Abstract
Amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) are associated with loss of nuclear transactive response DNA-binding protein 43 (TDP-43). Here we identify that TDP-43 regulates expression of the neuronal growth-associated factor stathmin-2. Lowered TDP-43 levels, which reduce its binding to sites within the first intron of stathmin-2 pre-messenger RNA, uncover a cryptic polyadenylation site whose utilization produces a truncated, non-functional mRNA. Reduced stathmin-2 expression is found in neurons trans-differentiated from patient fibroblasts expressing an ALS-causing TDP-43 mutation, in motor cortex and spinal motor neurons from patients with sporadic ALS and familial ALS with GGGGCC repeat expansion in the C9orf72 gene, and in induced pluripotent stem cell (iPSC)-derived motor neurons depleted of TDP-43. Remarkably, while reduction in TDP-43 is shown to inhibit axonal regeneration of iPSC-derived motor neurons, rescue of stathmin-2 expression restores axonal regenerative capacity. Thus, premature polyadenylation-mediated reduction in stathmin-2 is a hallmark of ALS–FTD that functionally links reduced nuclear TDP-43 function to enhanced neuronal vulnerability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others

Data availability
All RNA sequencing data generated and analyzed for this study have been deposited in the Gene Expression Omnibus database under accession number GSE122069. The data that support the findings of this study are readily available from the corresponding authors upon reasonable request.
References
Taylor, J. P., Brown, R. H. Jr & Cleveland, D. W. Decoding ALS: from genes to mechanism. Nature 539, 197–206 (2016).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Ling, S. C., Polymenidou, M. & Cleveland, D. W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 79, 416–438 (2013).
Lagier-Tourenne, C., Polymenidou, M. & Cleveland, D. W. TDP-43 and FUS/TLS: emerging roles in RNA processing and neurodegeneration. Hum. Mol. Genet. 19, R46–R64 (2010).
Alami, N. H. et al. Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron 81, 536–543 (2014).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Ling, J. P., Pletnikova, O., Troncoso, J. C. & Wong, P. C. TDP-43 repression of nonconserved cryptic exons is compromised in ALS-FTD. Science 349, 650–655 (2015).
Ayala, Y. M. et al. TDP-43 regulates its mRNA levels through a negative feedback loop. EMBO J. 30, 277–288 (2011).
Lagier-Tourenne, C. et al. Divergent roles of ALS-linked proteins FUS/TLS and TDP-43 intersect in processing long pre-mRNAs. Nat. Neurosci. 15, 1488–1497 (2012).
Kapitein, L. C. & Hoogenraad, C. C. Building the neuronal microtubule cytoskeleton. Neuron 87, 492–506 (2015).
Mitchison, T. & Kirschner, M. Dynamic instability of microtubule growth. Nature 312, 237–242 (1984).
Desai, A. & Mitchison, T. J. Microtubule polymerization dynamics. Annu. Rev. Cell. Dev. Biol. 13, 83–117 (1997).
Stein, R., Mori, N., Matthews, K., Lo, L. C. & Anderson, D. J. The NGF-inducible SCG10 mRNA encodes a novel membrane-bound protein present in growth cones and abundant in developing neurons. Neuron 1, 463–476 (1988).
Morii, H., Shiraishi-Yamaguchi, Y. & Mori, N. SCG10, a microtubule destabilizing factor, stimulates the neurite outgrowth by modulating microtubule dynamics in rat hippocampal primary cultured neurons. J. Neurobiol. 66, 1101–1114 (2006).
Riederer, B. M. et al. Regulation of microtubule dynamics by the neuronal growth-associated protein SCG10. Proc. Natl. Acad. Sci. USA 94, 741–745 (1997).
Belmont, L. D. & Mitchison, T. J. Identification of a protein that interacts with tubulin dimers and increases the catastrophe rate of microtubules. Cell 84, 623–631 (1996).
Chauvin, S. & Sobel, A. Neuronal stathmins: a family of phosphoproteins cooperating for neuronal development, plasticity and regeneration. Prog. Neurobiol. 126, 1–18 (2015).
Shin, J. E. et al. SCG10 is a JNK target in the axonal degeneration pathway. Proc. Natl. Acad. Sci. USA 109, E3696–E3705 (2012).
Shin, J. E., Geisler, S. & DiAntonio, A. Dynamic regulation of SCG10 in regenerating axons after injury. Exp. Neurol. 252, 1–11 (2014).
Graf, E. R., Heerssen, H. M., Wright, C. M., Davis, G. W. & DiAntonio, A. Stathmin is required for stability of the Drosophila neuromuscular junction. J. Neurosci. 31, 15026–15034 (2011).
Yusuf, M., Leung, K., Morris, K. J. & Volpi, E. V. Comprehensive cytogenomic profile of the in vitro neuronal model SH-SY5Y. Neurogenetics 14, 63–70 (2013).
Arnold, E. S. et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc. Natl. Acad. Sci. USA 110, E736–E745 (2013).
White, M. A. et al. TDP-43 gains function due to perturbed autoregulation in a Tardbp knock-in mouse model of ALS-FTD. Nat. Neurosci. 21, 552–563 (2018).
Fratta, P. et al. Mice with endogenous TDP-43 mutations exhibit gain of splicing function and characteristics of amyotrophic lateral sclerosis. EMBO J. 37, e98684 (2018).
Xue, Y. et al. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 152, 82–96 (2013).
Tian, B. & Graber, J. H. Signals for pre-mRNA cleavage and polyadenylation. Wiley Interdiscip. Rev. RNA 3, 385–396 (2012).
Sun, S. et al. Translational profiling identifies a cascade of damage initiated in motor neurons and spreading to glia in mutant SOD1-mediated ALS. Proc. Natl. Acad. Sci. USA 112, E6993–E7002 (2015).
Krach, F. et al. Transcriptome-pathology correlation identifies interplay between TDP-43 and the expression of its kinase CK1E in sporadic ALS. Acta Neuropathol. 136, 405–423 (2018).
Saberi, S., Stauffer, J. E., Schulte, D. J. & Ravits, J. Neuropathology of amyotrophic lateral sclerosis and its variants. Neurol. Clin. 33, 855–876 (2015).
Mackenzie, I. R. et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann. Neurol. 61, 427–434 (2007).
Sun, M. et al. Cryptic exon incorporation occurs in Alzheimer’s brain lacking TDP-43 inclusion but exhibiting nuclear clearance of TDP-43. Acta Neuropathol. 133, 923–931 (2017).
Jeong, Y. H. et al. Tdp-43 cryptic exons are highly variable between cell types. Mol. Neurodegener. 12, 13 (2017).
Humphrey, J., Emmett, W., Fratta, P., Isaacs, A. M. & Plagnol, V. Quantitative analysis of cryptic splicing associated with TDP-43 depletion. BMC Med. Genomics 10, 38 (2017).
Naftelberg, S., Schor, I. E., Ast, G. & Kornblihtt, A. R. Regulation of alternative splicing through coupling with transcription and chromatin structure. Annu. Rev. Biochem. 84, 165–198 (2015).
Licatalosi, D. D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Elkon, R., Ugalde, A. P. & Agami, R. Alternative cleavage and polyadenylation: extent, regulation and function. Nat. Rev. Genet. 14, 496–506 (2013).
Tian, B. & Manley, J. L. Alternative polyadenylation of mRNA precursors. Nat. Rev. Mol. Cell Biol. 18, 18–30 (2017).
Masuda, A. et al. Position-specific binding of FUS to nascent RNA regulates mRNA length. Genes Dev. 29, 1045–1057 (2015).
Prudencio, M. et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat. Neurosci. 18, 1175–1182 (2015).
Rot, G. et al. High-resolution RNA maps suggest common principles of splicing and polyadenylation regulation by TDP-43. Cell Rep. 19, 1056–1067 (2017).
Brunden, K. R. et al. Altered microtubule dynamics in neurodegenerative disease: Therapeutic potential of microtubule-stabilizing drugs. Neurobiol. Dis. 105, 328–335 (2017).
Wen, H. L. et al. Stathmin, a microtubule-destabilizing protein, is dysregulated in spinal muscular atrophy. Hum. Mol. Genet. 19, 1766–1778 (2010).
Strey, C. W. et al. Dysregulation of stathmin, a microtubule-destabilizing protein, and up-regulation of Hsp25, Hsp27, and the antioxidant peroxiredoxin 6 in a mouse model of familial amyotrophic lateral sclerosis. Am. J. Pathol. 165, 1701–1718 (2004).
Bellouze, S. et al. Stathmin 1/2-triggered microtubule loss mediates Golgi fragmentation in mutant SOD1 motor neurons. Mol. Neurodegener. 11, 43 (2016).
Mason, M. R., Lieberman, A. R., Grenningloh, G. & Anderson, P. N. Transcriptional upregulation of SCG10 and CAP-23 is correlated with regeneration of the axons of peripheral and central neurons in vivo. Mol. Cell. Neurosci. 20, 595–615 (2002).
Selvaraj, B. T., Frank, N., Bender, F. L., Asan, E. & Sendtner, M. Local axonal function of STAT3 rescues axon degeneration in the pmn model of motoneuron disease. J. Cell. Biol. 199, 437–451 (2012).
Duncan, J. E., Lytle, N. K., Zuniga, A. & Goldstein, L. S. The microtubule regulatory protein stathmin is required to maintain the integrity of axonal microtubules in Drosophila. PLoS One 8, e68324 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Xue, Y. et al. Sequential regulatory loops as key gatekeepers for neuronal reprogramming in human cells. Nat. Neurosci. 19, 807–815 (2016).
Minteer, S.D. & Moore, C.M. Microfluidic ethanol biobatteries on a microchip. Methods Mol. Biol. 321, 157–166 (2006).
Taylor, A. M. et al. A microfluidic culture platform for CNS axonal injury, regeneration and transport. Nat. Methods 2, 599–605 (2005).
Brooks, B.R., Miller, R.G., Swash, M. & Munsat, T.L. El Escorial revisited: revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph Lateral Scler Other Motor Neuron Disord. 1, 293–299 (2000).
Acknowledgements
The plasmids for expression of Brn2 and shPTB were kind gifts from X. D. Fu, UCSD. We thank the viral vector core facility at Sanford Burnham Prebys (SBP) Medical Discovery Institute for the kind contribution of the SIN18 vector. We thank B. Ren for providing the Illumina sequencing platform. We thank A. Goginashvili for valued input on the manuscript. We thank J. Nuovo at Phylogeny Inc. (Columbus, OH) for performing the chromogenic in situ hybridization experiment. This work was supported by grants from NINDS/NIH R01-NS27036 to D.W.C, Target ALS (S20A00) and NINDS/NIH (R01-NS087227) to C.L.-T. and R01-NS088578 to J.R. C.L.-T. was recipient of a Career Development grant from the Muscular Dystrophy Association (MDA). Z.M was recipient of EMBO long-term fellowship and is currently supported by the Human Frontiers Science Program (HFSP) long-term fellowship. J.L.-E. was recipient of a Milton Safenowitz postdoctoral fellowship. M.W.B. and M.S.B. were supported by the National Institute of General Medical Sciences of the National Institutes of Health Award T32-GM008666. M.A.M, C.F.B., and F.R. are employees and D.W.C. is a consultant for Ionis Pharmaceuticals.
Author information
Authors and Affiliations
Contributions
Z.M., J.L.-E., S.D.C, C.L.-T., and D.W.C. designed the research; Z.M., J.L.-E., M.W.B., K.D, O.Z., Y.S., S.D.C., C.L.-T., and D.W.C. analyzed the data; Z.M., J.L.-E., M.W.B., K.D., J.A., F.F., M.A.M., M.S.B., T.O, M.R., D.W., and N.L. performed the research; F.R., C.F.B., J.R., D.W., and N.L contributed key reagents and methodology; Z.M., C.L.-T., and D.W.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Expression of stathmin-2 protein is decreased more than 8 fold upon TDP-43 depletion in SH-SY5Y cells.
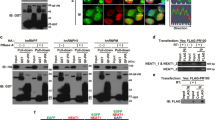
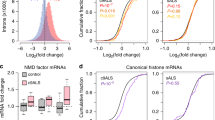
(a) Bar plot representing transcripts per million (TPM) values of the 15 most significant differentially expressed genes after TDP-43 depletion by siRNA treatment in SH-SY5Y cells. Mean values (−/+ STDEV) from 3 biologically independent experiments are plotted, DESeq2. (b) Serial dilution of total protein extracted from SH-SY5Y cells treated with siControl. Immunoblotting determines the expression of stathmin-2 in the sample diluted 1:8 as comparable to the non-diluted siTDP-43 sample. Experiment was repeated independently twice with similar results. For uncropped blots, see Supplementary Fig. 11.
Supplementary Figure 2 Genome-editing to produce SH-SY5Y cells expressing TDP-43N352S from both endogenous alleles.
(a) Histogram representation of the AAT codon (Asparagine) in position 352 of wild-type SH-SY5Y cells and its replacement by CRISPR-Cas9 to an AGT codon (Serine) in both endogenous TDP-43 alleles, as determined by sequencing in 3 independent biological samples (b) Nuclear to cytoplasmic TDP-43 ratios in SH-SY5YWT/WT (n = 34 cells, mean = 4.13) and SH-SY5YN352S/N352S (n = 31 cells, mean = 3.82) lines are plotted by using fluorescence intensity quantification. 3 biologically independent experiments were quantified. Statistical analysis was done using two-tailed t-test, P = 0.1427, SEM. (c) Alternative splicing alterations linked to TDP-43 loss of function 9 were tested in SH-SY5YWT/WT (black bars) and SH-SY5YN352S/N352S (red bars) lines. Representative images from 2% acrylamide gels are shown and inclusion/skipping band intensity ratios were plotted to the right (n = 5 independent biological experiments, two-tailed t-test, SEM). Mean values are represented for RWDD1 (2.08, P = 0.0002), CACNA1C (1.64, P = 0.00001), and ZNF569 (1.93, P = 0.003). **P < 0.01, ***P < 0.001. For uncropped gel images, see Supplementary Fig. 11. (d) Immunoblotting of stathmin-2 in SH-SY5YWT/WT and SH-SY5YN352S/N352S lines. α-Tubulin served as a loading control. Experiment was reproduced 3 times independently with similar results. For uncropped blot, see Supplementary Fig. 11.
Supplementary Figure 3 Stathmin-2 expression is reduced in fibroblasts of familial ALS patients and asymptomatic carriers of TDP-43 mutation.
(a) Family pedigree of ALS patients. Fibroblasts samples of four healthy controls and four carriers of the TDP-43 N352S mutation were collected for trans-differentiation into iNeurons. Two carriers of the mutation (hashed diamonds) were asymptomatic at the time of skin biopsy. (b) Quantitative real-time PCR analysis of stathmin-2 mRNA levels in individual controls (n = 4, white bar, mean = 1) and familial ALS patients’ fibroblasts (n = 7, turquoise bar, mean = 0.52). Red dots represent four family members carriers of the TDP-43N352S (from panel A) and three familial ALS patients heterozygously expressing one of the following mutations: TDP-43G298S, TDP-43A382T or TDP-43N390S. Each data point represents 2 independent biological experiments. P = 0.03, one-tailed t-test, SEM. *P < 0.05 (c) Representative images of control and ALS-patient fibroblasts lines. Immunofluorescence of TDP-43 (green) shows nuclear localization in fibroblasts from a control individual and an ALS patient carrying the TDP-43N352S mutation. Experiment was repeated independently with 3 control individuals and 3 familial ALS donor lines (with mutant TDP-43) with similar results. (d) Nuclear to cytoplasmic ratio of TDP-43 fluorescence intensity in control (n = 44 cells, mean = 8.45) and ALS patient fibroblasts (n = 47 cells, mean = 8.51) is shown. 4 biologically independent experiments were quantified, two-tailed t-test, P = 0.829 (e) Nuclear to cytoplasmic ratio of TDP-43 fluorescence intensity in control (n = 47 cells, mean = 3.76) and ALS patient (n = 47 cells, mean = 2.24) induced neurons (from Fig. 2a) is shown. 4 biologically independent experiments were quantified, two-tailed t-test, ***P < 0.001, SEM.
Supplementary Figure 4 Incorporation of exon 2a in stathmin-2 mRNA is not followed by splicing of downstream exons.
(a) Representation of possible stathmin-2 splice variants resulted from loss of TDP-43. (b) Schematics of possible stathmin-2 RNAs with exons 1 and 2. (c) RT-PCR demonstrating reduced stathmin-2 mRNA after TDP-43 depletion without expression of an isoform containing exon 2a spliced to exon 2, diagram is shown to the right. Experiment was repeated independently 3 times with similar results. Unprocessed gel image is shown in Supplementary Fig. 11. (d) Conservation scheme of the polyadenylation signal embedded within stathmin-2 exon 2a. Thirteen primate species, mouse and rat genomic regions are represented. (e) qPCR analysis of stathmin-2 transcripts demonstrates reduced full-length stathmin-2 pre-mRNA in SH-SY5Y cells expressing mutant TDP-43 in comparison to wild type SH-SY5Y cells. Mean values of 2 independent biological experiments are plotted. Error bars represent SD. Expression of the nuclear non-coding RNA Xist was used as endogenous control.
Supplementary Figure 5 Stathmin-2 mRNA is enriched in mice and human motor neurons.
(a) Data analyzed from Sun et al. 2015 30 showing enrichment of translated stathmin-2 mRNAs, determined as fragments per kilobase of transcript per million mapped reads (FPKM), in motor neurons relative to astrocytes (mean fold change = 25) and oligodendrocytes (mean fold change = 15) in wild-type mice (black bars) (two-tailed t-test, ****p < 0.0001, SEM). No change in stathmin-2 mRNA is observed in mice carrying the ALS-causing mutation G37R in the SOD1 gene (gray bars). (b) Bar plot representation of stathmin-2 mRNA expression rank (top 20 genes), based on RNA-seq 31 of laser-captured spinal motor neurons from 7 control individuals. (c) Expression of 4 stathmin genes from microarray analysis of human laser-captured spinal motor neurons (Rabin, S.J. et al. Hum. Mol. Genet. 2010), fetal spinal motor neurons and iPSC-derived motor neurons (Ho, R. et al. Nat. Neurosci. 2016) is plotted. Stathmin-2 mRNA (red) is enriched in human spinal motor neurons relative to mRNAs of other stathmin genes.
Supplementary Figure 6 Altered processing and usage of premature polyadenylation to produce short stathmin-2 RNA is a hallmark of sporadic ALS.
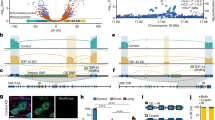
RNA sequencing of control and sporadic ALS laser-captured spinal motor neurons reveals robust signature of exon 2a incorporation in stathmin-2 mRNA in 9 sporadic ALS samples but none of the 3 healthy controls. Lower diagram shows the genomic region of stathmin-2. Data was analyzed from Krach et al., 2018 31.
Supplementary Figure 7 Expression of short stathmin-2 RNA is a hallmark of affected spinal motor neurons and motor cortex of sporadic ALS patients.
(a) RT-PCR using primers targeted to exon 1 and exon 2a confirms expression of stathmin-2 truncated RNA in anterior horns of thoracic spinal cord from sporadic ALS patients (n = 7), but not in control spinal cords (n = 5). (b) RT-PCR using primers targeted to exon 1 and exon 2a confirms expression of stathmin-2 truncated RNA in frontal cortex from sporadic and C9orf72 ALS patients diagnosed with frontotemporal dementia, but not in control. For uncropped gel images related to a and b panels, see Supplementary Fig. 11. (c) Motor cortex and (d) spinal cord sections isolated from healthy individuals and sporadic ALS patients were hybridized with LNA probes targeting intron one of stathmin-2 pre-mRNA (for truncated RNA) or exon 5 of stathmin-2 pre-mRNA. Signal is shown in blue, counterstain is nuclear fast red. Experiment was replicated with similar results in n = 5 ALS patients and n = 5 control individuals with a total of: 7 replicate experiments in motor cortex, 6 replicates in cervical spinal cord, 4 replicates in thoracic spinal cord, and 5 replicates in lumbar spinal cord. Each replicate tissue slide contained serial sections probed for the short and long RNA targets and each contemporaneous hybridization and staining batch included slides hybridized with control probes.
Supplementary Figure 8 Abnormal TDP-43 pathology is present in sporadic ALS and c9ALS patient spinal motor neurons, but not in SOD1-ALS.
Alpha motor neurons of the thoracic spinal cord stained by immunohistochemistry (IHC) for normal TDP-43 (using an antibody raised against the N-terminal amino acids 1-260, top row) and phosphorylated TDP-43 (pS409-410, bottom row). Hallmarks of TDP-43 pathology including nuclear clearance, phosphorylation and cytoplasmic punctate and skein-like inclusions are visible in the sporadic ALS and c9ALS/FTD motor neurons, but not in SOD1-ALS or control patients. Scale bar: 25 μm. Experiment was replicated with similar results in 3 ALS patients and 3 control individuals including tissue sections from lumbar spinal cord and motor cortex.
Supplementary Figure 9 Generation and characterization of induced pluripotent cells and motor neurons.
(a) Cytogenetic analysis of selected clone confirmed normal karyotype by G-banding staining. Representative image of 20 examined cells is shown. (b–d) Characterization of selected iPSC clone by immunofluorescence. Pluripotency was confirmed by staining for the following stem cells markers: (b) Nanog and Sox2 (c) Oct4 and SSEA4 (d) TRA-1-81 and SSEA1. Experiment was repeated 3 times independently with similar results. (e) Schematic timeline of iPS cells differentiation into motor neurons. (f) Immunofluorescence staining of motor neurons differentiated from human iPS cells demonstrates expression of the motor neurons precursor marker homeobox gene HB9, at day 21 of differentiation. (g) Bright field image of iPSC-derived motor neuron cultures. (h) Immunofluorescence staining demonstrates motor neurons maturation by expression of the axonal neurofilament heavy subunit (NF-H; green) and the microtubule-associated protein 2 (MAP2; red), at day 28 of differentiation. Experiments in f-h were reproduced 3 times independently with similar results. (i) Cell viability assay was performed in iPSC-derived motor neurons treated with control ASOs (mean = 78%) or with ASOs targeting TDP-43 (mean = 71.5%) or stathmin-2 (mean = 73.5%) for 20 days. Percentages of viable cells are plotted, 4 biologically independent motor neuron cultures are shown, error bars represent SEM.
Supplementary Figure 10
Scans of immunoblots and RT-PCR gels presented in the current study.
Supplementary Figure 11
Scans of immunoblots and RT-PCR gels presented in the current study.
Supplementary information
Supplementary Table 1
Differentially expressed genes in SH-SY5Y cells after TDP-43 knockdown by siRNA.
Supplementary Table 2
Differentially expressed genes in isogenic SH-SY5Y cells homozygously expressing mutant TDP-43N352S.
Supplementary Table 3
Information on fibroblast lines.
Supplementary Table 4
List of primers.
Supplementary Table 5
Summary of human samples.
Rights and permissions
About this article
Cite this article
Melamed, Z., López-Erauskin, J., Baughn, M.W. et al. Premature polyadenylation-mediated loss of stathmin-2 is a hallmark of TDP-43-dependent neurodegeneration. Nat Neurosci 22, 180–190 (2019). https://doi.org/10.1038/s41593-018-0293-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0293-z
This article is cited by
-
Nuclear-import receptors as gatekeepers of pathological phase transitions in ALS/FTD
Molecular Neurodegeneration (2024)
-
Stathmin 2 is a potential treatment target for TDP-43 proteinopathy in amyotrophic lateral sclerosis
Translational Neurodegeneration (2024)
-
Fluid biomarkers for amyotrophic lateral sclerosis: a review
Molecular Neurodegeneration (2024)
-
A fluid biomarker reveals loss of TDP-43 splicing repression in presymptomatic ALS–FTD
Nature Medicine (2024)
-
The nuclear import receptor Kapβ2 modifies neurotoxicity mediated by poly(GR) in C9orf72-linked ALS/FTD
Communications Biology (2024)