Abstract
Understanding the organization of the hippocampus is fundamental to understanding brain function related to learning, memory, emotions, and diseases such as Alzheimer’s disease. Physiological studies in humans and rodents have suggested that there is both structural and functional heterogeneity along the longitudinal axis of the hippocampus. However, the recent discovery of discrete gene expression domains in the mouse hippocampus has provided the opportunity to re-evaluate hippocampal connectivity. To integrate mouse hippocampal gene expression and connectivity, we mapped the distribution of distinct gene expression patterns in mouse hippocampus and subiculum to create the Hippocampus Gene Expression Atlas (HGEA). Notably, previously unknown subiculum gene expression patterns revealed a hidden laminar organization. Guided by the HGEA, we constructed the most detailed hippocampal connectome available using Mouse Connectome Project (http://www.mouseconnectome.org) tract tracing data. Our results define the hippocampus’ multiscale network organization and elucidate each subnetwork’s unique brain-wide connectivity patterns.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All images of mouse in situ hybridization gene expression data are available at the Allen Brain Atlas website (http://www.mouse.brain-map.org). Gene accession codes for all genes and the in situ hybridization probe sequences can be found by querying the gene name through the Allen Brain Atlas website (http://www.mouse.brain-map.org). All anatomical tracer image data is available through our iConnectome viewer as part of the Mouse Connectome Project at USC (http://www.mouseconnectome.org). The HGEA stereotaxic coordinate atlas, 3-D atlas viewer, and annotation data are available for download at http://www.mouseconnectome.org/MCP/page/papers. Additional informatics and visualization tools are available online at http://www.mouseconnectome.org/Analytics/page/matrix.
References
Fanselow, M. S. & Dong, H.-W. Are the dorsal and ventral hippocampus functionally distinct structures? Neuron 65, 7–19 (2010).
Maruszak, A. & Thuret, S. Why looking at the whole hippocampus is not enough-a critical role for anteroposterior axis, subfield and activation analyses to enhance predictive value of hippocampal changes for Alzheimer’s disease diagnosis. Front. Cell. Neurosci 8, 95 (2014).
Moser, M. B. & Moser, E. I. Functional differentiation in the hippocampus. Hippocampus 8, 608–619 (1998).
Poppenk, J., Evensmoen, H. R., Moscovitch, M. & Nadel, L. Long-axis specialization of the human hippocampus. Trends Cogn. Sci. 17, 230–240 (2013).
Risold, P. Y. & Swanson, L. W. Structural evidence for functional domains in the rat hippocampus. Science 272, 1484–1486 (1996).
Swanson, L. W. & Cowan, W. M. An autoradiographic study of the organization of the efferent connections of the hippocampal formation in the rat. J. Comp. Neurol. 172, 49–84 (1977).
Strange, B. A., Witter, M. P., Lein, E. S. & Moser, E. I. Functional organization of the hippocampal longitudinal axis. Nat. Rev. Neurosci. 15, 655–669 (2014).
Amaral, D. G. Emerging principles of intrinsic hippocampal organization. Curr. Opin. Neurobiol. 3, 225–229 (1993).
Dong, H.-W., Swanson, L. W., Chen, L., Fanselow, M. S. & Toga, A. W. Genomic-anatomic evidence for distinct functional domains in hippocampal field CA1. Proc. Natl. Acad. Sci. USA 106, 11794–11799 (2009).
Thompson, C. L. et al. Genomic anatomy of the hippocampus. Neuron 60, 1010–1021 (2008).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Cembrowski, M. S., Wang, L., Sugino, K., Shields, B. C. & Spruston, N. Hipposeq: a comprehensive RNA-seq database of gene expression in hippocampal principal neurons. eLife 5, e14997 (2016).
Habib, N. et al. Div-Seq: Single-nucleus RNA-Seq reveals dynamics of rare adult newborn neurons. Science 353, 925–928 (2016).
Lein, E. S., Callaway, E. M., Albright, T. D. & Gage, F. H. Redefining the boundaries of the hippocampal CA2 subfield in the mouse using gene expression and 3-dimensional reconstruction. J. Comp. Neurol. 485, 1–10 (2005).
Cembrowski, M. S. & Spruston, N. Integrating results across methodologies is essential for producing robust neuronal taxonomies. Neuron 94, 747–751.e1 (2017).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. seqFISH accurately detects transcripts in single cells and reveals robust spatial organization in the hippocampus. Neuron 94, 752–758.e1 (2017).
Zingg, B. et al. Neural networks of the mouse neocortex. Cell 156, 1096–1111 (2014).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Dong, H. W. The Allen Reference Atlas: A Digital Color Brain Atlas of the C57Bl/6J Male Mouse. (John Wiley & Sons Inc, Hoboken, NJ, 2008).
Saunders, A. et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174, 1015–1030 (2018).
Scharfman, H. E. The enigmatic mossy cell of the dentate gyrus. Nat. Rev. Neurosci. 17, 562–575 (2016).
Lorente de Nó, R. Studies on the structure of the cerebral cortex. II. Continuation of the study of the ammonic system. J. Psychol. Neurol. 46, 113–177 (1934).
Cui, Z., Gerfen, C. R. & Young, W. S. III Hypothalamic and other connections with dorsal CA2 area of the mouse hippocampus. J. Comp. Neurol. 521, 1844–1866 (2013).
Dudek, S. M., Alexander, G. M. & Farris, S. Rediscovering area CA2: unique properties and functions. Nat. Rev. Neurosci. 17, 89–102 (2016).
Kohara, K. et al. Cell type-specific genetic and optogenetic tools reveal hippocampal CA2 circuits. Nat. Neurosci. 17, 269–279 (2014).
Sik, A., Penttonen, M., Ylinen, A. & Buzsáki, G. Hippocampal CA1 interneurons: an in vivo intracellular labeling study. J. Neurosci. 15, 6651–6665 (1995).
Ding, S. L. Comparative anatomy of the prosubiculum, subiculum, presubiculum, postsubiculum, and parasubiculum in human, monkey, and rodent. J. Comp. Neurol. 521, 4145–4162 (2013).
Vann, S. D., Aggleton, J. P. & Maguire, E. A. What does the retrosplenial cortex do? Nat. Rev. Neurosci. 10, 792–802 (2009).
Weible, A. P. Remembering to attend: the anterior cingulate cortex and remote memory. Behav. Brain Res. 245, 63–75 (2013).
Kitamura, T. et al. Engrams and circuits crucial for systems consolidation of a memory. Science 356, 73–78 (2017).
Risold, P. Y. & Swanson, L. W. Connections of the rat lateral septal complex. Brain Res. Brain Res. Rev. 24, 115–195 (1997).
Cembrowski, M. S. et al. Dissociable structural and functional hippocampal outputs via distinct subiculum cell classes. Cell 173, 1280–1292.e18 (2018).
Gauthier, J. L. & Tank, D. W. A dedicated population for reward coding in the hippocampus. Neuron 99, 179–193.e7 (2018).
O’Keefe, J. & Dostrovsky, J. The hippocampus as a spatial map: preliminary evidence from unit activity in the freely-moving rat. Brain Res. 34, 171–175 (1971).
Lever, C., Burton, S., Jeewajee, A., O’Keefe, J. & Burgess, N. Boundary vector cells in the subiculum of the hippocampal formation. J. Neurosci. 29, 9771–9777 (2009).
Hafting, T., Fyhn, M., Molden, S., Moser, M.-B. & Moser, E. I. Microstructure of a spatial map in the entorhinal cortex. Nature 436, 801–806 (2005).
Sargolini, F. et al. Conjunctive representation of position, direction, and velocity in entorhinal cortex. Science 312, 758–762 (2006).
Taube, J. S. The head direction signal: origins and sensory-motor integration. Annu. Rev. Neurosci. 30, 181–207 (2007).
Okuyama, T., Kitamura, T., Roy, D. S., Itohara, S. & Tonegawa, S. Ventral CA1 neurons store social memory. Science 353, 1536–1541 (2016).
Hitti, F. L. & Siegelbaum, S. A. The hippocampal CA2 region is essential for social memory. Nature 508, 88–92 (2014).
Fanselow, M. S. & Poulos, A. M. The neuroscience of mammalian associative learning. Annu. Rev. Psychol. 56, 207–234 (2005).
Tovote, P., Fadok, J. P. & Lüthi, A. Neuronal circuits for fear and anxiety. Nat. Rev. Neurosci. 16, 317–331 (2015).
Swanson, L. W. Cerebral hemisphere regulation of motivated behavior. Brain Res. 886, 113–164 (2000).
Canteras, N. S., Pavesi, E. & Carobrez, A. P. Olfactory instruction for fear: neural system analysis. Front. Neurosci. 9, 276 (2015).
Gross, C. T. & Canteras, N. S. The many paths to fear. Nat. Rev. Neurosci. 13, 651–658 (2012).
de Lima, M. A. X., Baldo, M. V. C. & Canteras, N. S. A role for the anteromedial thalamic nucleus in the acquisition of contextual fear memory to predatory threats. Brain Struct. Funct. 222, 113–129 (2017).
Biag, J. et al. Cyto- and chemoarchitecture of the hypothalamic paraventricular nucleus in the C57BL/6J male mouse: a study of immunostaining and multiple fluorescent tract tracing. J. Comp. Neurol. 520, 6–33 (2012).
Thompson, R. H. & Swanson, L. W. Structural characterization of a hypothalamic visceromotor pattern generator network. Brain Res. Brain Res. Rev. 41, 153–202 (2003).
Li, S. & Kirouac, G. J. Sources of inputs to the anterior and posterior aspects of the paraventricular nucleus of the thalamus. Brain Struct. Funct. 217, 257–273 (2012).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Hintiryan, H. et al. The mouse cortico-striatal projectome. Nat. Neurosci. 19, 1100–1114 (2016).
Hintiryan, H. et al. Comprehensive connectivity of the mouse main olfactory bulb: analysis and online digital atlas. Front. Neuroanat. 6, 30 (2012).
Betzel, R. F. & Bassett, D. S. Multi-scale brain networks. Neuroimage 160, 73–83 (2017).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Gu, S. et al. Controllability of structural brain networks. Nat. Commun. 6, 8414 (2015).
Lancichinetti, A. & Fortunato, S. Consensus clustering in complex networks. Sci. Rep. 2, 336 (2012).
Droettboom, M. et al. Matplotlib/Matplotlib: V2. 0.0. https://doi.org/10.5281/zenodo.248351 (2017).
Acknowledgements
This manuscript is dedicated to the memory of Sigmund J. Bienkowski. The authors would like to thank A.W. Toga, L.W. Swanson, and H. Karten for advising the Mouse Connectome Project. This work was supported by NIH/NIMH R01 MH094360-01A1 (H.-W.D), NIH/NCI U01 CA198932-01 (H.-W.D), NIH/NIMH RF1 MH114112-01 (H.-W.D), NIH/NIMH U01 MH114829-01 (H.-W.D), NIH/NIMH F32 MH107071-01-A1 (M.S.B), and a Pilot fund (H.-W.D.) of ADRC center grant P50-AG05142.
Author information
Authors and Affiliations
Contributions
M.S.B. and H.-W.D. conceived, designed and managed the project. M.S.B. and H.-W.D. wrote the manuscript. M.S.B. performed manual analysis of all raw image data, including the gene expression and connectivity annotation, created the HGEA, and prepared figures for publication. I.B. led the informatics team and wrote the code for computational network analysis. M.S.B., I.B., and H.-W.D. constructed the neural networks diagrams. M.S.B., M.Y.S., L.G., M.Z., and N.L.B. performed stereotaxic surgeries to generate anatomical connectivity data. K.C. and M.S.B. performed integrative analysis of HGEA annotation with DropViz data. T.A. created the three-dimensional HGEA atlas rendering along with online three-dimensional atlas viewer and supplementary videos. S.Y. managed the iConnectome website and created online informatics and visualization tools. J.A.-J., S.A., and D.L. performed image processing for image data upload to the iConnectome viewer. N.L.B., N.N.F., and H.H. offered constructive guidance for the manuscript edits.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Coronal Hippocampus Gene Expression Atlas (HGEA).
Following the analysis of gene expression patterns from the Allen Gene Expression Atlas (www.brain-map.org), the interpreted hippocampal gene expression domains for the DG, CA3, CA1, and SUB were mapped onto all ARA sections. Hippocampal domains are generally color-coded with dorsal, intermediate, and ventral parts following a red, green, and blue color scheme, respectively. See figure legend for corresponding colors to each domain. For a list of annotated gene expression patterns, see Supplementary Table 2. SUB subregions corresponding to the presence or absence of specific gene expression layers are segregated by arrowheads.
Supplementary Figure 2 Sagittal Hippocampal Gene Expression Atlas (HGEA).
All hippocampal areas are cropped from corresponding Allen Reference Atlas sections and gene expression boundaries were interpreted from online sagittal gene expression data (www.brain-map.org). Corresponding representative sagittal planes through a coronal section are shown in the top left. Color coding scheme is the same as the coronal version (Supplementary Fig. 1) and listed in the figure legend. Atlas level 20 in the sagittal version of the Allen Reference Atlas does not exist so the hippocampus at this level was not included.
Supplementary Figure 3 Interpretation of sectioning angle in Allen Institute in situ hybridization tissue sections and HGEA boundary mapping.
(a) Histological processing of brain tissue requires careful sectioning for accurate registration of data to ARA levels. For example, a tissue section of Nov gene expression from the Allen Brain Atlas database does not accurately register to any one ARA atlas level (80, 82, 84), because of oblique cutting angle along the dorsoventral axis during histological sectioning (dashed red lines provide guides for comparison of dorsal, intermediate and ventral parts of the tissue across all images). Rather than register to one atlas level, we used multiple landmarks throughout the tissue section (1-10, 1 = shape of corpus callosum/ external capsule white matter, 2 = position of the CA1/SUB border relative to the tip of the DG granule layer, 3 = shape of the dorsal diencephalon, 4 = position of the posterior commisure, 5 = thalamic nuclei cytoarchitecture, 6 = ventral tip of the DG granule cell layer, 7 = separation of the CA3/CA1 pyramidal layer relative to the rhinal sulcus, 8 = shape of the MM and LM, 9 = position of the CA3/CA1/SUB pyramidal cell layers, 10 = amygdala nuclei cytoarchitecture) to register tissue locally to the appropriate ARA atlas section. Ultimately, the Nov tissue section is better described by a “composite” of ARA levels 80-84, where the ventral part of the hippocampus is closer to ARA 80 and dorsal part of the hippocampus is closer to 84 with a linear progression along the dorsoventral axis. (b) Histological sectioning angle is consistent among tissue sections cut from the same brain, but can be different across many other brains. Here, we show a different tissue section from the same brain showing Nov gene expression and a tissue section cut from a different brain showing Dcn gene expression. The composite ARA sections on the left show that the Nov tissue section contains a similar dorsoventral gradient of 4 atlas levels as shown in a with the dorsal part of the tissue more caudal than the ventral part of the tissue. In contrast, the Dcn tissue section contains a steeper gradient of 7 atlas levels cut where the ventral part of the hippocampus is more caudal than the dorsal part of the hippocampus (opposite of Nov). On the right, are corresponding composite HGEA atlas levels showing that the dorsal limit of positive Dcn gene expression corresponds to the mapping of the CA1i/CA1v boundary at HGEA level 85 (arrow). (c) Oblique tissue sectioning angle can be estimated using the composite ARA atlas levels (each ARA atlas level is 0.1mm apart). For the Nov and Dcn brain tissue sections in b, we can estimate the oblique sectioning angles relative to the ARA atlas (dashed black lines) for each brain as a right triangle where the tangent of the sectioning angle is the displacement of the ARA atlas level between the dorsal and ventral part of the hippocampus over the dorsoventral distance of the hippocampus (~5.0mm). For the Dcn tissue, the sectioning angle was calculated at -8.0 degrees whereas the Nov tissue sectioning angle was calculated at 4.6 degrees. Note, the greater the distance between the two hypotenuses, the greater the inaccuracy for comparison along the dorsoventral axis. (d) Tissue section registration is not only affected by oblique sectioning angle in the dorsoventral axis but also along the mediolateral axis. An example of Calb1 gene expression in a tissue section shows a 2 section ARA atlas difference in the dorsoventral direction and a 2 section ARA atlas difference between the left and right hippocampus (arrows point to corresponding ARA level numbers for each region). Below, a graphical illustration of the plane of sectioning for the Calb1 tissue section (shaded blue) relative to ARA level 91 (clear plane with dashed lines). (e) Because of differences in dorsoventral and mediolateral sectioning angles, it is important to examine the progression of gene expression across multiple rostrocaudal coronal tissue sections. Three adjacent tissue sections containing gene expression for the CA2 marker Amigo2 (with corresponding HGEA levels below) show the rostrocaudal progression of Amigo2 corresponds to the HGEA boundaries of the CA2 with the CA1d, CA1i, and CA1v. At the most caudal level (HGEA 82), a thin lamina of Amigo2 positive cells are located in the deepest part of the pyramidal layer (red and blue boxes correspond to magnified images on right). (f) Examination of the mediolateral progression of gene expression in sagittal-cut tissue can corroborate data from coronal sections (for sagittal-cut tissue sections, differences caused by oblique sectioning angles apply to the dorsoventral and rostrocaudal axis). Amigo2 expression in three adjacent sagittal sections are shown with corresponding HGEA sagittal atlas levels below. Similar to the coronal data, the mediolateral progression shows a similar descent of Amigo2 expression across the dorsoventral axis (dorsal and ventral CA2 are separated at HGEA 4 due to the curve of the hippocampus relative to the straight cut of the tissue). In the most lateral HGEA section 1, a laminar labeling pattern of Amigo2 is shown similar to the coronal section at HGEA 82 (d). Together, examination of both coronal and sagittal data provides corroboration when mapping HGEA boundaries.
Supplementary Figure 4 Gene expression patterns compared with the sagittal HGEA version.
(a) The HGEA sagittal atlas (Supplementary Figure 2) was interpreted from sagittal-sectioned in situ hybridization gene expression data (www.brain-map.org). (a) Gene expression patterns of Ttn and Rph3a are localized within CA3dd only and both CA3dd and CA3d, respectively. (b) Coch, Peg10, and Grp genes are expressed within the CA3vv, CA1vv, and DGv, respectively. (c) Gene expression patterns progressively demarcate all regions along the dorsal/ventral hippocampal axis, from left to right and top to bottom. (d) Gene expression patterns of subiculum at sagittal ARA level 4 clearly define the SUB gene expression layers (for comparison, see Fig. 3).
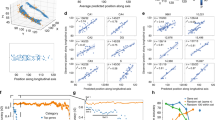
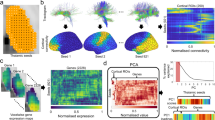
Supplementary Figure 5 Principal components analysis of DropViz data.
To compare the qualitative HGEA annotation to more quantitative single-cell RNAseq, we performed a PCA on the DropViz ‘meta-cell’ cell-types using all 32,307 genes listed in the DropViz database and compared that to a PCA using 248 HGEA genes that were also included in the DropViz database. The PCA using DropViz data from the 248 HGEA genes (top right) showed remarkablely similar clustering compared to the PCA using DropViz data from all 32,307 genes (top left). Each dot represents a DropViz ‘meta-cell’ cell-type and is colored according to their DropViz annotation as belonging to the DG, CA3/CA2, CA1, SUB, or interneuron (INT) groups (corresponding colored ellipses contain the distribution of each cell-type group). Both the 32,307 and 248 gene PCA plots show similar distribution of cell-types with HC_4-1 and HC_6-1 as extreme outliers compared to other cell-types. In the second set of PCA plots (middle row), we removed HC_4-1 and HC_6-1 (keeping everything within red box from top row plots) and clustering from both datasets remained relatively similar with minor differences in the overlap of some cell-type groups. In both plots, a dense cluster of INT overlapped with cell-types from other hippocampal regions. To try and get a clearer understanding of the distribution of cell-types within the dense INT cluster we performed a 3rd set of PCA plots (bottom row; using only cell-types within the middle row red box). In the 3rd set of PCA plots, both datasets show specific CA1 and SUB cell-types are distributed within the INT cluster (the 32,307 gene PCA also contains some CA3 cell-types). Overall, these PCA plots show that the more limited 248 HGEA genes largely recapitulate the overall organization of hippocampal cell-types described by more than 32,000 genes.
Supplementary Figure 6 Dentate gyrus gene expression and connectivity.
(a) Gene expression patterns within the DG revealed three gene expression domains within the granule layer and two distinct cell- groups within the polymorph layer. Images show DG mapped gene expression subdomains from HGEA level 89 aligned with example gene expression patterns (arrowheads mark the boundaries between regions in all images). Btg1 is highly expressed in all parts of the DG granule cell layer, whereas Slc26a10 and Trhr are restricted to dorsal and ventral parts of the granule cell layer respectively. The DGi is defined as the area of overlap between these two unique expression patterns. In the polymorph layer, we found two expression patterns that suggest dorsal (ex. Bid) and ventral (ex. Etv1) cell groups which are clearly segregated at rostral levels but more mixed at caudal levels (see Supplementary Figure 4). (b) DGpo was revealed to contain at least 2 distinct gene expression populations, the DGpod and DGpov. Tracer coinjection into the DGpod near the DGi produced bilateral anterogradely-labeled fibers in the inner molecular layer of the DGd and DGi and bilateral retrogradely-labeled neurons within the DGpov (SW110914-04A). In contrast, retrograde tracer injection in the DGd only labels neurons within the DGpod (SW110914-02A). Anterograde tracer injection within the DGpov produces bilateral fiber labeling within the DGv and DGi only (SW160909-01A). Taken together, the DGpo neuron populations provide a primarily unidirectional ventral-to-dorsal hippocampus pathway. (c) The three DG granule layer subregions (DGd, DGi, and DGv) are differentially connected to the CA3 regions as shown by multiple cases with DG anterograde tracer injections. All DGd granule cell mossy fibers display a hairpin connectivity motif, starting locally onto the DGpo, then travelling rostrally for up to 1mm before turning caudally (innervating multiple levels of CA3dd, CA3d, or CA3id along the way depending on DG granule cell location) to eventually terminate at a topographic location within the CA2 (yellow arrows follow the mossy fiber path across rostrocaudal sections). The geometric length of the mossy fiber pathway is limited by the rostral pole of the hippocampus. (d) In contrast to the hairpin connectivity motif of the DGd granule cells, granule cells within the DGi and DGv project rostrally in a linear orientation through the stratum lucidum of the CA3ic and CA3v/CA3vv, respectively. DGi and DGv mossy fiber pathways do not innervate the CA2. (e) Retrograde injection into the caudal ventral CA2 shows that the CA2 receives convergent input from DG granule cells across multiple rostrocaudal levels. Note the location of retrogradely-labeled DG neurons within the granule layer changes such that DG neurons at rostral levels (right) are close to CA3 and labeled neurons at caudal levels (left) are near the tip of the granule cell layer. (f) Retrograde injections within the CA3v and CA3vv retrogradely labels separate neuron populations within the DGv. For the number of tracer experiments and cross-validated results, see the Supplementary Methods.
Supplementary Figure 7 Complex relationship of CA3 input and SUB output within the CA1d.
(a) In case SW151029-03A, AAV (green) injection into the CA3dd (some spread into CA1d) and PHAL injection into CA3d (magenta) reveal bilateral topographically-organized projections within CA1d). CA3dd project to more distal parts of CA1d, whereas CA3d projects to more proximal parts. However, the proportional distribution within CA1d is different within CA1dr vs. CA1dc: CA3dd fibers are distributed more broadly in CA1dr (see b) whereas CA3d fibers are more prevalent in CA1dc. At caudal levels of the hippocampus (HGEA 87 below), CA3d fibers (magenta) cover a larger proportion of the CA1dc compared to CA3dd fibers (green). (b) Bilateral distribution of the CA3dd and CA3d fibers corresponds well to the distribution of proximally- and distally-enriched CA1d gene expression (Psrc1 more proximally enriched vs. Car2 more distally-enriched). Corresponding expression levels of each gene from Allen Brain Atlas are shown below. (c) SUBdd retrograde tracer injections labeled neurons within the CA1dr, while retrograde tracer injections within the ProSUB (at a similar rostrocaudal level) labeled neurons within the CA1dc and CA2. (d) Coinjections into the CA1dr and CA1dc confirmed the differential connectivity and further showed that the CA1dr sends its output specifically to SUBdd layer 1. In contrast, the CA1dc projects to ProSUB layers 3 and 4. Taken together, the proximodistal CA3 fiber topography within both the CA1dr and CA1dc suggest that both SUBdd and ProSUB receive multi-synaptic inputs from CA3dd and CA3d/CA3id, but SUBdd activity may be more biased by information from CA3dd, whereas ProSUB activity is more biased by information from CA3d/CA3id (see diagram a). In situ hybridization images from Allen Institute website (www.mouse.brain-map.org). For the number of tracer experiments and cross-validated results, see the Supplementary Methods.
Supplementary Figure 8 CA3-to-CA1 projection topography aligns with HGEA delineation.
Full map of CA3 subregion projections to progressively more ventral CA1 and SUB regions (arrowheads demarcate borders of HGEA atlas on the left). CA3dd anterograde labeling targets CA1d specifically at all rostrocaudal levels, but is denser within the medial CA1d (note the labeling at caudal levels ends at the CA1d/CA1i border). The distribution of CA3d and CA3id projection fibers within CA1 is highly similar. Both CA3d and CA3id fibers are complementary to the CA3dd fibers in CA1d (medial vs. lateral topography, see Supplementary Fig. 6), but also continues along the ventral axis to innervate CA2 and CA1i. Note the position of the CA1i/CA1v border aligns with the ventral extent of the CA3d fiber labeling whereas CA3id terminal fields do extend into the CA1v at caudal levels. In contrast to CA3id, CA3ic fibers do not target the CA1d, but instead target CA2, CA1i, CA1v, and the SUBv. Note the sharp boundaries adjacent to the CA1d border. Finally, CA3v fibers innervate the CA1v, SUBv, and SUBvv, whereas CA3vv fibers specifically target the CA1vv and ventral tip of the SUBvv layer 2. For the number of tracer experiments and cross-validated results, see the Supplementary Methods.
Supplementary Figure 9 Differential connectivity with the amygdala, nucleus accumbens, and BST.
(a) Amygdala-projecting neurons are distributed through layer 3 in the ProSUB, SUBv, and SUBvv whereas anterogradely-labeled fibers from the amygdala terminate primarily in the molecular layer (dashed white lines outline the pyramidal layer). Most cortical amygdala nuclei (COApl, PAA, COAa, etc.) terminate in the superficial part of the molecular layer, but the BMAp and BLAp innervate the deep part of the molecular layer and layer 3 and the caudal BLAa has an entirely unique projection pattern (see Fig. 7b). (b) Injection of anterograde and/or retrograde tracers into the SUBv and SUBvv reveals two distinct topographic connectivity patterns within the amygdala (dashed white lines outline the basolateral amygdala complex). SUBvv fibers and retrograde labeling are more robust in medial amygdala nuclei (particularly MEApd) whereas SUBv fibers and retrograde labeling are denser in BMAp, BLAp, and parts of the LA. (c) SUB projections target topographically distinct zones within the ACB (dashed white lines outline approximate boundaries of ACB). ProSUB fibers innervate the lateral core and shell of the ACB whereas SUBv fibers target the medial ACB core and middle part of the ACB shell. The SUBvv projects distinctly to the dorsomedial tip of the ACB shell and the CA1v projections overlap the area of the combined SUBv and SUBvv terminal fields. (d) Differential innervation of BST subnuclei by hippocampal inputs (white dashed lines outline BST subnuclei, anterior BST on top, posterior BST on bottom). The anterior BST is primarily innervated by the SUBv and SUBvv. Few ProSUB fibers enter into the BST. SUBv fibers are located throughout the anterior BST while SUBvv fibers are restricted to the BSTam/BSTdm. In the posterior BST, the BSTif/tr region is commonly innervated by ProSUB, SUBv, SUBvv, and CA1v. For the number of tracer experiments and cross-validated results, see the Supplementary Methods.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Supplementary Table 2
Gene Annotation List. Gene names are listed in alphabetical order in the left column and hippocampal regions are listed in the top row. Positive gene expression within a hippocampal subregion is marked by an ‘x’ within the cell. Empty cells denote the absence of gene expression.
Supplementary Table 3
Injection Site List. All case numbers reflect individual animals and are listed in numerical order. Since multiple tracers are injected in a single animal, injections sites and tracers are listed separately for each animal. Additionally, figure numbers that show images of labeling from an animal are listed. All raw image data is available at the MCP website.
Supplementary Table 4
Rogers-Tanimoto Dissimilarity Matrix comparing HGEA regions with DropViz cell-types. To compare the similarity/dissimilarity gene expression within HGEA subregions (binary annotation) with DropViz ‘meta-cell’ cell-types (quantitative counts of unique molecular identifiers (UMI), we first binarized the DropViz ‘meta-cell’ data after thresholding out UMI counts less than 30. The Rogers-Tanimoto dissimilarity matrix lists DropViz ‘meta-cells’ on the vertical axis and HGEA subregions on the horizontal axis (DropViz ‘meta-cells’ arranged so that similar meta-cells are closer together). Higher coefficient values represent greater dissimilarity (green shading) whereas lower coefficient values represent greater similarity (red shading). In most cases, DropViz meta-cells contain greater similarity for their corresponding HGEA subregions (cells outlined in black) and are dissimilar to other HGEA subregions (i.e. CA3 cell-types are similar to HGEA CA3 regions and not DG subregions). Notably, HGEA genes that were annotated as expressed within ‘putative interneurons’ showed a strong similarity for DropViz interneuron cell-types. Note HC_6-5 and HC_6-6 are annotated as questionable “Dentate hilum” neurons (also known as DGpo) but are also included in the DropViz CA3/CA2 ‘global cluster’.
Supplementary Table 5
HGEA subregion input/output connectivity annotation. Top row lists all hippocampal subregions and all regions that provide input and/or output to each hippocampal subregion are alphabetically listed below in columns.
Supplementary Table 6
Intrahippocampal Weighted Connectivity Annotation Data. Connections among the HGEA subregions were manually annotated and weighted (0-3). Connection weighting: 0 = absent, 1 = sparse, 2 = moderate, 3 = dense.
Supplementary Table 7
Combined lists of HGEA subregion gene expression, input, and output connectivity. Top row lists all hippocampal subregions and columns list gene expression and brain areas that provide input and/or output for each subregion (alphabetically listed). For SUB regions, genes expression is listed independently for each gene expression layer.
Supplementary Video 1
3-D rendering with all HGEA subregions. 3-D hippocampus composed of HGEA subregions within a whole mouse brain. Movie shows the DG/CA3, CA2, CA1, and SUB subregions as the camera rotates around the hippocampus. All subregion colors correspond to HGEA atlas in Supplementary Figure 1.
Supplementary Video 2
3-D rendering of DG/CA3 subregions. 3D hippocampus composed of HGEA subregions. Movie shows the DG/CA3 subregions as the camera rotates around the hippocampus. Other HGEA subregions are opaque. All subregion colors correspond to HGEA atlas in Supplementary Figure 1.
Supplementary Video 3
3-D rendering of the CA2 subregion. 3D hippocampus composed of HGEA subregions. Movie shows the CA2 subregion as the camera rotates around the hippocampus. Other HGEA subregions are opaque. All subregion colors correspond to HGEA atlas in Supplementary Figure 1.
Supplementary Video 4
3-D rendering of CA1 subregions. 3D hippocampus composed of HGEA subregions. Movie shows the CA1 subregions as the camera rotates around the hippocampus. Other HGEA subregions are opaque. All subregion colors correspond to HGEA atlas in Supplementary Figure 1.
Supplementary Video 5
3-D rendering of subiculum subregions. 3D hippocampus composed of HGEA subregions. Movie shows the SUB subregions as the camera rotates around the hippocampus. Other HGEA subregions are opaque. All subregion colors correspond to HGEA atlas in Supplementary Figure 1.
Supplementary Video 6
3-D Rabies-labeled neurons within the SUBdd and ProSUB. 3-D flythrough from a 200μm coronal Z-stack through the dorsal subiculum. At the beginning of the movie, tissue is oriented so that medial is left, lateral is right. RSPv-projecting neurons (red, G-deleted rabies-RFP) are distributed within the SUBdd largely segregated from LHA-projecting neurons (green, G-deleted rabies-GFP) within the ProSUB (blue cells stained with Neurotrace blue).
Rights and permissions
About this article
Cite this article
Bienkowski, M.S., Bowman, I., Song, M.Y. et al. Integration of gene expression and brain-wide connectivity reveals the multiscale organization of mouse hippocampal networks. Nat Neurosci 21, 1628–1643 (2018). https://doi.org/10.1038/s41593-018-0241-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0241-y
This article is cited by
-
A hippocampus-accumbens code guides goal-directed appetitive behavior
Nature Communications (2024)
-
Immunohistochemical field parcellation of the human hippocampus along its antero-posterior axis
Brain Structure and Function (2024)
-
Specific AAV2/PHP.eB-mediated gene transduction of CA2 pyramidal cells via injection into the lateral ventricle
Scientific Reports (2023)
-
Combinatorial quantification of distinct neural projections from retrograde tracing
Nature Communications (2023)
-
Inferring neuron-neuron communications from single-cell transcriptomics through NeuronChat
Nature Communications (2023)