Abstract
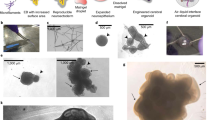
Neural organoids have the potential to improve our understanding of human brain development and neurological disorders. However, it remains to be seen whether these tissues can model circuit formation with functional neuronal output. Here we have adapted air–liquid interface culture to cerebral organoids, leading to improved neuronal survival and axon outgrowth. The resulting thick axon tracts display various morphologies, including long-range projection within and away from the organoid, growth-cone turning, and decussation. Single-cell RNA sequencing reveals various cortical neuronal identities, and retrograde tracing demonstrates tract morphologies that match proper molecular identities. These cultures exhibit active neuronal networks, and subcortical projecting tracts can innervate mouse spinal cord explants and evoke contractions of adjacent muscle in a manner dependent on intact organoid-derived innervating tracts. Overall, these results reveal a remarkable self-organization of corticofugal and callosal tracts with a functional output, providing new opportunities to examine relevant aspects of human CNS development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Code availability
All custom code for the MEA analysis in this paper is publicly available at https://github.com/Timothysit/organoids. The code used for latency analysis can be accessed at the following link: https://github.com/jboulanger/stimulation-motion.
Data availability
The data that support the findings of this study are included in the Supplementary Information with the paper. All scRNA-seq data has been deposited on GEO, accession number GSE124174. Raw data (for example, raw images and electrophysiological recordings) is available upon request from the corresponding author.
References
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013).
Birey, F. et al. Assembly of functionally integrated human forebrain spheroids. Nature 545, 54–59 (2017).
Lancaster, M. A. et al. Guided self-organization and cortical plate formation in human brain organoids. Nat. Biotechnol. 35, 659–666 (2017).
Quadrato, G. et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53 (2017).
Mansour, A. A. et al. An in vivo model of functional and vascularized human brain organoids. Nat. Biotechnol. 36, 432–441 (2018).
Gähwiler, B. H., Capogna, M., Debanne, D., McKinney, R. A. & Thompson, S. M. Organotypic slice cultures: a technique has come of age. Trends Neurosci. 20, 471–477 (1997).
Renner, M. et al. Self-organized developmental patterning and differentiation in cerebral organoids. EMBO J. 36, 1316–1329 (2017).
Bak, M. & Fraser, S. E. Axon fasciculation and differences in midline kinetics between pioneer and follower axons within commissural fascicles. Development 130, 4999–5008 (2003).
Polleux, F. & Snider, W. Initiating and growing an axon. Cold Spring Harb. Perspect. Biol. 2, a001925 (2010).
Piper, M. et al. Neuropilin 1-Sema signaling regulates crossing of cingulate pioneering axons during development of the corpus callosum. Cereb. Cortex 19(Suppl. 1), i11–i21 (2009).
Chédotal, A. & Richards, L. J. Wiring the brain: the biology of neuronal guidance. Cold Spring Harb. Perspect. Biol. 2, a001917 (2010).
Shu, T. & Richards, L. J. Cortical axon guidance by the glial wedge during the development of the corpus callosum. J. Neurosci. 21, 2749–2758 (2001).
Keeble, T. R. et al. The Wnt receptor Ryk is required for Wnt5a-mediated axon guidance on the contralateral side of the corpus callosum. J. Neurosci. 26, 5840–5848 (2006).
Schroeter, M. S., Charlesworth, P., Kitzbichler, M. G., Paulsen, O. & Bullmore, E. T. Emergence of rich-club topology and coordinated dynamics in development of hippocampal functional networks in vitro. J. Neurosci. 35, 5459–5470 (2015).
Cotterill, E., Charlesworth, P., Thomas, C. W., Paulsen, O. & Eglen, S. J. A comparison of computational methods for detecting bursts in neuronal spike trains and their application to human stem cell-derived neuronal networks. J. Neurophysiol. 116, 306–321 (2016).
Streit, J., Spenger, C. & Lüscher, H. R. An organotypic spinal cord–dorsal root ganglion–]skeletal muscle coculture of embryonic rat. II. functional evidence for the formation of spinal reflex arcs in vitro. Eur. J. Neurosci. 3, 1054–1068 (1991).
Koh, T. H. & Eyre, J. A. Maturation of corticospinal tracts assessed by electromagnetic stimulation of the motor cortex. Arch. Dis. Child. 63, 1347–1352 (1988).
Daza, R. A. M., Englund, C. & Hevner, R. F. Organotypic slice culture of embryonic brain tissue. CSH Protoc. 2007, t4914 (2007).
Sorkin, R. et al. Compact self-wiring in cultured neural networks. J. Neural Eng. 3, 95–101 (2006).
Gonzalez, C. et al. Modeling amyloid beta and tau pathology in human cerebral organoids. Mol. Psychiatry 23, 2363–2374 (2018).
Qian, X. et al. Brain-region-specific organoids using mini-bioreactors for modeling ZIKV exposure. Cell 165, 1238–1254 (2016).
Rezakhaniha, R. et al. Experimental investigation of collagen waviness and orientation in the arterial adventitia using confocal laser scanning microscopy. Biomech. Model. Mechanobiol. 11, 461–473 (2012).
Mátés, L. et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 41, 753–761 (2009).
Meijering, E., Dzyubachyk, O. & Smal, I. Methods for cell and particle tracking. Meth. Enzymol. 504, 183–200 (2012).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Watanabe, M. et al. Self-organized cerebral organoids with human-specific features predict effective drugs to combat Zika virus infection. Cell Rep. 21, 517–532 (2017).
Pollen, A. A. et al. Molecular identity of human outer radial glia during cortical development. Cell 163, 55–67 (2015).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Camp, J. G. et al. Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl Acad. Sci. USA 112, 15672–15677 (2015).
Cutts, C. S. & Eglen, S. J. Detecting pairwise correlations in spike trains: an objective comparison of methods and application to the study of retinal waves. J. Neurosci. 34, 14288–14303 (2014).
Acknowledgements
The authors would like to thank members of the Lancaster lab for helpful discussions and D. Jabaudon for insightful comments, as well as members of the LMB mouse facility for help with timed matings and tissue collections. We also thank members of the H. McMahon laboratory for plasmids and the LMB light microscopy facility, in particular B. Sutcliffe, for assistance with imaging. We are grateful to P. Coupland and S. Ballereau (Cancer Research UK) for technical assistance and to M. Galardini and P. Beltrao (European Bioinformatics Institute) for helping with computing resources. This work was supported by the Medical Research Council MC_UP_1201/9 (to M.A.L.), European Research Council ERC STG 757710 (to M.A.L.), Medical Research Council MR/P008658/1 (to A.L.), Wellcome Trust ISSF_RRZC/115_RG89529 (to A.L.), Newton Trust RRZC/115_RG89305 (to A.L.), MRC Clinician Scientist Fellowship (to A.L.), grants from the Biotechnology and Biological Sciences Research Council (BBSRC) (to O.P.), Medical Research Council MC_UP_1201/2 (to M.T.), European Research Council ERC Starting Grant, STG 677029 (to M.T.), ERANET-NEURON Micronet consortium (to M.T.), Medical Research Council MC_UP_1201/13 (to E.D.), and HFSP CDA (to E.D.).
Author information
Authors and Affiliations
Contributions
S.L.G. planned and performed experiments, analyzed data, and wrote the manuscript. S.B.M. planned and performed MEA and stimulation experiments, analyzed data and wrote the manuscript. G.M.G. performed the organoid cell dissociation and provided quality control for the single-cell preparation, and L.M.D.W. analyzed the scRNA-seq data. L.M. performed whole-cell electrophysiology experiments and analyzed data under the supervision of M.T. T.S. analyzed MEA electrophysiology data. M.S. generated and maintained organoids and ALI-CO cultures, and analyzed data. J.B. analyzed time-lapse image data. E.D. performed and analyzed time-lapse imaging. O.P. supervised electrophysiology experiments and interpreted data. A.L. planned experiments, analyzed and interpreted data, and supervised G.M.G and L.M.D.W. M.A.L. conceived the project, planned and performed experiments, supervised the project, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Neuroscience thanks Alexander Jaworski and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Culture at the air–liquid interface maintains survival and morphology over extended time periods.
a. Representative images of SMI312 staining of day 105 whole organoids compared with ALI-COs (20 days ALI, 85 days total) used for OrientationJ analysis shown in Fig. 1d. Shown are representative images out of six samples from two independent batches each. b. Comparison of more extended cultures of 169 days total whole organoid compared with ALI-CO (100 days ALI, 169 days total). Note the more abundant neuronal staining (TUBB3) and extensive tracts in the ALI-CO at this time point compared with whole organoid. GFAP staining marks glia including astrocytes. c. Higher magnification imaging reveals a dense TUBB3+ neuropil containing axons (SMI312), dendrites (MAP2) and astrocytes (GFAP) in ALI-COs, while whole organoids exhibit disrupted morphology with decreased neuronal processes. b-c are representative images of three ALI-COs compared with three whole organoids from the same batch, with similar results. Scale bars, 500 μm (a, b) and 100 μm (c).
Supplementary Figure 2 ALI–COs exhibit synapses, interneuron populations, and electrophysiological activity.
a. Sparse labeling of a 1-year ALI-CO with Sendai-emGFP reveals extensive dendritic trees with dendritic spines (arrows). Maximum intensity projection shown. b. ALI-COs from organoids electroporated with fGFP to label neuronal processes harbour mature synapses with juxtaposed pre- (SYT1, Bassoon, SYP) and post- (Homer1, PSD95) synaptic termini (arrowheads). ALI-CO age: 55 + 40 days ALI (95 total) and 89 + 23 (112 total). c. Immunostaining for the markers VGAT and Calretinin reveals numerous interneurons and VGAT positive punctae suggesting GABA-ergic synapses. ALI-CO age: 57 + 49 day ALI. Maximum intensity projection shown. d. Staining for Somatostatin and GAD67 demonstrates the presence of other interneuron types. ALI-CO age: 57 + 49 day ALI. e. ALI-CO electroporated with fGFP reveals a pyramidal neuron with dendritic spines decorated by punctae of the presynaptic structural protein PCLO and the presence of VGAT punctae suggesting GABA-ergic synapses onto the pyramidal neuron (arrowhead). ALI-CO age: 55 + 40 day ALI. Sparse labelling in a-e with emGFP or fGFP was performed on three ALI-COs from three organoids with similar results. Staining for pre- and post-synaptic markers was performed on three samples, and staining for interneuron markers in c and d was performed on three samples. Lower magnification overview is a maximum intensity projection. f. c-Fos staining for active neurons and MAP2 (dendrites) in a 54 + 60 day ALI-CO. Representative image shown of three experiments on three independently prepared ALI-COs with similar results. g. Image of an ALI-CO after 65 days at ALI (125 days total) then placed on the 3D multielectrode array (MEA). h. Five-second trace from an individual electrode recorded from a 40 + 85 day ALI-CO with threshold (red line) for detected spikes (black hash marks above trace); inset, expanded view of a single spike. i. Heatmap of average spike frequency per electrode over 75-second recording before (top) and after (bottom) application of 2 μM tetrodotoxin (TTX) showing spatial arrangement of spontaneous activity. j. Rasterplot of spike frequency (averaged in 5-ms time bins for each electrode) in the ALI-CO 75 seconds before (left) and after (right) TTX illustrating the temporal distribution of spontaneous activity. k. Quantification of number of active electrodes before and after TTX application in one such ALI-CO recording where TTX was applied. l. Quantification of average spike frequency (error bars are s.e.m.) in the eight active electrodes of k. before and after TTX application. Scale bars, 20 μm (a, c, d), 10 μm (b, e), 50 μm (f), 1 mm (g), 1 μm (b insets, e inset) and 5 μm (c inset, d insets).
Supplementary Figure 3 Growth cones within tracts exhibit various behaviors including turning and crossing.
a. Images of an electroporated organoid just before sectioning at day 64 (upper left), after placement at the ALI (upper right) and stills from long-term live imaging (Supplementary Video 1) of this same tissue (bottom panels) after 2 days at the ALI. Time stamp is hrs:min. b. Temporal projection showing highly disorganized axon outgrowth at this very early stage (2 days ALI, 66 days total). a-b are representative images out of 5 similarly staged and live imaged ALI-COs. c. Temporal projection of a time-lapse (Supplementary Video 5) of a region of the ALI-CO shown in Fig. 3e and boxed in d. Overlay includes tracings of individual growth cones (colored lines) and boxed regions used for kymograph analysis (right panels) showing velocities of growth cones (yellow arrowheads). Black arrowhead points to the turning point of an axon. d. Image of the same region shown in Fig. 3e at an earlier time point of 14 days at the ALI (84 days total). e. Fluorescence image of fGFP+ tracts used for directional image analysis shown in Fig. 3f. f. Image of fGFP labelled axons with costaining for DAPI in a 90 day ALI-CO (of which 26 days at ALI), revealing tracts extending away from the main body of the organoid (outlined). g. Another example of a fGFP electroporated ALI-CO (90 days total of which 20 days at ALI) displaying both internal (yellow arrows) and escaping (white arrows) tract morphologies. h. Tracing of individual growth cones within intersecting and interdigitated fiber tracts showing crossing behaviour reminiscent of decussation (arrow) (Supplementary Video 6). Age: 17 days at the ALI, 100 days total. i. Temporal projection (left panel) and growth cone tracing (right panel) showing instances of growth cone turning (arrowheads) (Supplementary Video 7). Age: 9 days at the ALI, 79 days total. c-i are representative images out of 7 similarly staged and imaged ALI-COs. Scale bars, 500 μm (a, d, e, f, g) and 100 μm (b, c, h, i).
Supplementary Figure 4 ALI–COs exhibit both intracortical and escaping tract morphologies with specific patterns of guidance cues.
a. Costaining of NRP1 and SMI312 reveals a specific subset of tracts that are positive for NRP1 suggesting callosal identity. Age: 55 days at the ALI, 147 days total. Representative image of four ALI-COs stained with similar results. b. Costaining for NRP1 and RYK reveals a subset of axons with a putative callosal identity expressing the WNT receptor RYK. Age: 19 days at the ALI, 84 days total. c. Staining of labeled tracts (fGFP+) for EphrinB1 demonstrates the presence of internally projecting axons that are also positive for this receptor (arrow). Age: 19 days at the ALI, 84 days total. b-c are representative images of three independent experiments. d. Axon (SMI312) and dendrite (MAP2) staining of an independent ALI-CO (34 days at the ALI, 89 days total) also exhibits the presence of tracts projecting away (magnified inset) from the main mass. Representative image of seven ALI-COs stained with similar results. e. Retrograde labeling with DiI crystals (black arrow, site of injection) of an escaping tract reveals the morphology of the tract (white arrow) and the location of projecting cells (magnified inset, arrowheads) within the ALI-CO (57 days at the ALI, 149 days total). DiI labeling was successful in one ALI-CO shown here. Scale bars, 200 μm (a, d inset), 100 μm (b, a inset, c inset, e inset), 50 μm (b inset) and 500 μm (c, d).
Supplementary Figure 5 Resemblances of the developmental gene-expression profile between the organoids and the fetal brain.
a. Heatmaps show normalized layer-specific gene expression in ALI-COs and b. in 12–13 week old human fetal brain cells24 ordered by pseudotime. Colour bars reflect gene expression levels. c. Matrices demonstrate Pearson correlation of gene expression for developmental layer-specific markers in the ALI-COs in comparison with d. and e. datasets5,24 derived from two other studies on 3D cerebral organoids (blue = negative correlation, yellow = positive correlation) comprising 508 cells from 9 brain organoid samples (Camp et al. study) and 66,889 cells from 19 brain organoids (Quadrato et al. study). Colour bar represents correlation values. f. Feature plots for the top 5 differentially expressed genes per cluster in the organoids. g. Featuremap shows cell group specific areas in different colours and feature plots demonstrate the distribution of cells expressing region-specific genes across multiple clusters.
Supplementary Figure 6 ALI–COs display correlated activity and can elicit muscle contractions in mouse tissue co-cultures.
a. Rasterplot of spike frequency (averaged in 5-ms time bins for each electrode) during a 6-minute recording. Network burst shown in Fig. 6b is marked by the red arrowhead. b. High resolution 50-ms traces from 12 electrodes at the time of a network burst. c. Correlation matrix shows clusters of synchronous activity. Correlated activity in the 59 electrodes of the ALI-CO shown in Figure a-b was determined using the spike time tiling coefficient as detailed in methods. d. Human ALI-CO (lower tissue)-mouse spinal (upper right) co-culture at 36 days at the ALI, before (blue box) and after (red box) axotomy (red dash line). Representative image of three such co-cultures with axotomy performed. Spontaneous contractions (blue trace, Supplementary Video 9) show a decrease in amplitude after axotomy (red trace, Supplementary Video 10). Total organoid age is 100 days. e. ALI-CO and mouse co-culture at 40 days at the ALI (total organoid age: 104 days) in which innervation has not occurred (blue box) and image after axotomy-style cut was made (red box, red dashed line) near mouse tissue. This axotomy-style cut on non-innervated co-culture was performed once. Spontaneous small amplitude muscle contractions before (blue trace) and after (red trace) cut show similar amplitude and frequency, suggesting that disturbing the mouse tissue through nearby manipulation is not sufficient to eliminate these low-amplitude contractions. Yellow box indicates ROI used for quantification of displacement due to muscle contraction. f. Image of 31-day ALI-CO-mouse co-culture (total organoid age: 79 days) with a stimulation electrode (yellow dashed line) placed on axon tracts projecting from the organoid. Large amplitude muscle contractions, recorded from ROI (yellow), were evoked by brief current pulses (upper trace, black arrow and dotted line, 3.2 mA, 120 µs-long) applied at 30, 60, 75, 90, 105, 120, 135 s. Repeated stimulation at 1 Hz (lower trace, black hash marks, 3.2 mA, 120 µs-long) could also drive muscle contractions. g. Higher magnification image of the mouse spinal section shown in white-dashed box in Fig. 6h, stained with Map2 for mouse neuronal cell bodies and dendrites (green). STEM121 stains human axon bundles (magenta) revealing innervation of the mouse spinal cord (white arrows). Maximum intensity projection shown. h. Staining of the same co-culture as in f. and i. after axotomy and fixation showing human axon tracts (STEM121+) innervating (arrows) the Map2+ mouse spinal cord. Maximum intensity projection shown. i. Precise electrode placement (yellow dashed line outlining the electrode) determines the ability to evoke muscle contractions in a 31-day mouse spinal cord-ALI-CO co-culture. Top-left image shows correct electrode placement in the innervating tract with evoked contractions traced at bottom-right (blue trace), while electrode displacement from the axon tracts (top-right panel, light blue outline), or axotomy (bottom-left, red outline) abolishes the ability to elicit a response. Red dashed line indicates the site of axotomy. Stimulation (hash marks above traces) before and after electrode displacement and axotomy was done with 1 Hz TTL-stimulated current pulses (120 µs, increasing current amplitude every 10 s for 0.2, 0.8, 1.6, 3.2 mA) with the final 1Hz TTL stimulation at 3.2 mA lasting 14 s for the control trace (dark blue hash marks). f-i are representative results of six such co-culture experiments with stimulation, two of which included axotomy. j. Confirmation of stimulation electrode placement before (dark blue outline), after TTX application (orange outline), and after wash-out (green outline) for recording shown in Fig. 6n. Shown are representative results for two independent experiments with TTX application. Yellow boxes denote the ROI used for quantification of muscle contractions. Purple-dashed box with corresponding inset shows the axonal tracts (arrowhead) joining the organoid and the mouse spinal co-culture. Scale bars, 500 μm (d, e, f, I, j), 100 μm (g, j inset) and 200 μm (h).
Supplementary information
Supplementary Video 1
Time lapse of axon outgrowth at an early stage of culture at the ALI. Related to Supplementary Fig. 3a, b and Figure 3d. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 2
Time lapse of axon outgrowth at an early stage of culture at the ALI. Related to Figure 3b. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 3
Time lapse of axon outgrowth at an intermediate stage of culture at the ALI. Related to Figure. 3c. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 4
Time lapse of axon outgrowth at a later stage of culture at the ALI with established tracts. Related to Figure. 3d. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 5
Time lapse of axon outgrowth at a later stage of culture at the ALI with established tracts. Related to Supplementary Fig. 3c. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 6
Time lapse of axon outgrowth in established tracts with decussation. Related to Supplementary Fig. 3h. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 7
Time lapse of axon outgrowth within established tracts with turning behaviors. Related to Supplementary Fig. 3i. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 8
Time lapse of outgrowth of an axon tract. Related to Figure 3g. Each time point is a maximum intensity projection. Time stamp = h:min.
Supplementary Video 9
Live imaging of spontaneous muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture. Related to Supplementary Fig. 6d, pre-axotomy. Muscle displacement trace and time shown below.
Supplementary Video 10
Live imaging of muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture post-axotomy. Related to Supplementary Fig. 6d, post-axotomy. Muscle displacement trace and time shown below.
Supplementary Video 11
Live imaging of evoked muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture. Related to Figure 6j. Muscle displacement trace and time shown below.
Supplementary Video 12
Live imaging of evoked muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture. Related to Figure 6k. Muscle displacement trace and time shown below.
Supplementary Video 13
Live imaging of evoked muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture. Related to Figure 6l. Muscle displacement trace and time shown below.
Supplementary Video 14
Live imaging of failure to evoke muscle contractions within an innervated ALI–CO–spinal cord–muscle co-culture post-axotomy. Related to Figure 6l, post-axotomy. Muscle displacement trace and time shown below.
Rights and permissions
About this article
Cite this article
Giandomenico, S.L., Mierau, S.B., Gibbons, G.M. et al. Cerebral organoids at the air–liquid interface generate diverse nerve tracts with functional output. Nat Neurosci 22, 669–679 (2019). https://doi.org/10.1038/s41593-019-0350-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0350-2
This article is cited by
-
A cell fate decision map reveals abundant direct neurogenesis bypassing intermediate progenitors in the human developing neocortex
Nature Cell Biology (2024)
-
Kirigami electronics for long-term electrophysiological recording of human neural organoids and assembloids
Nature Biotechnology (2024)
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Brain exposure to SARS-CoV-2 virions perturbs synaptic homeostasis
Nature Microbiology (2024)
-
Single-cell mapping of lipid metabolites using an infrared probe in human-derived model systems
Nature Communications (2024)