Abstract
In addition to cognitive impairments, neurodevelopmental disorders often result in sensory processing deficits. However, the biological mechanisms that underlie impaired sensory processing associated with neurodevelopmental disorders are generally understudied and poorly understood. We found that SYNGAP1 haploinsufficiency in humans, which causes a sporadic neurodevelopmental disorder defined by cognitive impairment, autistic features, and epilepsy, also leads to deficits in tactile-related sensory processing. In vivo neurophysiological analysis in Syngap1 mouse models revealed that upper-lamina neurons in somatosensory cortex weakly encode information related to touch. This was caused by reduced synaptic connectivity and impaired intrinsic excitability within upper-lamina somatosensory cortex neurons. These results were unexpected, given that Syngap1 heterozygosity is known to cause circuit hyperexcitability in brain areas more directly linked to cognitive functions. Thus, Syngap1 heterozygosity causes a range of circuit-specific pathologies, including reduced activity within cortical neurons required for touch processing, which may contribute to sensory phenotypes observed in patients.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Sahin, M. & Sur, M. Genes, circuits, and precision therapies for autism and related neurodevelopmental disorders. Science 350, aab3897 (2015).
Hull, J. V. et al. Resting-state functional connectivity in autism spectrum disorders: a review. Front. Psychiatry 7, 205 (2017).
Just, M. A., Cherkassky, V. L., Keller, T. A. & Minshew, N. J. Cortical activation and synchronization during sentence comprehension in high-functioning autism: evidence of underconnectivity. Brain 127, 1811–1821 (2004).
Zoghbi, H. Y. & Bear, M. F. Synaptic dysfunction in neurodevelopmental disorders associated with autism and intellectual disabilities. Cold Spring Harb. Perspect. Biol. 4, a009886 (2012).
Robertson, C. E. & Baron-Cohen, S. Sensory perception in autism. Nat. Rev. Neurosci. 18, 671–684 (2017).
Javitt, D. C. & Sweet, R. A. Auditory dysfunction in schizophrenia: integrating clinical and basic features. Nat. Rev. Neurosci. 16, 535–550 (2015).
Peron, S. P., Freeman, J., Iyer, V., Guo, C. & Svoboda, K. A cellular resolution map of barrel cortex activity during tactile behavior. Neuron 86, 783–799 (2015).
Feldmeyer, D. et al. Barrel cortex function. Prog. Neurobiol. 103, 3–27 (2013).
Pluta, S. R., Lyall, E. H., Telian, G. I., Ryapolova-Webb, E. & Adesnik, H. Surround integration organizes a spatial map during active sensation. Neuron 94, 1220–1233.e5 (2017).
Ogden, K. K., Ozkan, E. D. & Rumbaugh, G. Prioritizing the development of mouse models for childhood brain disorders. Neuropharmacology 100, 2–16 (2016).
Hoischen, A., Krumm, N. & Eichler, E. E. Prioritization of neurodevelopmental disease genes by discovery of new mutations. Nat. Neurosci. 17, 764–772 (2014).
Deciphering Developmental Disorders, S.; Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
Hamdan, F. F. et al. Mutations in SYNGAP1 in autosomal nonsyndromic mental retardation. N. Engl. J. Med. 360, 599–605 (2009).
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Deciphering Developmental Disorders, S.; Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433–438 (2017).
O’Roak, B. J. et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nat. Commun. 5, 5595 (2014).
Hamdan, F. F. et al. De novo SYNGAP1 mutations in nonsyndromic intellectual disability and autism. Biol. Psychiatry 69, 898–901 (2011).
Berryer, M. H. et al. Mutations in SYNGAP1 cause intellectual disability, autism, and a specific form of epilepsy by inducing haploinsufficiency. Hum. Mutat. 34, 385–394 (2013).
Parker, M. J. et al. De novo, heterozygous, loss-of-function mutations in SYNGAP1 cause a syndromic form of intellectual disability. Am. J. Med. Genet. A. 167A, 2231–2237 (2015).
Mignot, C. et al. Genetic and neurodevelopmental spectrum of SYNGAP1-associated intellectual disability and epilepsy. J. Med. Genet. 53, 511–522 (2016).
Carvill, G. L. et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat. Genet. 45, 825–830 (2013).
von Stülpnagel, C. et al. SYNGAP1 mutation in focal and generalized epilepsy: a literature overview and a case report with special aspects of the EEG. Neuropediatrics 46, 287–291 (2015).
Clement, J. P. et al. Pathogenic SYNGAP1 mutations impair cognitive development by disrupting maturation of dendritic spine synapses. Cell 151, 709–723 (2012).
Ozkan, E. D. et al. Reduced cognition in Syngap1 mutants is caused by isolated damage within developing forebrain excitatory neurons. Neuron 82, 1317–1333 (2014).
Agency for Healthcare Research and Quality. Registries for Evaluating Patient Outcomes: A User’s Guide [Internet]: Rare Disease Registries. (Gliklich, R. E., Dreyer, N. A., & Leavy, M. B.) Chapter 20 (US Department of Health & Human Services, Rockville, MD, USA, 2014).
Berryer, M. H. et al. Decrease of SYNGAP1 in GABAergic cells impairs inhibitory synapse connectivity, synaptic inhibition and cognitive function. Nat. Commun. 7, 13340 (2016).
Clement, J. P., Ozkan, E. D., Aceti, M., Miller, C. A. & Rumbaugh, G. SYNGAP1 links the maturation rate of excitatory synapses to the duration of critical-period synaptic plasticity. J. Neurosci. 33, 10447–10452 (2013).
Dana, H. et al. Thy1-GCaMP6 transgenic mice for neuronal population imaging in vivo. PLoS One 9, e108697 (2014).
Gonçalves, J. T., Anstey, J. E., Golshani, P. & Portera-Cailliau, C. Circuit level defects in the developing neocortex of Fragile X mice. Nat. Neurosci. 16, 903–909 (2013).
Wolfe, J., Houweling, A. R. & Brecht, M. Sparse and powerful cortical spikes. Curr. Opin. Neurobiol. 20, 306–312 (2010).
Margolis, D. J. et al. Reorganization of cortical population activity imaged throughout long-term sensory deprivation. Nat. Neurosci. 15, 1539–1546 (2012).
Gorski, J. A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Gray, L. T. et al. Layer-specific chromatin accessibility landscapes reveal regulatory networks in adult mouse visual cortex. eLife 6, e21883 (2017).
Diamond, M. E. & Arabzadeh, E. Whisker sensory system - from receptor to decision. Prog. Neurobiol. 103, 28–40 (2013).
Nelson, S. B. & Valakh, V. Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87, 684–698 (2015).
Muhia, M., Yee, B. K., Feldon, J., Markopoulos, F. & Knuesel, I. Disruption of hippocampus-regulated behavioural and cognitive processes by heterozygous constitutive deletion of SynGAP. Eur. J. Neurosci. 31, 529–543 (2010).
Guo, X. et al. Reduced expression of the NMDA receptor-interacting protein SynGAP causes behavioral abnormalities that model symptoms of schizophrenia. Neuropsychopharmacology 34, 1659–1672 (2009).
Sachidhanandam, S., Sermet, B. S. & Petersen, C. C. H. Parvalbumin-expressing GABAergic neurons in mouse barrel cortex contribute to gating a goal-directed sensorimotor transformation. Cell Rep. 15, 700–706 (2016).
Yang, H., Kwon, S. E., Severson, K. S. & O’Connor, D. H. Origins of choice-related activity in mouse somatosensory cortex. Nat. Neurosci. 19, 127–134 (2016).
Sachidhanandam, S., Sreenivasan, V., Kyriakatos, A., Kremer, Y. & Petersen, C. C. Membrane potential correlates of sensory perception in mouse barrel cortex. Nat. Neurosci. 16, 1671–1677 (2013).
Ben-Sasson, A. et al. A meta-analysis of sensory modulation symptoms in individuals with autism spectrum disorders. J. Autism Dev. Disord. 39, 1–11 (2009).
Marco, E. J., Hinkley, L. B., Hill, S. S. & Nagarajan, S. S. Sensory processing in autism: a review of neurophysiologic findings. Pediatr. Res. 69, 48R–54R (2011).
Robertson, A. E. & Simmons, D. R. The relationship between sensory sensitivity and autistic traits in the general population. J. Autism Dev. Disord. 43, 775–784 (2013).
Araki, Y., Zeng, M., Zhang, M. & Huganir, R. L. Rapid dispersion of SynGAP from synaptic spines triggers AMPA receptor insertion and spine enlargement during LTP. Neuron 85, 173–189 (2015).
Kopanitsa, M. V. et al. Chronic treatment with a MEK inhibitor reverses enhanced excitatory field potentials in Syngap1 +/– mice. Pharmacol. Rep. 70, 777–783 (2018).
Rumbaugh, G., Adams, J. P., Kim, J. H. & Huganir, R. L. SynGAP regulates synaptic strength and mitogen-activated protein kinases in cultured neurons. Proc. Natl. Acad. Sci. USA 103, 4344–4351 (2006).
Vazquez, L. E., Chen, H. J., Sokolova, I., Knuesel, I. & Kennedy, M. B. SynGAP regulates spine formation. J. Neurosci. 24, 8862–8872 (2004).
Walkup, W. G. et al. A model for regulation by SynGAP-alpha1 of binding of synaptic proteins to PDZ-domain ‘Slots’ in the postsynaptic density. eLife 5, e16813 (2016).
Zeng, M., Bai, G. & Zhang, M. Anchoring high concentrations of SynGAP at postsynaptic densities via liquid-liquid phase separation. Small GTPases https://doi.org/10.1080/21541248.2017.1320350 (2017).
Aceti, M. et al. Syngap1 haploinsufficiency damages a postnatal critical period of pyramidal cell structural maturation linked to cortical circuit assembly. Biol. Psychiatry 77, 805–815 (2015).
Kim, J. H., Lee, H. K., Takamiya, K. & Huganir, R. L. The role of synaptic GTPase-activating protein in neuronal development and synaptic plasticity. J. Neurosci. 23, 1119–1124 (2003).
Harrison, T. C., Sigler, A. & Murphy, T. H. Simple and cost-effective hardware and software for functional brain mapping using intrinsic optical signal imaging. J. Neurosci. Methods 182, 211–218 (2009).
Chen-Bee, C. H., Kwon, M. C., Masino, S. A. & Frostig, R. D. Areal extent quantification of functional representations using intrinsic signal optical imaging. J. Neurosci. Methods 68, 27–37 (1996).
Chen-Bee, C. H. et al. Visualizing and quantifying evoked cortical activity assessed with intrinsic signal imaging. J. Neurosci. Methods 97, 157–173 (2000).
Chen-Bee, C. H. & Frostig, R. D. Variability and interhemispheric asymmetry of single-whisker functional representations in rat barrel cortex. J. Neurophysiol. 76, 884–894 (1996).
Goldey, G. J. et al. Removable cranial windows for long-term imaging in awake mice. Nat. Protoc. 9, 2515–2538 (2014).
Holtmaat, A. et al. Imaging neocortical neurons through a chronic cranial window. Cold Spring Harb. Protoc. 2012, 694–701 (2012).
Dubbs, A., Guevara, J. & Yuste, R. moco: fast motion correction for calcium imaging. Front. Neuroinform. 10, 6 (2016).
Patel, T. P., Man, K., Firestein, B. L. & Meaney, D. F. Automated quantification of neuronal networks and single-cell calcium dynamics using calcium imaging. J. Neurosci. Methods 243, 26–38 (2015).
Desai, N. S., Siegel, J. J., Taylor, W., Chitwood, R. A. & Johnston, D. Matlab-based automated patch-clamp system for awake behaving mice. J. Neurophysiol. 114, 1331–1345 (2015).
Desai, N. S. & Walcott, E. C. Synaptic bombardment modulates muscarinic effects in forelimb motor cortex. J. Neurosci. 26, 2215–2226 (2006).
Ozkan, E. D. et al. Input-specific regulation of hippocampal circuit maturation by non-muscle myosin IIB. J. Neurochem. 134, 429–444 (2015).
Agmon, A. & Connors, B. W. Thalamocortical responses of mouse somatosensory (barrel) cortex in vitro. Neuroscience 41, 365–379 (1991).
Heyser, C. J. & Chemero, A. Novel object exploration in mice: not all objects are created equal. Behav. Processes 89, 232–238 (2012).
Wu, H. P., Ioffe, J. C., Iverson, M. M., Boon, J. M. & Dyck, R. H. Novel, whisker-dependent texture discrimination task for mice. Behav. Brain Res. 237, 238–242 (2013).
Gaffield, M. A., Amat, S. B., Bito, H. & Christie, J. M. Chronic imaging of movement-related Purkinje cell calcium activity in awake behaving mice. J. Neurophysiol. 115, 413–422 (2016).
Slotnick, B. A simple 2-transistor touch or lick detector circuit. J. Exp. Anal. Behav. 91, 253–255 (2009).
He, C. X. et al. Tactile defensiveness and impaired adaptation of neuronal activity in the Fmr1 knock-out mouse model of autism. J. Neurosci. 37, 6475–6487 (2017).
Acknowledgements
This work was supported in part by NIH grants from the National Institute of Mental Health (MH096847 and MH108408 to G.R. and MH105400 to C.A.M.), the National Institute for Neurological Disorders and Stroke (NS064079 to G.R. and NS083894 to J.M.C.), and the National Institute for Drug Abuse (DA034116 and DA036376 to C.A.M.). J.L.H. is supported by a National Institute for Neurological Disorders and Stroke Mentored Clinical Scientist Research Career Development Award (NS091381). The SYNGAP1 Natural History Study and Patient Registry is supported by a grant to M.W., J.L.H., and G.R. from The National Organization of Rare Disorders (NORD).
Author information
Authors and Affiliations
Contributions
S.D.M. performed experiments, designed experiments, analyzed data, co-wrote the manuscript, and edited the manuscript. E.D.O. performed experiments, designed experiments, analyzed data, and edited the manuscript. M.A. performed experiments, designed experiments and analyzed data. S.M. performed experiments, designed experiments, and analyzed data. E.M. performed experiments, designed experiments, and analyzed data. M.W. performed experiments, designed experiments, and analyzed data. N.L. performed experiments and analyzed data. T.V. performed experiments and analyzed data. M.A.G. designed experiments and interpreted data. J.M.C. designed experiments and interpreted data. J.L.H. performed experiments, designed experiments, analyzed data, interpreted data, and edited the manuscript. C.A.M. designed experiments, interpreted data, and edited the manuscript. G.R. conceived the study, designed experiments, interpreted data, co-wrote the manuscript, and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests. M.W. is a paid employee of Bridge-the-GAP Educational Research Foundation. G.R. and J.L.H. are unpaid scientific advisors to Bridge-the-GAP Educational Research Foundation.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Impaired sensory-evoked GCaMP6s dynamics in SSC of Syngap1 mice.
(a) Example wide-field ΔF/F images superimposed over craniotomy pictures of WT and Het mice. (b-d) Traces of individual (b) and averaged (c) GCaMP6s responses, as well as a scatter plot showing the integrated ΔF/F signal amplitude (d) in response to 5 pulses at 5 Hz (Student’s t-test t(12)=4.09 p=0.0015). (e-g) Average time courses of GCaMP6s dynamics in response to a single whisker stimulation (e), 5 pulses at 5 Hz (f) and 5 pulses at 1 Hz (g) stimulation protocols. (h) Scatter plot showing ratio of fourth pulse to first pulse in 5 pulse at 1 Hz paradigm (Student’s t-test t(12)=4.16 p=0.0013). Peak (i, j) and integrated (k, l) ΔF/F responses for increasing number of pulses at 10 Hz (j, l) and increasing frequency of pulses at 5 pulses (i, k) (2-way RM-ANOVA : (i) F(1,12)=3.90 p=0.07 for genotype and F(2,24)=7.44 p=0.003 for genotype and frequency interaction; (j) F(1,12)=2.27 p=0.16 for genotype and F(4,48)=2.92 p=0.031 for genotype and frequency interaction; (k) F(1,12)=7.11 p=0.02 for genotype and F(2,24)=4.92 p=0.01 for genotype and frequency interaction; (l) F(1,12)=4.22 p=0.06 for genotype and F(4,48)=5.96 p=0.001 for genotype and frequency interaction). Open circles represent animal means, black lines or closed circles indicate population means and error bars or shaded areas indicate SEMs. Data were obtained n= 6 WT and n=8 Het mice from two cohorts of animals. All statistical tests were two-sided.
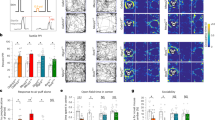
Supplementary Figure 2 Cluster analysis of ongoing and whisker-generated spike counts in SSC L2/3 neurons from awake Syngap1 mice.
(a) Relating to Fig. 2, graph showing the fraction of neurons clustered into low- (white), medium- (gray), and high-spiking (black) activity clusters. Mean spike count for each cluster is noted in the graph legend. Number of neurons in each cluster is denoted on the bar graph for each group. (b) Results of Chi-square analysis showing overall effect on how the neurons clustered across the four groups. (c) Results of Chi-square analyses demonstrating differences in each of the three activity clusters when comparing the four experimental groups. (d) Results of Chi-square analyses comparing the four groups to each other for all three activity clusters.
Supplementary Figure 3 Cell-type-specific recombination in Cux2-CreERT2 and Rbp4-Cre lines.
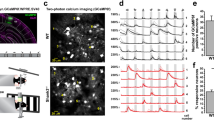
(a) Representative coronal brain sections of a Cux2-Cre+/-;Ai9+/- mouse, where Cre activity is reported by red fluorescence (black) from the TdTomato expression at PND60 (TMX injection at PND2). (b) Representative coronal brain section from a Cux2-Cre+/-;Ai9+/- mouse, where Cre activity is reported by TdTomato expression in the somatosensory cortex (red) at PND60 (TMX injection at PND2). (c) Representative coronal brain sections of a Rbp4Cre+/-;Ai6+/- mouse, where Cre activity is reported by green fluorescence (black) from the zsGreen expression at PND60. (d) Representative coronal brain section from a Rbp4Cre+/-;Ai9+/- mouse where Cre activity is reported by TdTomato expression in the somatosensory cortex (red) at PND60. Validation of the Cre driver lines were repeated in 2-3 mice per genotype.
Supplementary Figure 4 Additional performance metrics of Syngap1 mice in the go–no-go task.
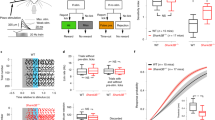
Representative learning curves during Step 2 training for a WT (a, b) and a Het (c, d) mouse depicting probability of licking on Go (black) and NoGo (blue) trials (a, c) and discrimination index (b, d). Learning curves were similar for 6/7 WT and 7/7 Het mice. (e) Normalized bodyweights of mice from one day before water restriction to the completion of the experiment (2-way RM-ANOVA with Bonferroni’s multiple comparison, Genotype: F(1,12)=1.29, p=0.28; Day: F(45,540)=6.64, p= 1.19E-29; Genotype*Day interaction: F(45,540)=0.49, p=0.99). (f, g) Scatter plots of licking behavior during lick-port training depicting mean number of water licks (f, Unpaired t-test: t(12)=0.17, p=0.87) and total reward licks (g, Unpaired t-test t(12)=0.88, p=0.40). (h, i) Averaged response times across sessions during Go (h, 2-way RM-ANOVA with Bonferroni’s multiple comparison, Genotype: F(1,12)=0.33, p=0.58; Session: F(15,180)=16.84, p= 5.965E-27; Genotype*Session interaction: F(15,180)=0.89, p=0.57) and NoGo (i, 2-way RM-ANOVA with Bonferroni’s multiple comparison, Genotype: F(1,12)=0.0059, p=0.94; Session: F(15,180)=13.94, p= 4.88E-23; Genotype*Session interaction: F(15,180)=0.95, p=0.51) trials. (j-l) For Step 2 training, mean number of trials plotted against session (j, 2-way RM-ANOVA with Bonferroni’s multiple comparison, Genotype: F(1,12)=0.88 p=0.37; Session: F(15,180)=1.88 p=0.028; Genotype*Session interaction: F(15,180)=1.56 p=0.088), scatter plots of mean trials per session (k: Unpaired t-test t(12)=1.29, p=0.22) and total trials performed (l: Unpaired t-test t(12)=0.59, p=0.56). (m) Mean number of total licks for each session in Step 2 training (2-way RM-ANOVA with Bonferroni’s multiple comparison, Genotype: F(1,12)=2.08 p=0.18; Session: F(15,180)=3.51 p= 2.73E-5; Genotype*Session interaction: F(15,180)=2.58 p=0.0016). (n) Scatter plot of mean licks per trial in Step 2 training (Mann-Whitney test, U=11, p= 0.097). (o) Scatter plot of total number of licks in Step 2 training (Unpaired t-test t(12)=1.44, p=0.18). (e-o) Open circles represent individual animals, closed circles or solid black lines indicate population means and error bars or shaded area represent the SEMs. (a,c) Solid black and blue dashed lines indicate performance criteria for hits and FAs, respectively. (b,d) Solid black line indicates performance criteria for d’. All data obtained from n=7 WT and n=7 Het mice, from two cohorts of animals. All statistical tests were two-sided.
Supplementary information
Supplementary Table 1
Registry entries with a completed sensory profile
Supplementary Table 2
Genotype of registry entries denoting touch-related sensory abnormalities
Supplementary Table 3
Up/down state properties of L2/3 barrel cortex neurons
Supplementary Video 1
Head-fixed wild-type mouse without a Botox injection
Supplementary Video 2
Head-fixed wild-type mouse after a Botox injection
Rights and permissions
About this article
Cite this article
Michaelson, S.D., Ozkan, E.D., Aceti, M. et al. SYNGAP1 heterozygosity disrupts sensory processing by reducing touch-related activity within somatosensory cortex circuits. Nat Neurosci 21, 1–13 (2018). https://doi.org/10.1038/s41593-018-0268-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0268-0
This article is cited by
-
Cortex-restricted deletion of Foxp1 impairs barrel formation and induces aberrant tactile responses in a mouse model of autism
Molecular Autism (2023)
-
Non-synaptic function of the autism spectrum disorder-associated gene SYNGAP1 in cortical neurogenesis
Nature Neuroscience (2023)
-
Kv7/KCNQ potassium channels in cortical hyperexcitability and juvenile seizure-related death in Ank2-mutant mice
Nature Communications (2023)
-
Clinical and behavioural features of SYNGAP1-related intellectual disability: a parent and caregiver description
Journal of Neurodevelopmental Disorders (2022)
-
Cortical interneurons in autism
Nature Neuroscience (2021)