Abstract
Adaptive fear responses to external threats rely upon efficient relay of computations underlying contextual encoding to subcortical circuits. Brain-wide analysis of highly coactivated ensembles following contextual fear discrimination identified the dorsolateral septum (DLS) as a relay of the dentate gyrus–CA3 circuit. Retrograde monosynaptic tracing and electrophysiological whole-cell recordings demonstrated that DLS somatostatin-expressing interneurons (SST-INs) receive direct CA3 inputs. Longitudinal in vivo calcium imaging of DLS SST-INs in awake, behaving mice identified a stable population of footshock-responsive SST-INs during contextual conditioning whose activity tracked and predicted non-freezing epochs during subsequent recall in the training context but not in a similar, neutral context or open field. Optogenetic attenuation or stimulation of DLS SST-INs bidirectionally modulated conditioned fear responses and recruited proximal and distal subcortical targets. Together, these observations suggest a role for a potentially hard-wired DLS SST-IN subpopulation as arbiters of mobility that calibrate context-appropriate behavioral fear responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study and custom MATLAB codes are available from the corresponding author upon reasonable request.
References
Likhtik, E. & Gordon, J. A. Circuits in sync: decoding theta communication in fear and safety. Neuropsychopharmacology 39, 235–236 (2014).
Mahan, A. L. & Ressler, K. J. Fear conditioning, synaptic plasticity and the amygdala: implications for posttraumatic stress disorder. Trends Neurosci. 35, 24–35 (2012).
Risold, P. Y. & Swanson, L. W. Structural evidence for functional domains in the rat hippocampus. Science 272, 1484–1486 (1996).
Risold, P. Y. & Swanson, L. W. Connections of the rat lateral septal complex. Brain Res. Rev. 24, 115–195 (1997).
Sheehan, T. P., Chambers, R. A. & Russell, D. S. Regulation of affect by the lateral septum: implications for neuropsychiatry. Brain Res. Rev. 46, 71–117 (2004).
Risold, P. Y. & Swanson, L. W. Chemoarchitecture of the rat lateral septal nucleus. Brain Res. Rev. 24, 91–113 (1997).
Zhao, C., Eisinger, B. & Gammie, S. C. Characterization of GABAergic neurons in the mouse lateral septum: a double fluorescence in situ hybridization and immunohistochemical study using tyramide signal amplification. PLoS One 8, e73750 (2013).
Luo, A. H., Tahsili-Fahadan, P., Wise, R. A., Lupica, C. R. & Aston-Jones, G. Linking context with reward: a functional circuit from hippocampal CA3 to ventral tegmental area. Science 333, 353–357 (2011).
Calandreau, L., Desgranges, B., Jaffard, R. & Desmedt, A. Switching from contextual to tone fear conditioning and vice versa: the key role of the glutamatergic hippocampal-lateral septal neurotransmission. Learn. Mem. 17, 440–443 (2010).
Thomas, E., Yadin, E. & Strickland, C. E. Septal unit activity during classical conditioning: a regional comparison. Brain Res. 547, 303–308 (1991).
Bender, F. et al. Theta oscillations regulate the speed of locomotion via a hippocampus to lateral septum pathway. Nat. Commun. 6, 8521 (2015).
Tingley, D. & Buzsáki, G. Transformation of a spatial map across the hippocampal-lateral septal circuit. Neuron 98, 1229–1242.e5 (2018).
Anthony, T. E. et al. Control of stress-induced persistent anxiety by an extra-amygdala septohypothalamic circuit. Cell 156, 522–536 (2014).
Guzmán, Y. F. et al. Fear-enhancing effects of septal oxytocin receptors. Nat. Neurosci. 16, 1185–1187 (2013).
Menon, R. et al. Oxytocin signaling in the lateral septum prevents social fear during lactation. Curr. Biol. 28, 1066–1078.e6 (2018).
Shin, S. et al. Drd3 signaling in the lateral septum mediates early life stress-induced social dysfunction. Neuron 97, 195–208.e6 (2018).
McHugh, T. J. et al. Dentate gyrus NMDA receptors mediate rapid pattern separation in the hippocampal network. Science 317, 94–99 (2007).
Sahay, A. et al. Increasing adult hippocampal neurogenesis is sufficient to improve pattern separation. Nature 472, 466–470 (2011).
Besnard, A., Laroche, S. & Caboche, J. Comparative dynamics of MAPK/ERK signalling components and immediate early genes in the hippocampus and amygdala following contextual fear conditioning and retrieval. Brain Struct. Funct. 219, 415–430 (2014).
Wheeler, A. L. et al. Identification of a functional connectome for long-term fear memory in mice. PLoS Comput. Biol. 9, e1002853 (2013).
Fanselow, M. S. & Dong, H. W. Are the dorsal and ventral hippocampus functionally distinct structures? Neuron 65, 7–19 (2010).
Kim, Y. et al. Brain-wide maps reveal stereotyped cell-type-based cortical architecture and subcortical sexual dimorphism. Cell 171, 456–469.e22 (2017).
Wickersham, I. R., Finke, S., Conzelmann, K. K. & Callaway, E. M. Retrograde neuronal tracing with a deletion-mutant rabies virus. Nat. Methods 4, 47–49 (2007).
Sun, Y. et al. Cell-type-specific circuit connectivity of hippocampal CA1 revealed through Cre-dependent rabies tracing. Cell Reports 7, 269–280 (2014).
Seidler, B. et al. A Cre-loxP-based mouse model for conditional somatic gene expression and knockdown in vivo by using avian retroviral vectors. Proc. Natl Acad. Sci. USA 105, 10137–10142 (2008).
Resendez, S. L. et al. Visualization of cortical, subcortical and deep brain neural circuit dynamics during naturalistic mammalian behavior with head-mounted microscopes and chronically implanted lenses. Nat. Protoc. 11, 566–597 (2016).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Thomas, E., Burock, D., Knudsen, K., Deterding, E. & Yadin, E. Single unit activity in the lateral septum and central nucleus of the amygdala in the elevated plus-maze: a model of exposure therapy? Neurosci. Lett. 548, 269–274 (2013).
Frankland, P. W., Cestari, V., Filipkowski, R. K., McDonald, R. J. & Silva, A. J. The dorsal hippocampus is essential for context discrimination but not for contextual conditioning. Behav. Neurosci. 112, 863–874 (1998).
Strange, B. A., Witter, M. P., Lein, E. S. & Moser, E. I. Functional organization of the hippocampal longitudinal axis. Nat. Rev. Neurosci. 15, 655–669 (2014).
Bannerman, D. M. et al. Regional dissociations within the hippocampus--memory and anxiety. Neurosci. Biobehav. Rev. 28, 273–283 (2004).
Kheirbek, M. A. et al. Differential control of learning and anxiety along the dorsoventral axis of the dentate gyrus. Neuron 77, 955–968 (2013).
Besnard, A. & Sahay, A. Adult hippocampal neurogenesis, fear generalization, and stress. Neuropsychopharmacology 41, 24–44 (2016).
Guo, N. et al. Dentate granule cell recruitment of feedforward inhibition governs engram maintenance and remote memory generalization. Nat. Med. 24, 438–449 (2018).
Raam, T., McAvoy, K. M., Besnard, A., Veenema, A. H. & Sahay, A. Hippocampal oxytocin receptors are necessary for discrimination of social stimuli. Nat. Commun. 8, 2001 (2017).
Roy, D. S. et al. Distinct neural circuits for the formation and retrieval of episodic memories. Cell 170, 1000–1012.e19 (2017).
Cai, D. J. et al. A shared neural ensemble links distinct contextual memories encoded close in time. Nature 534, 115–118 (2016).
Carus-Cadavieco, M. et al. Gamma oscillations organize top-down signalling to hypothalamus and enable food seeking. Nature 542, 232–236 (2017).
Kepecs, A. & Fishell, G. Interneuron cell types are fit to function. Nature 505, 318–326 (2014).
Dehorter, N. et al. Tuning of fast-spiking interneuron properties by an activity-dependent transcriptional switch. Science 349, 1216–1220 (2015).
Gielow, M. R. & Zaborszky, L. The input-output relationship of the cholinergic basal forebrain. Cell Reports 18, 1817–1830 (2017).
Poulin, J. F., Tasic, B., Hjerling-Leffler, J., Trimarchi, J. M. & Awatramani, R. Disentangling neural cell diversity using single-cell transcriptomics. Nat. Neurosci. 19, 1131–1141 (2016).
Seidenbecher, T., Laxmi, T. R., Stork, O. & Pape, H. C. Amygdalar and hippocampal theta rhythm synchronization during fear memory retrieval. Science 301, 846–850 (2003).
Arriaga, M. & Han, E. B. Dedicated hippocampal inhibitory networks for locomotion and immobility. J. Neurosci. 37, 9222–9238 (2017).
Urban-Ciecko, J. & Barth, A. L. Somatostatin-expressing neurons in cortical networks. Nat. Rev. Neurosci. 17, 401–409 (2016).
Yeung, M. & Treit, D. The anxiolytic effects of somatostatin following intra-septal and intra-amygdalar microinfusions are reversed by the selective sst2 antagonist PRL2903. Pharmacol. Biochem. Behav. 101, 88–92 (2012).
Markowitz, J. E. et al. The striatum organizes 3D behavior via moment-to-moment action selection. Cell 174, 44–58.e17 (2018).
Xu, C. et al. Distinct hippocampal pathways mediate dissociable roles of context in memory retrieval. Cell 167, 961–972.e16 (2016).
Felix-Ortiz, A. C. et al. BLA to vHPC inputs modulate anxiety-related behaviors. Neuron 79, 658–664 (2013).
Yu, K. et al. The central amygdala controls learning in the lateral amygdala. Nat. Neurosci. 20, 1680–1685 (2017).
Adhikari, A. et al. Basomedial amygdala mediates top-down control of anxiety and fear. Nature 527, 179–185 (2015).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Grewe, B. F. et al. Neural ensemble dynamics underlying a long-term associative memory. Nature 543, 670–675 (2017).
Mukamel, E. A., Nimmerjahn, A. & Schnitzer, M. J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Zhou, P. et al. Efficient and accurate extraction of in vivo calcium signals from microendoscopic video data. eLife 7, e28728 (2018).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Sheintuch, L. et al. Tracking the same neurons across multiple days in Ca2+ imaging data. Cell Reports 21, 1102–1115 (2017).
Jimenez, J. C. et al. Anxiety cells in a hippocampal-hypothalamic circuit. Neuron 97, 670–683.e6 (2018).
Acknowledgements
We thank Y. Ziv, C. Harvey, M. Kheirbek and members of the Sahay lab for their comments on the manuscript. A.B. acknowledges support from 2014 NARSAD Young Investigator Award, Bettencourt-Schueller Foundation, Philippe Foundation and 2016 MGH ECOR Fund for Medical Discovery (FMD) Postdoctoral Fellowship Awards. W.F. was supported by 2013 HSCI Harvard Internship Program Award. A.R. was supported by 2017 HSCI Harvard Internship Program Award. A.S. acknowledges support from the NIH Biobehavioral Research Awards for Innovative New Scientists (BRAINS; grant R01MH104175), NIH–R01AG048908, NIH-1R01MH111729, the Ellison Medical Foundation New Scholar in Aging, the Whitehall Foundation, an Inscopix Decode award, a NARSAD Independent Investigator Award, Ellison Family Philanthropic support, the Blue Guitar Fund, a Harvard Neurodiscovery Center–MADRC Center Pilot Grant award, Alzheimer’s Association Research Grant, a Harvard Stem Cell Institute Development grant and HSCI seed grant.
Author information
Authors and Affiliations
Contributions
A.B., Y.G., M.T.K., H.T., A.K.R., T.L., W.F., and I.D. performed experiments. X.X., D.S. and L.S.Z. contributed reagents. A.S. and A.B. codeveloped the concept, analyzed data and wrote the manuscript. A.S. conceived the project and supervised all aspects of the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Brain-wide c-Fos expression associated with contextual fear discrimination (corresponds to Fig. 1).
a) Immunohistochemistry for c-Fos in mice previously exposed to context B or A’ (30 min prior) as well as a naive control group (behavioral analysis shown in Supplementary Fig. 2c-e). Note the increase in number of c-Fos positive cells in dorsal DG following exposure to context B. Representative images for 3–6 independent animals per group (single experiment). Yellow arrowheads indicate representative c-Fos immunopositive cells. Scale bar: 100 μm. b-i) Immunohistochemistry for c-Fos in mice previously exposed to context B or A’ (60 min prior) as well as a naive control group. Representative images for 5 independent animals per group (single experiment) in dorsal DG (b), dorsal CA3 (c), dorsal CA1 (d), anterior DLS medial and lateral (e), anterior CPU medial (f), BNST dorsal and ventral (g), amygdala lateral and basolateral (h) and PAG lateral, dorsolateral and dorsomedial (i). Yellow arrowheads point to representative c-Fos immunopositive cells. Scale bar: 100 μm.
Supplementary Figure 2 Immunohistochemical analysis of brain-wide c-Fos expression associated with contextual fear discrimination (corresponds to Fig. 1).
a) Brain-wide c-Fos analysis in Context B and Context A’ groups as compared to naive controls (see Fig. 1f). Detailed quantifications in brain regions categorized according to changes in levels of c-Fos immunoreactivity: no change (blue), B and A’ significantly different from naive controls (grey) and B and A’ significantly different from each other (red). Data (means ± SEM; n = 5, 5, 5 mice per group) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05, context B or A’ versus controls, #p < 0.05, context B versus context A’. b) Examples of robust inter-regional correlations between c-Fos immunopositive cells (expressed as % controls) using within and between subject design in context A’ and B. c) Schematic representation of CFCDL timeline consisting of 5 block trainings (10 days) at the end of which mice were sacrificed 30 min following exposure to the safe context for c-Fos analysis (day 11). d) Freezing behavior upon final exposure to the safe context. Data (means ± SEM; n = 6, 4 mice per group) were analyzed with unpaired Student two-tailed T-test, ***p < 0.001, context B versus A’ (Statistics detailed in Supplementary Table 1). e) c-Fos analysis in dorsal, medial and ventral DG of Context B and A’ groups as compared to naive controls. Data (means ± SEM; n = 3, 6, 4 mice per group) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Immunohistochemistry shown in Supplementary Fig. 1a; Statistics detailed in supplementary Table 1). *p < 0.05, context B or A’ versus controls, #p < 0.05, context B versus context A’. f) Schematic representation of CFCDL timeline consisting of 5 block trainings with or without footshocks (10 days) at the end of which mice were sacrificed 60 min following exposure to context A (no footshock) or B for c-Fos analysis (day 11). g) Freezing behavior upon final exposure to context A or B. Data (means ± SEM; n = 4, 4, 4, 4 mice per group) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). **p < 0.01, context A shocked throughout versus all groups. h) Immunohistochemistry for c-Fos in mice previously exposed to context B or A’ as well as a naive control group. Note the marked increase in the number of c-Fos positive cells in the DLS in context B mice. Representative images for 4 independent animals per group. Scale bar: 50 μm. i-j) Detailed quantifications in DLS medial and DLS lateral according to changes in levels of c-Fos immunoreactivity: no change (blue), B and A’ significantly different from naive controls (grey) and B and A’ significantly different from each other (red). Data (means ± SEM; n = 4, 4, 4, 4, 4 mice per group) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05, context B or A versus controls, #p < 0.05, context B shocked versus context B no-shock.
Supplementary Figure 3 DLS c-Fos expression following contextual fear recall (corresponds to Fig. 1).
a-c) c-Fos Immunohistochemistry in the anterior DLS of naïve mice (a) or mice previously exposed to context B (b) or A’(c) 60 min prior to sacrifice. Note the marked increase in number of c-Fos positive cells in context B mice. Representative images for 5 independent mice (M1–5) per group (single experiment). Scale bar: 50 μm.
Supplementary Figure 4 Cav2 mapping of DLS afferents (corresponds to Fig. 2).
a) Ai14 mice were injected with a Canine associated virus-2 encoding Cre recombinase (Cav2-Cre) virus in the DLS (red arrow). Cre-expressing cells (presynaptic partners) were retrogradely labeled with tdTomato at the site of injection (DLS) and in different brain regions such as the nucleus of the diagonal band, prefrontal cortex, anteromedial thalamus, nucleus reuniens and supramammillary nucleus. Representative images for 3 independent mice (single experiment). Scale bar: 100 μm. b) Presynaptic partners were identified in both CA3 and CA2 (as defined by RGS14 immunopositive cells). Scale bar: 100 μm. c) Quantifications of tdTomato expressing cells revealed presynaptically labeled cells in the Sub (subiculum) MS/DBN (medial septum/diagonal band nucleus), CA3/CA2, PFCx (prefrontal cortex), AM (anteromedial thalamic nucleus), SUM (supramammillary nucleus), Re (nucleus reuniens), LH (lateral hypothalamus) and CA1. Sparse tdTomato cells were found in the LM (lateral mammillary nucleus), Raphe, VTA (ventral tegmental area). No tdTomato cells were found in DG, AD (anterodorsal thalamic nucleus), PAG (periaqueductal gray), AV (anteroventral thalamic nucleus), CeA (central amygdala), CPu (caudate putamen), LA (lateral amygdala), NAcc (nucleus accumbens) and BNST (bed nucleus of stria terminalis). Data (means ± SEM; n = 3 mice per group) d) No presynaptic partners were detected in DG of Ai14 mice injected with Canine virus. Scale bar: 100 μm. e) No presynaptic partners were detected in CA3 of Ai14 mice not injected with Canine virus (Cav2-Cre). Nuclei are counterstained with DAPI. Representative images for 3 independent animals (single experiment). Scale bar: 100 μm.
Supplementary Figure 5 Characterization of distribution of SST-INs in DLS (corresponds to Fig. 2).
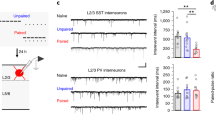
a) Immunohistochemistry for calbindin (CB), somatostatin (SST), calretinin (CR), parvalbumin (PV), neuropeptide Y (NPY), and choline acetyltransferase (ChAT) in the anterior DLS. Outlines denote medial and lateral parts of DLS and correspond to dorsal and ventral CA3 inputs to DLS (Besnard and Sahay, unpublished observations). Note the distribution of different neuronal populations throughout DLS except for ChAT, which is only expressed in the medial septum. Representative images for 3 independent animals per group (single experiment). Scale bar: 100 μm. b) Genetic labeling of Gad2 positive cells (Gad2-Cre::Ai14) and SST positive cells (SST-Cre::Ai14) with tdTomato reveals that SST-INs comprise a large proportion of DLS-INs. Representative images for 3 independent animals per group (single experiment). Scale bar: 100 μm. c-d) Detailed quantifications of tdTomato positive cells in DLS medial and DLS lateral along the rostro-caudal axis in Gad2-Cre::Ai14 and SST-Cre::Ai14 mice. Data (means ± SEM; n = 3, 3 mice per group) were analyzed using mixed factor two-way ANOVA (repeated measure over time) followed by Bonferroni’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). e) Immunohistochemistry for somatostatin (SST), endogenous expression of tdTomato in the DLS of Gad2-Cre::Ai14, cell nuclei are counterstained with DAPI. Note the overlap between SST and tdTomato (white arrowheads) confirming that SST-INs are inhibitory in the DLS. Representative images for 3 independent animals (single experiment). Scale bar: 25 μm. f) Immunohistochemistry for EYFP in the DLS, NAcc, SUM, LH, PVNH, MS/DBN and BNST of Gad2-Cre mice and SST-Cre mice injected with AAV5-DIO-ChR2-EYFP. Representative images for 3 independent animals (single experiment). Scale bar: 100 μm.
Supplementary Figure 6 Calcium imaging of SST-INs in DLS (corresponds to Fig. 3).
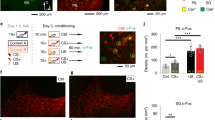
a) Immunohistochemistry for GFP (GCaMP6m), tdTomato and c-Fos in the DLS of SST-Cre::Ai14 mice injected with AAV-DJ-DIO-GCaPM6m and perfused 30 min following CFC2. Cell nuclei are counterstained with DAPI. Note the expression of cytoplasmic GCaMP6m restricted to SST-INs expressing tdTomato (white arrowheads) and nuclear expression of c-Fos in a subset of these cells (yellow arrowheads). Representative images for 4 independent animals. Scale bar: 25 μm. b) Example of calcium transients detected with CNMF-E from 7 different cells during contextual fear conditioning in context A. Note the transients observed in response to some (not all) footshocks (vertical red lines). Scale bar: x axis:10 sec, y axis:5 sd. c-g) Percent time spent in the different compartments in the open-field (c), elevated plus maze (d), time spent freezing in context A and B, freezing bout duration (f) and number of freezing bouts (g) across two consecutive recording sessions (e). Data (means ± SEM; n = 8 mice) were analyzed using mixed factor two-way ANOVA (repeated measure over time)(Statistics detailed in supplementary Table 1). h) Average calcium transient frequency (Hz) for the overall population (n = 118 cells) identified with PCA-ICA in the OF, EPM, context A and context B (average of two consecutive sessions). Data (means ± SEM; n = 118 cells per group, and data distribution represented as violin plots) were analyzed using one-way ANOVA (repeated measures over time) followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05 for specific comparisons. i) Sorting of the 118 cells detected with PCA-ICA based on calcium transients observed in response to footshocks delivered during CFC1 and CFC2. Cells that never responded to foothsocks are shown in grey, cells that responded to 1, 2 or 3 footshocks during CFC1 in blue and CFC2 in green. Cells that responded to footshocks during CFC1 and CFC2 are shown in red. Linear regression was used to test correlations between responses to footshocks during CFC1 and CFC2 across foothsock responsive cells)(Statistics detailed in supplementary Table 1). r2 = 0.00; NS. j-k) Statistical comparison for the overlap of footshock responsive (SSTFSH) cells (63/118) and footshock unresponsive (SSTnon-FSH) cells (25/118) detected with PCA-ICA across CFC1 and CFC2 with a truncated null distribution (Statistics detailed in supplementary Table 1). l) Maximum calcium transient amplitude (df/f) for the population including cells that were found active across all behavioral states as identified with CNMF-E in the OF, EPM, context A and context B. Data (means ± SEM and data distribution represented as violin plots; n = 11–87 cells) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05 for specific comparisons. Area under the curve (df/f) for the overall population (n=121 cells) as identified with CNMF-E in the OF, EPM, context A and context B. Data (means ± SEM and data distribution represented as violin plots; n = 121 cells) were analyzed using one-way ANOVA (repeated measures over time) followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05 for specific comparisons. m) Sorting of the 121 cells detected with CNMF-E based on calcium transients frequency (Hz) during movement epochs in the OF on two consecutive sessions. Inactive cells are shown in grey, cells that were active exclusively during OF1 in blue and OF2 in green. Cells that were active during OF1 and OF2 are shown in red. Linear regression was used to test correlations between calcium transient frequency during OF1 and OF2 across cells active during both sessions (Statistics detailed in supplementary Table 1). r2 = 0.182; p < 0.01. n) Statistical comparison for the overlap of cells active during movement epochs (SSTMOB) cells detected with CNMF-E across OF1 and OF2 with a truncated null distribution (Statistics detailed in supplementary Table 1). o) Sorting of the 121 cells detected with CNMF-E based on calcium transients frequency (Hz) during exploration of the open arms in the EPM on two consecutive sessions. Inactive cells are shown in grey, cells that were active exclusively during EPM1 in blue and EPM2 in green. Cells that were active during EPM1 and EPM2 are shown in red. Linear regression was used to test correlations between calcium transient frequency during EPM1 and EPM2 across cells active during both sessions (Statistics detailed in supplementary Table 1). r2 = 0.126; NS. p) Statistical comparison for the overlap of cells active during exploration of the open arms (SSTOA) cells detected with CNMF-E across EPM1 and EPM2 with a truncated null distribution (Statistics detailed in supplementary Table 1). q) Direct comparisons of decoder analysis based on calcium transient frequency local minima (Onset) and maxima (Offset) predicting behavioral state transitions (freezing) in context A using 1, 2 or 3 SSTFSH cells (red) and SSTnon-FSH (orange) cells per mouse. Data (means ± SEM; n = 6,8 mice per group) were analyzed using mixed factor two-way ANOVA (repeated measure over time)(Statistics detailed in Supplementary Table 1).
Supplementary Figure 7 Ca2+ imaging across multiple sessions (corresponds to Fig. 3).
a) Spatial maps of segmented cells from CNMF-E were tracked across multiple imaging sessions (top to bottom) for each mouse (left to right). Individual cell maps show the maximum projection for Ca2+ transients observed during each session. Red and white arrowheads point to active and inactive cells, respectively. Note that each arrowhead (4 per mouse) indicates the same cell across 10 sessions for each mouse. b) Maximum projection of cells across sessions. Scale bar: 100 μm.
Supplementary Figure 8 Cell registration across multiple sessions (corresponds to Fig. 3).
a) Spatial maps of segmented cells from CNMF-E were uploaded to CellReg, which then probabilistically tracked cells across multiple sessions. b) Cell registration performed across 10 sessions allowed to generate a reliable cell map for each mouse. Two parameters were used to verify that a cell-pair represents indeed the same cell or different cells, namely, spatial correlation and the center of mass distance.
Supplementary Figure 9 Optogenetic interrogation of SST-INs in DLS (corresponds to Figs. 4 and 5).
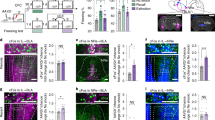
a) Immunohistochemistry for somatostatin (SST) and enhanced yellow fluorescent protein (EYFP) in the DLS of SST-Cre mice injected with AAV5-DIO-EYFP. Note the overlap between SST and EYFP (white arrowheads). Representative images for 7 independent animals. Scale bar: 25 μm. b) Post-mortem histological control (related to Fig. 4) of AAV5-DIO-EYFP (virus: blue crosses; fiber optic: blue circles) as well as DIO-eNpHR3.0 (virus: red crosses; fiber optic: red circles) along the rostro-caudal axis of DLS. c) Post-mortem histological control (related to Fig. 5) of AAV5-DIO-EYFP (virus: blue crosses; fiber optic: blue circles) as well as DIO-ChR2 (virus: red crosses; fiber optic: red circles) along the rostro-caudal axis of DLS. d-e) Freezing analysis on block 7 for the effect of light silencing SST-INs in DLS across contexts for freezing bouts number (d) and duration (e). Data (means ± SEM; n = 7,7 mice per group) were analyzed using mixed factor two-way ANOVA (Statistics detailed in supplementary Table 1). f-g) In-depth analysis of the effect of SST cell body silencing on freezing bout duration revealed a main effect on short bouts (<10 sec) in context B but not A. Data (means ± SEM; n = 7, 7 mice per group) were analyzed using mixed factor two-way ANOVA (repeated measure over time) followed by Bonferroni’s multiple comparisons post-hoc test (detailed in Supplementary Table 1). *p < 0.05, DIO-EYFP versus DIO-eNpHR3.0. h-i) Freezing analysis on block 7 for the effect of light stimulating SST-INs (15 Hz) in DLS across contexts for freezing bouts number (h) and duration (i). Data (means ± SEM; n = 7,6 mice per group) were analyzed using mixed factor two-way ANOVA (Statistics detailed in supplementary Table 1). j-k) In-depth analysis of the effect of SST cell body stimulation (15 Hz) on freezing bout duration revealed a main effect on long bouts (<20 sec) in context A but not B. Data (means ± SEM; n = 7, 6 mice per group) were analyzed using mixed factor two-way ANOVA (repeated measure over time) followed by Bonferroni’s multiple comparisons post-hoc test (Statistics detailed in Supplementary Table 1). *p < 0.05, DIO-EYFP versus DIO-ChR2. l) Effect of 15 Hz light stimulation of SST cell bodies in DLS on SST-INs c-Fos expression 60 min following exposure to context B (day 21). Immunohistochemistry for SST and c-Fos (white arrowheads) in home cage controls (2 EYFP and 2 ChR2), EYFP light stimulated and ChR2 light stimulated animals. Representative images for 4,5,5 independent animals per group. Scale bar: 25 μm. m) ChR2 (Ch) light-stimulated animals showed greater overall expression of c-Fos in SST-INs as compared to controls (C) or EYFP (E) animals. Data (means ± SEM; n = 4, 5, 5 mice per group) were analyzed using one-way ANOVA followed by Tukey’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05, DIO-EYFP or DIO-ChR2 versus controls, #p < 0.05, DIO-ChR2 versus DIO-EYFP. n) In-depth analysis of c-Fos levels in SST-INs revealed distributions of cells with varying c-Fos levels in both groups and uncovered a significant increase in c-Fos expression in ChR2 expressing mice compared to controls and EYFP injected animals. Data (means ± SEM; n = 4, 5, 5 mice per group) were analyzed using mixed factor two-way ANOVA (repeated measure over time): c-Fos intensity F (10, 110) = 417.6, p < 0.001; treatment F (2, 11) = 31.7, p < 0.001; interaction F (20,110) = 11.07, p < 0.001 followed by Bonferroni’s multiple comparisons post-hoc test (Statistics detailed in supplementary Table 1). *p < 0.05, DIO-EYFP or DIO-ChR2 versus controls, #p < 0.05, DIO-EYFP versus DIO-ChR2.
Supplementary information
Supplementary Video 1
Imaging during recall in Context A (raw calcium dynamics and df/f synced to behavior. Accelerated 4×).
Supplementary Video 2
Imaging during recall in Context B (raw calcium dynamics and df/f synced to behavior. Accelerated 4×).
Rights and permissions
About this article
Cite this article
Besnard, A., Gao, Y., TaeWoo Kim, M. et al. Dorsolateral septum somatostatin interneurons gate mobility to calibrate context-specific behavioral fear responses. Nat Neurosci 22, 436–446 (2019). https://doi.org/10.1038/s41593-018-0330-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0330-y
This article is cited by
-
Loss of mGlu5 receptors in somatostatin-expressing neurons alters negative emotional states
Molecular Psychiatry (2024)
-
TrkB-dependent regulation of molecular signaling across septal cell types
Translational Psychiatry (2024)
-
Social trauma engages lateral septum circuitry to occlude social reward
Nature (2023)
-
Lateral septum adenosine A2A receptors control stress-induced depressive-like behaviors via signaling to the hypothalamus and habenula
Nature Communications (2023)
-
Hippocampus: Molecular, Cellular, and Circuit Features in Anxiety
Neuroscience Bulletin (2023)