cross-coupling (CC) of the spike trains (bottom) shows the time course of the relationship between the two neurons’ spike rates. a2 shows the same situation as (a1), but with the two neurons’ spiking also positively modulated by global population rate fluctuations (red line). The population rate strongly impacts the Pearson CC, whose zero-lag deflection changes from negative to positive. Conversely, the GLM CC retains a similar shape to the CC in (a1). b: Relationship between zero-lag Pearson (r-value) and zero-lag GLM coupling coefficient (β0) for grid–grid (top) and HD–HD (bottom) cell pairs. Note that for cell pairs with a near-zero Pearson r-value, the GLM coupling is typically negative, demonstrating the GLM method’s ability to recover negative couplings by discounting the cell pair’s common positive coupling to the population rate. c: Coupling of single unit spiking to population spike rate. A Poisson GLM was fitted to the spike rate of each unit, with the population spike rate as the single regressor. The plots show the resultant distributions of zero-lag GLM couplings for the population rate (βPOP). Population rates were z-score transformed before fitting the model, such that results were comparable across different mean population rates. Note the almost exclusively positive βPOP-values. Firing rates of units in each neuronal class were overwhelmingly positively coupled to the population rate (P < 10-10, z > 6 in all states, binomial sign test, n = 138 grid cells, 95 HD cells, 39 grid/HD cells).
Supplementary Figure 3 Coupling between pairs of grid cells.
a: Scatter plots showing the lag time of the largest peak for each GLM cross-coupling (CC), versus the zero-lag GLM coupling (β0). For cell pairs with positive β0, the largest positive peak was identified; for cell pairs with negative β0, the largest negative peak was identified. Most of the peaks falling far from zero lag are for cell pairs with small absolute β0, indicating weak coupling. b: Histograms of the peak times shown in (a). c: Histograms of β0 distributions (n = 138 grid cells, 95 HD cells, 39 grid/HD cells). d: Relationship between the grid phase offset ϕG of grid–grid cell pairs and their zero-lag coupling, measured with the Pearson correlation coefficient. The r-values shown are Spearman Rank (all P-values < 10−9, n = 135 grid–grid pairs). Note the absence of strong negative couplings, in accordance with the positive bias expected from common positive coupling to the population rate (Supplementary Fig. 2). Note the lack of strong negative couplings, in contrast to the strong correlations evident when using the GLM approach in Fig. 1c. e: Relationship between grid–grid cell pair rate map similarity (measured as the r-value of the Pearson correlation between each cell pair’s respective rate maps), and β0. f: Colour-mapped CC for all intramodular grid–grid cell pairs, ranked by the cell-pair spatial phase offset ϕG. Each row of the matrix is the colour-coded CC of one cell pair. These results are equivalent to those in Fig. 1b, but here the Pearson correlation coefficient is plotted instead of β. g: Spatial firing patterns and cross-coupling of an example grid–grid/HD cell pair, as in Fig. 1a. The marked asymmetry of the RUN cross-correlorams is likel
Supplementary Figure 4 Signatures of hexagonal geometry in grid-cell coupling.
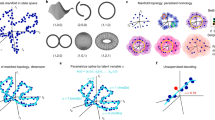
a: A simulated triangular grid pattern. Each grey dot represents a vertex (receptive field) of the grid. The hexagonal tiles surrounding the vertices are Voronoi cells which form the phase space of the grid pattern. The black arrow marked “s” denotes the distance between adjacent vertices (the grid spacing). Within the central tile are drawn a series of concentric rings, indicating zones corresponding to different values of ϕG, the magnitude of a grid phase offset vector. Note that the phase tile border encroaches on the outermost ϕG zone, meaning that the largest values of ϕG have a nonuniform radial distribution. b: Illustration of the four spatial kernel types used to simulate grid firing fields. c: Correlation of simulated pairs of grid patterns as a function of the phase offset between the two patterns. Each position on the hexagonal phase tile represents a particular phase offset of one grid from the other. The colour at each point indicates the Pearson correlation r-value for two grid patterns with that phase offset. Each plot shows the result of simulations using a different spatial kernel type. In each case, the normalized field width parameter w was 0.19, equal to the empirically estimated value. Contour lines follow equal correlation values. Note the circle formed by the phases where correlation is zero (dashed black line), and the non-concentric deviations in the correlation pattern near the phase tile’s edge. Inset: examples of simulated rate maps. d: Relationship of grid pattern Pearson correlation r-values with ϕG and the grid field width. For all kernel types, the field width is defined as the radius enclosing 50% of the field’s mass, as a proportion of the grid spacing. The dashed line traces the path of the value of ϕG at which RMS crosses zero (ϕG0) Note the convergence of ϕG0 on a value of approximately 0.32 as field width increases. The black circle on the first plot indicates ϕG0 at the empirically estimated average field width for the sample of grid cells (see i). e: Negative correlations predominate on the grid phase tile. Plotted is the proportion of the area of the grid phase tile (as shown in (c)) containing positive Pearson r-values, for grid patterns of different field widths and spatial kernels. Note that for all widths and kernel types, the fraction of positive r-values is below 50%. f: Empirical relationship between ϕG and spatial rate-map Pearson correlation r-values among grid–grid pairs (black dots). The grey line indicates the values obtained from simulated rate maps for grid–grid pairs with uniform-randomly distributed phases. g: The empirical transition between positive and negative couplings occurs at a similar phase offset during all states. Plotted are Matthews correlation coefficient (MCC) values which indicate how accurately the distribution of grid–grid pair GLM β0-values is separated into positive and negative values by splitting at different values of ϕG (see Methods). The vertical dotted grey line indicates the ϕG0-value of 0.32 determined from simulations, as shown in (d). The shaded regions show 95% bootstrap confidence intervals. h: ϕG -values that maximise the MCC shown in (g), with 95% bootstrap confidence intervals. The ϕG0 values for each state (RUN 0.32, SWS 0.33, REM 0.33) are similar to the value of 0.32 obtained from simulations (see d) (P > 0.05 in all cases, bootstrap, n = 135 grid–grid pairs). i: Estimated spacing and field width (w) for all grid cells. Field width was estimated with an algorithm which fitted a mixture of 2D Gaussian functions to a grid cell’s 2D firing rate map. The field width was estimated as the value of the Gaussian sigma parameter which achieved the best fit to the cell’s rate map, as a proportion of the cell’s grid spacing.
Supplementary Figure 5 Effect of intersection/union (I/U) module classification threshold on zero-lag coupling comparisons.
a: Classifying a pair of grid cells as intramodular or transmodular was based on geometry of the ellipses (black) fitted to the six innermost peaks of each cell’s spatial autocorrelogram. The ratio between the areas of the intersection (I) and the union (U) of their respective ellipses. b: Illustration of the module classification process. The autocorrelograms of two cells’ 2D rate maps during RUN are shown in superimposition. Ellipses are fitted to each autocorrelogram’s innermost ring of peaks, and the intersection/union area ratio (I/U) of the two ellipses is calculated (a). The I/U ratio of the cell pair shown is below the threshold of 0.75 that defines them as originating from separate modules. To ensure that the outcomes shown in Fig. 2c were not dependent on the particular threshold value used, we also examined the results of using different threshold values (e,f). c: Distribution of ellipse I/U ratios across all grid–grid pairs. The stippled vertical line indicates the module classification threshold. d: Relationship of grid–grid cell pairs’ GLM zero lag coupling coefficient (β0) values during SWS with the I/U ratio of their fitted ellipses. Note wider range of β0 -values for high I/U ratios, in orange (intramodular comparisons). This result was not dependent on the particular threshold chosen for classifying module membership, since we observed the same outcome across a wide range of thresholds (e,f). e: Relationship of I/U ratio module classification threshold with the two-sample Kolmogorov-Smirnov test P-value (two-sided, n = 198 grid–grid pairs) when comparing the cumulative distributions of zero-lag couplings (β0) for intramodular and transmodular pairs, as shown in (c). Each line shows the resultant P-values for each threshold in a given arousal state. The bars at the top of the plot indicate the threshold values for which P < 0.05. The dashed black line indicates the actual threshold that was used in analyses (0.75). f: Same as (e), but showing P-values for the 2-sample Student’s t-test.
Supplementary Figure 6 Spike rate cross-coupling between grid and conjunctive grid/HD cells.
a: Colour-mapped GLM cross-coupling (CC) for all intramodular grid–grid/HD pairs, ranked by their spatial phase offset magnitude ϕG. Each row of the matrix is the CC of one cell pair. The black line to the right of the matrix indicates the value of ϕG for each pair. b: Relationship between grid phase magnitude ϕG and GLM zero-lag coupling (β0) during each state. Displayed statistics are for Spearman rank correlation (n = 16 grid–grid/HD pairs). c: Same as (a), but with cell pairs ranked in order of the β0-value during RUN. d: Same as (b), but showing relationship between β0 during RUN and β0 during SWS and REM.
Supplementary Figure 7 Stability of spike rate coupling in MEC and CA1 cell pairs.
a: Same as Fig. 5a, with subsampled spike rates. Since mean spike rates are known to influence the magnitude of observed correlations34, it is possible that differences in observed coupling stability could merely reflect differences in mean spike rate. To discount this possibility, we randomly subsampled each unit’s mean spike rate to 0.2 Hz during each 15-minute window of the first hour of SWS. β0-values are compared between the first 15 (β0_0-15) and last 15 minutes (β0_45-60) of the first hour of SWS. Displayed r-values are for Spearman rank correlation. b: Stability of β0 for different Gaussian kernel widths, for intramodular grid–grid, HD–HD and CA1–CA1 cell pairs, calculated as the correlation between the cell pairs’ β0_0-15 and β0_45-60 values. The shaded regions indicate 95% bootstrap confidence intervals. c: Coupling stability for CA1 putative pyramidal cell pairs within each recording. Each plot shows, for one recording, all CA1 pyramidal cell pair GLM spiking correlations from the first 15 (β0_0-15) and last 15 minutes (β0_45-60) of the first hour of SWS.
Supplementary Figure 8 Temporal relationships between MEC sharp-waves (MEC-SW), CA1 ripples and MEC spike rates.
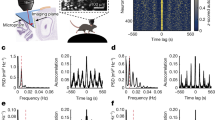
a: Event-triggered LFP traces from an example recording. a1 shows MEC 10–40 Hz LFP traces anchored to the times of CA1-SWR events detected during SWS (n = 4141 events). Events are ordered from lowest-amplitude (bottom) to highest-amplitude (top). The average response is shown above. a2 same as (a1), but showing CA1 125–250 Hz amplitude, anchored to the times of identified MEC-SW events (n = 1935 events). Note that CA1-ripple and MEC-SW events are precisely aligned, but correspondence only exists for a subset of events. b: Average LFP power spectrograms from CA1 and MEC, anchored respectively to MEC-SW (b1) and CA1-ripples (b2). The horizontal stippled lines indicate the 10–40 Hz sharp-wave band. c: Amplitude relationship between synchronous CA1-ripple and MEC-SW activity from an example recording, demonstrating weak correlation between MEC and CA1 event amplitudes. c1: amplitudes of detected CA1-ripple events are compared with amplitude of the 10–40 Hz bandpass-filtered MEC LFP. The horizontal red line indicates the mean MEC-SW amplitude across the perievent window. c2: same as (c1), but showing detected MEC-SW events, versus the synchronous CA1-ripple amplitude. r- and P- values for Spearman rank correlation are shown. The horizontal red line indicates the mean CA1-ripple amplitude across the perievent window. d: Event-triggered LFP histogram, d1: MEC 10-40 Hz LFP values at the times of detected CA1 ripples. d2: CA1 125-250 Hz LFP amplitude at the time of detected MEC-SW. e: Time-course of MEC-SW-triggered spike rates across all units (n = 95 HD cells, 134 grid cells, 39 grid/HD cells). Plotted values are mean ± 95% bootstrap confidence intervals. The grey bar indicates the ± 25 ms perievent time range defining the MEC-SW period used in Fig. 6e. f: Time course of mean spike rates anchored to MEC-SW events. Each row represents the mean MEC-SW-triggered spike rate of one unit. Rates were normalized by dividing by the mean value across the perievent time window. Units are displayed ordered by the timing of their maximal firing rate. g–j: same as (a–d), but showing data from a separate paired MEC–CA1 recording (n = 402 CA1-ripples, 1507 MEC-SW).
Supplementary information
Rights and permissions
About this article
Cite this article
Gardner, R.J., Lu, L., Wernle, T. et al. Correlation structure of grid cells is preserved during sleep. Nat Neurosci 22, 598–608 (2019). https://doi.org/10.1038/s41593-019-0360-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0360-0
This article is cited by
-
A consistent map in the medial entorhinal cortex supports spatial memory
Nature Communications (2024)
-
Local origin of excitatory–inhibitory tuning equivalence in a cortical network
Nature Neuroscience (2024)
-
Toroidal topology of population activity in grid cells
Nature (2022)
-
Attractor and integrator networks in the brain
Nature Reviews Neuroscience (2022)
-
Perineuronal nets stabilize the grid cell network
Nature Communications (2021)