Abstract
Coding variants in the triggering receptor expressed on myeloid cells 2 (TREM2) are associated with late-onset Alzheimer’s disease (AD). We demonstrate that amyloid plaque seeding is increased in the absence of functional Trem2. Increased seeding is accompanied by decreased microglial clustering around newly seeded plaques and reduced plaque-associated apolipoprotein E (ApoE). Reduced ApoE deposition in plaques is also observed in brains of AD patients carrying TREM2 coding variants. Proteomic analyses and microglia depletion experiments revealed microglia as one origin of plaque-associated ApoE. Longitudinal amyloid small animal positron emission tomography demonstrates accelerated amyloidogenesis in Trem2 loss-of-function mutants at early stages, which progressed at a lower rate with aging. These findings suggest that in the absence of functional Trem2, early amyloidogenesis is accelerated due to reduced phagocytic clearance of amyloid seeds despite reduced plaque-associated ApoE.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Jucker, M. & Walker, L. C. Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 501, 45–51 (2013).
Meyer-Luehmann, M. et al. Exogenous induction of cerebral beta-amyloidogenesis is governed by agent and host. Science 313, 1781–1784 (2006).
Butovsky, O. & Weiner, H. L. Microglial signatures and their role in health and disease. Nat. Rev. Neurosci. 19, 622–635 (2018).
Song, W. M. & Colonna, M. The identity and function of microglia in neurodegeneration. Nat. Immunol. 19, 1048–1058 (2018).
Ulrich, J. D., Ulland, T. K., Colonna, M. & Holtzman, D. M. Elucidating the role of TREM2 in Alzheimer’s disease. Neuron 94, 237–248 (2017).
Guerreiro, R. et al. TREM2 variants in Alzheimer’s disease. N. Engl. J. Med. 368, 117–127 (2013).
Jonsson, T. et al. Variant of TREM2 associated with the risk of Alzheimer’s disease. N. Engl. J. Med. 368, 107–116 (2013).
Mazaheri, F. et al. TREM2 deficiency impairs chemotaxis and microglial responses to neuronal injury. EMBO Rep. 18, 1186–1198 (2017).
Ulland, T. K. et al. TREM2 maintains microglial metabolic fitness in Alzheimer’s disease. Cell 170, 649–663.e13 (2017).
Kleinberger, G. et al. The FTD-like syndrome causing TREM2 T66M mutation impairs microglia function, brain perfusion, and glucose metabolism. EMBO J. 36, 1837–1853 (2017).
Kleinberger, G. et al. TREM2 mutations implicated in neurodegeneration impair cell surface transport and phagocytosis. Sci. Transl. Med. 6, 243ra86 (2014).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290.e17 (2017).
Krasemann, S. et al. The TREM2-APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581.e9 (2017).
Liu, C. C. et al. ApoE4 accelerates early seeding of amyloid pathology. Neuron 96, 1024–1032.e3 (2017).
Huynh, T. V. et al. Age-dependent effects of apoE reduction using antisense oligonucleotides in a model of β-amyloidosis. Neuron 96, 1013–1023.e4 (2017).
Ma, J., Yee, A., Brewer, H. B. Jr, Das, S. & Potter, H. Amyloid-associated proteins α1-antichymotrypsin and apolipoprotein E promote assembly of Alzheimer β-protein into filaments. Nature 372, 92–94 (1994).
Liao, F. et al. Targeting of nonlipidated, aggregated apoE with antibodies inhibits amyloid accumulation. J. Clin. Invest. 128, 2144–2155 (2018).
Jay, T. R. et al. Disease progression-dependent effects of TREM2 deficiency in a mouse model of Alzheimer’s disease. J. Neurosci. 37, 637–647 (2017).
Wang, Y. et al. TREM2-mediated early microglial response limits diffusion and toxicity of amyloid plaques. J. Exp. Med. 213, 667–675 (2016).
Jay, T. R. et al. TREM2 deficiency eliminates TREM2+inflammatory macrophages and ameliorates pathology in Alzheimer’s disease mouse models. J. Exp. Med. 212, 287–295 (2015).
Wang, Y. et al. TREM2 lipid sensing sustains the microglial response in an Alzheimer’s disease model. Cell 160, 1061–1071 (2015).
Kane, M. D. et al. Evidence for seeding of β-amyloid by intracerebral infusion of Alzheimer brain extracts in β-amyloid precursor protein-transgenic mice. J. Neurosci. 20, 3606–3611 (2000).
Radde, R. et al. Abeta42-driven cerebral amyloidosis in transgenic mice reveals early and robust pathology. EMBO Rep. 7, 940–946 (2006).
Turnbull, I. R. et al. Cutting edge: TREM-2 attenuates macrophage activation. J. Immunol. 177, 3520–3524 (2006).
Guerreiro, R. J. et al. Using exome sequencing to reveal mutations in TREM2 presenting as a frontotemporal dementia-like syndrome without bone involvement. JAMA Neurol. 70, 78–84 (2013).
Ulrich, J. D. et al. Altered microglial response to Aβ plaques in APPPS1-21 mice heterozygous for TREM2. Mol. Neurodegener. 9, 20 (2014).
Brendel, M. et al. Increase of TREM2 during aging of an Alzheimer’s disease mouse model is paralleled by microglial activation and amyloidosis. Front. Aging Neurosci. 9, 8 (2017).
Dickens, A. M. et al. Detection of microglial activation in an acute model of neuroinflammation using PET and radiotracers 11C-(R)-PK11195 and 18F-GE-180. J. Nucl. Med. 55, 466–472 (2014).
Holtman, I. R. et al. Induction of a common microglia gene expression signature by aging and neurodegenerative conditions: a co-expression meta-analysis. Acta Neuropathol. Commun. 3, 31 (2015).
Grathwohl, S. A. et al. Formation and maintenance of Alzheimer’s disease β-amyloid plaques in the absence of microglia. Nat. Neurosci. 12, 1361–1363 (2009).
Heppner, F. L. et al. Experimental autoimmune encephalomyelitis repressed by microglial paralysis. Nat. Med. 11, 146–152 (2005).
Sims, R. et al. Rare coding variants in PLCG2, ABI3, and TREM2 implicate microglial-mediated innate immunity in Alzheimer’s disease. Nat. Genet. 49, 1373–1384 (2017).
Ulrich, J. D. et al. ApoE facilitates the microglial response to amyloid plaque pathology. J. Exp. Med. 215, 1047–1058 (2018).
Cheng-Hathaway, P. J. et al. The Trem2 R47H variant confers loss-of-function-like phenotypes in Alzheimer’s disease. Mol. Neurodegener. 13, 29 (2018).
Lavisse, S. et al. Reactive astrocytes overexpress TSPO and are detected by TSPO positron emission tomography imaging. J. Neurosci. 32, 10809–10818 (2012).
Chau, W. F. et al. Exploration of the impact of stereochemistry on the identification of the novel translocator protein PET imaging agent [18F]GE-180. Nucl. Med. Biol. 42, 711–719 (2015).
Feeney, C. et al. Kinetic analysis of the translocator protein positron emission tomography ligand [18F]GE-180 in the human brain. Eur. J. Nucl. Med. Mol. Imaging 43, 2201–2210 (2016).
Vomacka, L. et al. TSPO imaging using the novel PET ligand [18F]GE-180: quantification approaches in patients with multiple sclerosis. EJNMMI Res. 7, 89 (2017).
Boutin, H. et al. 18F-GE-180: a novel TSPO radiotracer compared to 11C-R-PK11195 in a preclinical model of stroke. Eur. J. Nucl. Med. Mol. Imaging 42, 503–511 (2015).
James, M. L. et al. [18F]GE-180 PET detects reduced microglia activation after LM11A-31 therapy in a mouse model of Alzheimer’s disease. Theranostics 7, 1422–1436 (2017).
Yeh, F. L., Wang, Y., Tom, I., Gonzalez, L. C. & Sheng, M. TREM2 binds to apolipoproteins, including APOE and CLU/APOJ, and thereby facilitates uptake of amyloid-beta by microglia. Neuron 91, 328–340 (2016).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016).
Fagan, A. M. et al. Human and murine ApoE markedly alters Aβ metabolism before and after plaque formation in a mouse model of Alzheimer’s disease. Neurobiol. Dis. 9, 305–318 (2002).
Sperling, R., Mormino, E. & Johnson, K. The evolution of preclinical Alzheimer’s disease: implications for prevention trials. Neuron 84, 608–622 (2014).
Ziegler-Waldkirch, S. et al. Seed-induced Aβ deposition is modulated by microglia under environmental enrichment in a mouse model of Alzheimer’s disease. EMBO J. 37, 167–182 (2018).
Song, W. M. et al. Humanized TREM2 mice reveal microglia-intrinsic and -extrinsic effects of R47H polymorphism. J. Exp. Med. 215, 745–760 (2018).
Suárez-Calvet, M. et al. Early changes in CSF sTREM2 in dominantly inherited Alzheimer’s disease occur after amyloid deposition and neuronal injury. Sci. Transl. Med. 8, 369ra178 (2016).
Sevigny, J. et al. The antibody aducanumab reduces Aβ plaques in Alzheimer’s disease. Nature 537, 50–56 (2016).
Styren, S. D., Hamilton, R. L., Styren, G. C. & Klunk, W. E. X-34, a fluorescent derivative of Congo red: a novel histochemical stain for Alzheimer’s disease pathology. J. Histochem. Cytochem. 48, 1223–1232 (2000).
Brendel, M. et al. Glial activation and glucose metabolism in a transgenic amyloid mouse model: a triple-tracer PET study. J. Nucl. Med. 57, 954–960 (2016).
Overhoff, F. et al. Automated spatial brain normalization and hindbrain white matter reference tissue give improved [18F]-florbetaben PET quantitation in Alzheimer’s model mice. Front. Neurosci. 10, 45 (2016).
Deussing, M. et al. Coupling between physiological TSPO expression in brain and myocardium allows stabilization of late-phase cerebral [18F]GE180 PET quantification. Neuroimage 165, 83–91 (2018).
Page, R. M. et al. Generation of Aβ38 and Aβ42 is independently and differentially affected by familial Alzheimer disease-associated presenilin mutations and γ-secretase modulation. J. Biol. Chem. 283, 677–683 (2008).
Daria, A. et al. Young microglia restore amyloid plaque clearance of aged microglia. EMBO J. 36, 583–603 (2017).
Wiśniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Bruderer, R., Bernhardt, O. M., Gandhi, T. & Reiter, L. High-precision iRT prediction in the targeted analysis of data-independent acquisition and its impact on identification and quantitation. Proteomics 16, 2246–2256 (2016).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (DFG) within the framework of the Munich Cluster for Systems Neurology (EXC 1010 SyNergy), a DFG funded Koselleck Project (HA1737/16-1 to C.H.), by the Helmholtz-Gemeinschaft Zukunftsthema ‘Immunology and Inflammation’ (ZT-0027 to C.H.), the FOR2290 (to S.F.L. and C.H.), and by a dedicated PET imaging grant to M.B. and A.R. (BR4580/1-1 & RO5194/1-1). This project has received additional funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement No. 115976. Startup funding for this project came from LMUexcellent. Additional funding came from the general legacy of S. Ammer, the MetLife award, and the Cure Alzheimer’s fund. M.M.-L. is supported by the Emmy Noether Program of the DFG (ME 3542/1-1). M.M.-L. and S.F.L. are supported by the Hans and Ilse Breuer Foundation. S.T. got support from Ono Pharmaceuticals, Japan. D.M.H. is supported by NIH grants NS090934 and AG047644, the JPB Foundation, and the Cure Alzheimer’s Fund. S.F.L. is supported by the Centers of Excellence in neurodegeneration and the Helmholtz-Israel program. O.B. is supported by NIH grants NINDS (R01NS088137), NIH-NIA (R01AG051812), NIH-NIA (R01AG054672) and the Cure Alzheimer’s Fund. The APPPS1 colony was established from a breeding pair kindly provided by M. Jucker (Hertie-Institute for Clinical Brain Research, University of Tübingen and DZNE-Tübingen). The authors thank M. Colonna for the Trem2–/– mice. Thanks to the Queen Square Brain Bank for access to tissue: this resource is funded in part by the Weston Foundation and the MRC. D.E. is supported by the European Community’s Health Seventh Framework Programme under grant agreement 617198 [DPR-MODELS].
Author information
Authors and Affiliations
Contributions
C.H., M.M.-L., G.K., and S.P. conceived the study and analyzed the results. C.H. wrote the manuscript with help from M.M.-L., S.P., T.A., G.K., M.B., and D.M.H. and further input from all co-authors. S.P. and N.K. performed seeding experiments; S.P. and M.X. performed the ApoE stainings. B.N. performed Aβ ELISA. M.B., M.D., C.F., P.B., and A.R. performed PET imaging and quantitative PET analyses. G.E. made GE180 cassettes available through an early access model. A.E., A.G. and S.P. performed the three-dimensional image analyses. O.B. and S.K. provided independent immunohistochemical data and interpretation on ApoE. D.M.H. interpreted the ApoE stainings and provided appropriate ApoE antibodies. T.A. and J.H. provided human brain sections and interpreted the immunohistochemical analyses. G.K. and N.P. performed TREM2 sequencing and D.E. performed PSEN1, PSEN2 and APP sequencing of human autopsy cases. S.T., A.C., L.S.M., S.A.M., and S.F.L. prepared primary microglial lysates and measured ApoE levels by mass spectrometry. M.W. provided technical advice for protein extraction and ApoE experiments. S.A.G. and J.J.N. provided brain sections from microglia-depleted mice.
Corresponding authors
Ethics declarations
Competing interests
C.H. collaborates with Denali and received a speaker honorarium of Roche and Novartis. D.M.H. co-founded and is on the scientific advisory board of C2N Diagnostics. D.M.H. consults for Denali, Eli Lilly, Glaxosmithkline, and AbbVie. A.R. received consultant and speaker honoria from Piramal Imaging and GE Healthcare. O.B. collaborates with Sanofi. S.T. collaborates with Ono Pharmaceuticals (Japan). S.A.G is an employee of Neurimmune AG. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplemetary Figure 1 Amyloid seeding occurs in a host-dependent manner.
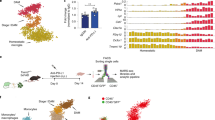
a, Top: ThioS and 4G8 co-staining for amyloid plaques in C57BL6 mice injected with APPPS1 brain homogenate (BH) (n = 7 mice) show no plaques in the hippocampus or cortex ten weeks after injection. Middle: In contrast, amyloid seeding is induced in the hippocampus of APPPS1/Trem2+/+ mice at four months (n = 8 mice). Bottom: Age-matched C57BL6 brain homogenate-injected APPPS1 mice did not induce amyloid seeding in the hippocampus, although cortical plaques are present in the transgenic APPPS1 mice (n = 6 mice). b, Quantification of Aβ42 in formic acid fractions of C57BL6 brain homogenate-injected hippocampi by Meso Scale Discovery electrochemiluminescence assay confirm significantly lower plaque load compared to APPPS1 brain homogenate-injected hippocampi in APPPS1/Trem2+/+ mice at four months. Data represent mean ± s.e.m. (APPPS1 BH-injected C57BL6 n = 6 mice, C57BL6 BH-injected APPPS1/Trem2+/+ n = 6 mice; APPPS1-injected APPPS1/Trem2+/+ n = 8 mice; F2,17 = 36.8, P = 1.1 × 10–5). One-way ANOVA, Dunnett’s post hoc analysis; ***P < 0.0001. c, APPPS1 brain homogenate-injected APPPS1/Trem2+/+, APPPS1/Trem2-/- and APPPS1/Trem2p.T66M mice show no hippocampal or cortical plaque staining two weeks after injection (n = 4 mice/genotype). d, Uninjected APPPS1/Trem2+/+ mice show neither ThioS nor immunopositive amyloid plaques in hippocampus compared to cortex. Similar seeding patterns are not observed in the cortex because amyloid seeding is region-dependent2 but note pre-existing non-experimentally seeded plaques within the cortex of four months old transgenic mice. Similarly, uninjected APPPS1/Trem2-/- and APPPS1/Trem2p.T66M mice show no ThioS or 4G8-positive plaques in the hippocampus compared to cortex (n = 6 mice/genotype).
Supplemetary Figure 2 Trem2 deficiency impairs microglial clustering and CD68 upregulation around cortical plaques.
a, APPPS1/Trem2+/+ mice show increased microglia clustering around cortical plaques as compared to APPPS1/Trem2-/- and APPPS1/Trem2p.T66M mice. b, Number of IBA1-positive microglia cells quantified per plaque in the cortex (n+/+ = 9 mice, n-/- = 10 mice, np.T66M = 10 mice, F2,26 = 83.07, P = 5.1 × 10–12). c, APPPS1/Trem2+/+ mice show increased CD68-positive microglia clustering around cortical plaques as compared to APPPS1/Trem2-/- and APPPS1/Trem2p.T66M mice. d, Number of CD68-positive microglia cells quantified per plaque in the cortex (n+/+ = 8 mice, n-/- = 6 mice, np.T66M = 6 mice; F2,17 = 109.7, P = 1.9 × 10–10). e, Number of IBA1-positive microglia cells quantified in plaque barren thalamic nuclei (n = 6 mice/genotype; F2,15 = 0.1553, P = 0.8576). Data represent mean ± s.e.m.. One-way ANOVA, Dunnett’s post hoc analysis; n.s. P > 0.05; ***p < 0.0001.
Supplemetary Figure 3 Regression correlation analysis with longitudinal imaging of microglial activity as well as fibrillar amyloidogenesis in individual mice.
a,b, Coronal and axial slices show serial TSPO (a) and Amyloid μPET (b) Z-score increases (versus C57BL6) from three to twelve months of age for APPPS1/Trem2+/+ and APPPS1/Trem2-/- mice. APPPS1/Trem2-/- mice indicate decreased microglial activity, most pronounced at twelve months of age, and higher fibrillar amyloidogenesis at three and six months of age when compared to APPPS1/Trem2+/+ mice. c,d, Spaghetti plots show individual longitudinal time courses of cortical microglial activity (c) and fibrillar amyloidogenesis (d) as assessed by in vivo μPET in APPPS1/Trem2+/+ (purple highlighted data point corresponds to representative image shown in e and f) and APPPS1/Trem2-/- mice (dark blue highlighted data point corresponds to representative image shown in a and b). e,f, x34/CD68/IBA1 co-stained microglia (n+/+ = 8 mice; n-/- = 7 mice) as well as (e) fibrillar and immunostained amyloid plaques (f) (n+/+ = 8 mice; n-/- = 7 mice) of twelve months old APPPS1 mice previously included in longitudinal TSPO and Amyloid μPET imaging. g,h, Regression correlation analysis of CD68-positive phagocytic microglia staining and TSPO μPET imaging (n+/+ = 5 mice, n-/- = 4 mice, P = 0.0026) (g) and x34-positive fibrillar Aβ staining and amyloid μPET imaging (n+/+ = 4 mice, n-/- = 4 mice, P = 0.0003) (h)in three months old APPPS1 mice. i,j, Regression correlation analysis of CD68 and TSPO μPET imaging (n+/+ = 10 mice, n-/- = 5 mice, p = 5.2E-5) (i)and x34 and Amyloid μPET imaging (n+/+ = 4 mice, n-/- = 4 mice, P = 0.0088) (j)in six months old APPPS1 mice. k,l Regression correlation analysis of CD68 and TSPO μPET imaging (n+/+ = 8 mice, n-/- = 7 mice, P = 1.0 x 10-6) (k) and x34 and Amyloid μPET imaging (n+/+ = 7 mice, n-/- = 7 mice, P = 2.0 × 10–5) (l) in twelve months old APPPS1 mice. Purple dots represent APPPS1/Trem2+/+ and blue dots indicate APPPS1/Trem2-/- mice.
Supplemetary Figure 4 Confocal images of individual microglia-depleted APPPS1/TK+ and control APPPS1/TK– mice.
a, IMARIS 3D reconstructed high-resolution confocal images of x34/ApoE/Aβ stained cortical amyloid plaques of control APPPS1/TK- (left) and age-matched microglia-depleted APPPS1/TK + mice (right) (n = 3 mice/genotype). Numbers at the top left of each image indicate three individual mice per group. b, Left: 3D reconstructed image of x34/ApoE/IBA1 staining shows higher ApoE and IBA1 colocalisation in APPPS1/TK- mice indicated by the degree of white merged staining, compared to; right: APPPS1/TK + mice (n = 3 mice/genotype). c, x34/ApoE/GFAP co-staining in APPPS1/TK- mice looks comparable to APPPS1/TK + mice (n = 3 mice/genotype).
Supplemetary Figure 5 Reduced ApoE levels in Aβ plaques and impaired microglial clustering in TREM2 mutation carriers.
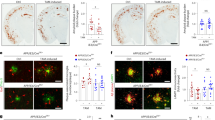
a, Temporal neocortex of additional human AD patients with and without the indicated examples of TREM2 variants stained for Aβ by 4G8 immunohistochemistry. b, Reduced ApoE immunoreactivity within amyloid plaques compared to no CV shown in sections consecutive to those of the left column. Red boxes indicate the area in each staining that is magnified as inset. c, Additional examples of ApoE stainings comparing the same region in consecutive Aβ stained sections. d, Impaired clustering of microglial cells (brown) around and within amyloid plaques (red) in AD cases with different TREM2 coding variants compared to no CV cases (last 2 rows). Dotted black boxes indicate the area magnified as inset. e, Quantification of Aβ42 in formic acid fractions of temporal neocortex by Meso Scale Discovery electrochemiluminescence assay (nR47H = 2 cases, nR62H = 4 cases; nR62C = 1 case; nno CV = 4 cases). f, ApoE/Aβ42 ratio quantified from temporal neocortex plaque-enriched formic acid fraction (nR47H = 2 cases, nR62H = 4 cases; nR62C = 1 case, nno CV = 4 cases). Of note, frozen material from p.D87N cases was not available and therefore not included. One of the no CV cases could not be included in the study due to diagnosed Hepatitis. (g) Number of IBA1-positive microglia per plaque in temporal neocortex quantified from images shown in d and Fig. 8 (nR47H = 2 cases, nR62H = 4 cases; nR62C = 1 case; nD87N = 2 cases; nno CV = 3 cases). Noteworthy, sections from only three no CV cases were available in comparison to frozen material. No subjects were excluded in this analysis. (h) Aβ42 in formic acid fractions quantified from temporal neocortex grouped according to APOE status (nE3/E3 = 4 cases, nE3/E4 = 4 cases; nE4/E4 = 3 cases). (i) Number of IBA1-positive microglia per plaque in temporal neocortex grouped according to APOE status (nE3/E3 = 3 cases, nE3/E4 = 6 cases; nE4/E4 = 3 cases). Medial temporal cortex at the level of anterior hippocampus was used for all experiments. Data represent as mean ± s.d.
Supplemetary Figure 6 Schematic summary.
Schematic figure of the consequences of a Trem2 loss-of-function (LoF) on microglial clustering and ApoE accumulation in amyloid plaques.
Supplementary information
Rights and permissions
About this article
Cite this article
Parhizkar, S., Arzberger, T., Brendel, M. et al. Loss of TREM2 function increases amyloid seeding but reduces plaque-associated ApoE. Nat Neurosci 22, 191–204 (2019). https://doi.org/10.1038/s41593-018-0296-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0296-9
This article is cited by
-
Regulation of human microglial gene expression and function via RNAase-H active antisense oligonucleotides in vivo in Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
Taurine reduces microglia activation in the brain of aged senescence-accelerated mice by increasing the level of TREM2
Scientific Reports (2024)
-
Immunological aspects of central neurodegeneration
Cell Discovery (2024)
-
Associations between sex, body mass index and the individual microglial response in Alzheimer’s disease
Journal of Neuroinflammation (2024)
-
Microglial function, INPP5D/SHIP1 signaling, and NLRP3 inflammasome activation: implications for Alzheimer’s disease
Molecular Neurodegeneration (2023)