Abstract
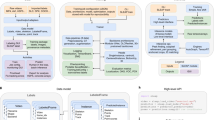
Quantifying behavior is crucial for many applications in neuroscience. Videography provides easy methods for the observation and recording of animal behavior in diverse settings, yet extracting particular aspects of a behavior for further analysis can be highly time consuming. In motor control studies, humans or other animals are often marked with reflective markers to assist with computer-based tracking, but markers are intrusive, and the number and location of the markers must be determined a priori. Here we present an efficient method for markerless pose estimation based on transfer learning with deep neural networks that achieves excellent results with minimal training data. We demonstrate the versatility of this framework by tracking various body parts in multiple species across a broad collection of behaviors. Remarkably, even when only a small number of frames are labeled (~200), the algorithm achieves excellent tracking performance on test frames that is comparable to human accuracy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data are available from the corresponding author upon reasonable request.
References
Tinbergen, N. On aims and methods of ethology. Z. Tierpsychol. 20, 410–433 (1963).
Bernstein, N. A. The Co-ordination and Regulation of Movements Vol. 1 (Pergamon, Oxford and New York, 1967).
Krakauer, J. W., Ghazanfar, A. A., Gomez-Marin, A., MacIver, M. A. & Poeppel, D. Neuroscience needs behavior: correcting a reductionist bias. Neuron 93, 480–490 (2017).
Ota, N., Gahr, M. & Soma, M. Tap dancing birds: the multimodal mutual courtship display of males and females in a socially monogamous songbird. Sci. Rep. 5, 16614 (2015).
Wade, N. J. Capturing motion and depth before cinematography. J. Hist. Neurosci. 25, 3–22 (2016).
Dell, A. I. et al. Automated image-based tracking and its application in ecology. Trends Ecol. Evol. 29, 417–428 (2014).
Gomez-Marin, A., Paton, J. J., Kampff, A. R., Costa, R. M. & Mainen, Z. F. Big behavioral data: psychology, ethology and the foundations of neuroscience. Nat. Neurosci. 17, 1455–1462 (2014).
Anderson, D. J. & Perona, P. Toward a science of computational ethology. Neuron 84, 18–31 (2014).
Winter, D. A. Biomechanics and Motor Control of Human Movement (Wiley, Hoboken, NJ, USA, 2009).
Vargas-Irwin, C. E. et al. Decoding complete reach and grasp actions from local primary motor cortex populations. J. Neurosci. 30, 9659–9669 (2010).
Wenger, N. et al. Closed-loop neuromodulation of spinal sensorimotor circuits controls refined locomotion after complete spinal cord injury. Sci. Transl. Med. 6, 255ra133 (2014).
Maghsoudi, O. H., Tabrizi, A. V., Robertson, B. & Spence, A. Superpixels based marker tracking vs. hue thresholding in rodent biomechanics application. Preprint at https://arxiv.org/abs/1710.06473 (2017).
Pérez-Escudero, A., Vicente-Page, J., Hinz, R. C., Arganda, S. & de Polavieja, G. G. idTracker: tracking individuals in a group by automatic identification of unmarked animals. Nat. Methods 11, 743–748 (2014).
Nakamura, T. et al. A markerless 3D computerized motion capture system incorporating a skeleton model for monkeys. PLoS One 11, e0166154 (2016).
de Chaumont, F. et al. Computerized video analysis of social interactions in mice. Nat. Methods 9, 410–417 (2012).
Matsumoto, J. et al. A 3D-video-based computerized analysis of social and sexual interactions in rats. PLoS One 8, e78460 (2013).
Uhlmann, V., Ramdya, P., Delgado-Gonzalo, R., Benton, R. & Unser, M. FlyLimbTracker: An active contour based approach for leg segment tracking in unmarked, freely behaving Drosophila. PLoS One 12, e0173433 (2017).
Felzenszwalb, P. F. & Huttenlocher, D. P. Pictorial structures for object recognition. Int. J. Comput. Vis. 61, 55–79 (2005).
Toshev, A. & Szegedy, C. DeepPose: human pose estimation via deep neural networks. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 1653–1660 (IEEE, Piscataway, NJ, USA, 2014).
Dollár, P., Welinder, P. & Perona, P. Cascaded pose regression. in IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2010 1078–1085 (IEEE, Piscataway, NJ, USA, 2010).
Machado, A. S., Darmohray, D. M., Fayad, J., Marques, H. G. & Carey, M. R. A quantitative framework for whole-body coordination reveals specific deficits in freely walking ataxic mice. Elife 4, e07892 (2015).
Guo, J. Z. et al. Cortex commands the performance of skilled movement. Elife 4, e10774 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. in Advances in Neural Information Processing Systems Vol. 25 (eds. Pereira, F. et al.) 1097–1105 (Curran Associates, Red Hook, NY, USA, 2012).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. in P roceedings of the IEEE Conference on Computer Vision and Pattern Recognition 770–778 (IEEE, Piscataway, NJ, USA, 2016).
Wei, S.-E., Ramakrishna, V., Kanade, T. & Sheikh, Y. Convolutional pose machines. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 4724–4732 (IEEE, Piscataway, NJ, USA, 2016).
Pishchulin, L. et al. DeepCut: joint subset partition and labeling for multi person pose estimation. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 4929–4937 (IEEE, Piscataway, NJ, USA, 2016).
Insafutdinov, E., Pishchulin, L., Andres, B., Andriluka, M. & Schiele, B. DeeperCut: a deeper, stronger, and faster multi-person pose estimation model. in European Conference on Computer Vision 34–50 (Springer, New York, 2016).
Feichtenhofer, C., Pinz, A. & Zisserman, A. Detect to track and track to detect. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 3038–3046 (IEEE, Piscataway, NJ, USA, 2017).
Insafutdinov, E. et al. ArtTrack: articulated multi-person tracking in the wild. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 1293–1301 (IEEE, Piscataway, NJ, USA, 2017).
Andriluka, M., Pishchulin, L., Gehler, P. & Schiele, B. 2D human pose estimation: new benchmark and state of the art analysis. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 3686–3693 (IEEE, Piscataway, NJ, USA, 2014).
Donahue, J. et al. DeCaf: a deep convolutional activation feature for generic visual recognition. in I nternational Conference on Machine Learning 647–655 (PMLR, Beijing, 2014).
Yosinski, J., Clune, J., Bengio, Y. & Lipson, H. How transferable are features in deep neural networks? in Advances in Neural Information Processing Systems 3320–3328 (Curran Associates, Red Hook, NY, USA, 2014).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning Vol. 1 (MIT Press, Cambridge, MA, USA, 2016).
Kümmerer, M., Wallis, T. S. & Bethge, M. DeepGaze II: reading fixations from deep features trained on object recognition. Preprint at https://arxiv.org/abs/1610.01563 (2016).
Khan, A. G., Sarangi, M. & Bhalla, U. S. Rats track odour trails accurately using a multi-layered strategy with near-optimal sampling. Nat. Commun. 3, 703 (2012).
Li, Y. et al. Neuronal representation of social information in the medial amygdala of awake behaving mice. Cell 171, 1176–1190.e17 (2017).
Robie, A. A., Seagraves, K. M., Egnor, S. E. & Branson, K. Machine vision methods for analyzing social interactions. J. Exp. Biol. 220, 25–34 (2017).
Mathis, M. W., Mathis, A. & Uchida, N. Somatosensory cortex plays an essential role in forelimb motor adaptation in mice. Neuron 93, 1493–1503.e6 (2017).
Drai, D. & Golani, I. SEE: a tool for the visualization and analysis of rodent exploratory behavior. Neurosci. Biobehav. Rev. 25, 409–426 (2001).
Sousa, N., Almeida, O. F. X. & Wotjak, C. T. A hitchhiker’s guide to behavioral analysis in laboratory rodents. Genes Brain Behav. 5 (Suppl. 2), 5–24 (2006).
Gomez-Marin, A., Partoune, N., Stephens, G. J., Louis, M. & Brembs, B. Automated tracking of animal posture and movement during exploration and sensory orientation behaviors. PLoS One 7, e41642 (2012).
Ben-Shaul, Y. OptiMouse: a comprehensive open source program for reliable detection and analysis of mouse body and nose positions. BMC Biol. 15, 41 (2017).
Zhang, C., Bengio, S., Hardt, M., Recht, B. & Vinyals, O. Understanding deep learning requires rethinking generalization. Preprint at https://arxiv.org/abs/1611.03530 (2016).
Berman, G. J. Measuring behavior across scales. BMC Biol. 16, 23 (2018).
Kim, C. K., Adhikari, A. & Deisseroth, K. Integration of optogenetics with complementary methodologies in systems neuroscience. Nat. Rev. Neurosci. 18, 222–235 (2017).
Stauffer, C. & Grimson, W.E.L. Adaptive background mixture models for real-time tracking. in IEEE Computer Society Conference on C omputer Vision and Pattern Recognition, 1999 Vol. 2, 246–252 (IEEE, Piscataway, NJ, USA, 1999).
Ristic, B., Arulampalam, S. & Gordon, N. Beyond the Kalman Filter: Particle Filters for Tracking Applications (Artech House, Norwood, MA, USA, 2003).
Carreira, J. & Zisserman, A. Quo vadis, action recognition? A new model and the kinetics dataset. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 4724–4733 (IEEE, Piscataway, NJ, USA, 2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Abadi, M. et al. TensorFlow: a system for large-scale machine learning. Preprint at https://arxiv.org/abs/1605.08695 (2016).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Acknowledgements
We are grateful to E. Insafutdinov and C. Lassner for suggestions on how to best use the TensorFlow implementation of DeeperCut. We thank N. Uchida for generously providing resources for the joystick behavior and R. Axel for generously providing resources for the Drosophila research. We also thank A. Hoffmann, J. Rauber, T. Nath, D. Klindt and T. DeWolf for a critical reading of the manuscript, as well as members of the Bethge lab, especially M. Kümmerer, for discussions. We also thank the β-testers for trying our toolbox and sharing their results with us. Funding: Marie Sklodowska-Curie International Fellowship within the 7th European Community Framework Program under grant agreement No. 622943 and DFG grant MA 6176/1-1 (A.M.); Project ALS (Women and the Brain Fellowship for Advancement in Neuroscience) and a Rowland Fellowship from the Rowland Institute at Harvard (M.W.M.); German Science Foundation (DFG) through the CRC 1233 on “Robust Vision” and from IARPA through the MICrONS program (M.B.).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.M., M.W.M. and M.B. Software: A.M. and M.W.M. Formal analysis: A.M. Experiments: A.M. and V.N.M. (trail-tracking), M.W.M. (mouse reaching), K.M.C. (Drosophila). Labeling: P.M., K.M.C., T.A., M.W.M., A.M. Writing: A.M. and M.W.M. with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
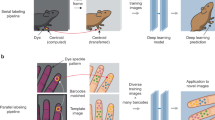
Supplementary Figure 1 Illustration of labeled data set.
a: Example images with human applied labels (first and second trial by the same labeler colored in yellow and cyan) illustrating the variability. Top row shows full frames that were labeled and illustrate typical examples of the 1,080 random frames, which comprise the data set. Rows below are cropped for visibility of the labels and indicate the average RMSE in pixels across all body parts above the image. The scorer was highly accurate, as illustrated by b. b: x-axis and y-axis difference in pixels between the first - second trial. Only a few labels strongly deviate between the two trials. Most errors are smaller than 5 pixels as can be seen in the histogram of trial-by-trial labeling errors (cropped at 20 pixels), c).
Supplementary Figure 2 Cross-validating model parameters and testing deeper networks.
a: Average training and test error for the same 6 splits with 10% training set size as in Fig. 2f with standard augmentation (i.e. scaling), augmentation by rotations as well as rotations and translations (see Methods). Although with augmentation there are 8 and 24 times as many training samples (rotations and rotations + translations, respectively), the training and test errors remained comparable. b: Average training and test error for the same 3 splits with 50% training set size for three different architectures: ResNet-50, ResNet-101 as well as ResNet-101ws, where part loss layers are added to conv4 bank29. For these networks the training error is strongly reduced, and the test performance modestly improved, indicating that the deeper networks do not over-fit (but do not offer radical improvement). Averaged over 3 splits, individual simulation results shown in as faint lines. The deeper networks reach human level accuracy on test set. The data for ResNet-50 is also depicted in Fig. 2d. c: Cross validating model parameters for ResNet-50 and 50%-training set fraction. We varied the distance variable ϵ, which determines the width of the score-map template during training around the ground-truth value with scale variable 100% (otherwise the scale ratio of the output layer was set to 80% relative to the input image size). Varying distance parameters only mildly improves the test performance (after 500k training steps). The average performance for scale 0.8 and ϵ=17 is indicated by horizontal lines (from Fig. 2d). In particular, for smaller distance parameters the RMSE increases and learning proceeds much slower (c,d). d-e: Evolution of the training and test errors at various states of the network training for various distance variables ϵ corresponding to c.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2
Supplementary Video 1
Odor guided navigation task with automated tracking of the snout. (Related to Fig. 3.)
Supplementary Video 2
Drosophila egg-laying behavior with automated tracking of various body parts. (Related to Fig. 6.)
Supplementary Video 3
Skilled reach and pull task with automated tracking of the hand. (Related to Fig. 7.)
Rights and permissions
About this article
Cite this article
Mathis, A., Mamidanna, P., Cury, K.M. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat Neurosci 21, 1281–1289 (2018). https://doi.org/10.1038/s41593-018-0209-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0209-y
This article is cited by
-
Replacement of tibialis cranialis tendon with polyester, silicone-coated artificial tendon preserves biomechanical function in rabbits compared to tendon excision only
Journal of Orthopaedic Surgery and Research (2024)
-
Enhancing surgical performance in cardiothoracic surgery with innovations from computer vision and artificial intelligence: a narrative review
Journal of Cardiothoracic Surgery (2024)
-
NSF DARE—transforming modeling in neurorehabilitation: a patient-in-the-loop framework
Journal of NeuroEngineering and Rehabilitation (2024)
-
Characterization of early markers of disease in the mouse model of mucopolysaccharidosis IIIB
Journal of Neurodevelopmental Disorders (2024)
-
A systematic review of instrumented assessments for upper limb function in cerebral palsy: current limitations and future directions
Journal of NeuroEngineering and Rehabilitation (2024)