Abstract
Lateral hypothalamus (LH) neurons containing the neuropeptide hypocretin (HCRT; orexin) modulate affective components of arousal, but their relevant synaptic inputs remain poorly defined. Here we identified inputs onto LH neurons that originate from neuronal populations in the bed nuclei of stria terminalis (BNST; a heterogeneous region of extended amygdala). We characterized two non-overlapping LH-projecting GABAergic BNST subpopulations that express distinct neuropeptides (corticotropin-releasing factor, CRF, and cholecystokinin, CCK). To functionally interrogate BNST→LH circuitry, we used tools for monitoring and manipulating neural activity with cell-type-specific resolution in freely behaving mice. We found that Crf-BNST and Cck-BNST neurons respectively provide abundant and sparse inputs onto Hcrt-LH neurons, display discrete physiological responses to salient stimuli, drive opposite emotionally valenced behaviors, and receive different proportions of inputs from upstream networks. Together, our data provide an advanced model for how parallel BNST→LH pathways promote divergent emotional states via connectivity patterns of genetically defined, circuit-specific neuronal subpopulations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jennings, J. H., Rizzi, G., Stamatakis, A. M., Ung, R. L. & Stuber, G. D. The inhibitory circuit architecture of the lateral hypothalamus orchestrates feeding. Science 341, 1517–1521 (2013).
Nieh, E. H. et al. Decoding neural circuits that control compulsive sucrose seeking. Cell 160, 528–541 (2015).
de Lecea, L. et al. The hypocretins: hypothalamus-specific peptides with neuroexcitatory activity. Proc. Natl. Acad. Sci. USA 95, 322–327 (1998).
Sakurai, T. et al. Orexins and orexin receptors: a family of hypothalamic neuropeptides and G protein-coupled receptors that regulate feeding behavior. Cell 92, 573–585 (1998).
Baimel, C. et al. Orexin/hypocretin role in reward: implications for opioid and other addictions. Br. J. Pharmacol. 172, 334–348 (2015).
Giardino, W. J. & de Lecea, L. Hypocretin (orexin) neuromodulation of stress and reward pathways. Curr. Opin. Neurobiol. 29, 103–108 (2014).
Li, S. B., Giardino, W. J. & de Lecea, L. Hypocretins and arousal. Curr. Top. Behav. Neurosci. 33, 93–104 (2017).
Bourgin, P. et al. Hypocretin-1 modulates rapid eye movement sleep through activation of locus coeruleus neurons. J. Neurosci. 20, 7760–7765 (2000).
Peyron, C. et al. Neurons containing hypocretin (orexin) project to multiple neuronal systems. J. Neurosci. 18, 9996–10015 (1998).
Boutrel, B. et al. Role for hypocretin in mediating stress-induced reinstatement of cocaine-seeking behavior. Proc. Natl. Acad. Sci. USA 102, 19168–19173 (2005).
Hata, T. et al. Intra-ventral tegmental area or intracerebroventricular orexin-A increases the intra-cranial self-stimulation threshold via activation of the corticotropin-releasing factor system in rats. Eur. J. Neurosci. 34, 816–826 (2011).
Borgland, S. L. et al. Orexin A/hypocretin-1 selectively promotes motivation for positive reinforcers. J. Neurosci. 29, 11215–11225 (2009).
Muschamp, J. W. et al. Hypocretin (orexin) facilitates reward by attenuating the antireward effects of its cotransmitter dynorphin in ventral tegmental area. Proc. Natl. Acad. Sci. USA 111, E1648–E1655 (2014).
González, J. A., Iordanidou, P., Strom, M., Adamantidis, A. & Burdakov, D. Awake dynamics and brain-wide direct inputs of hypothalamic MCH and orexin networks. Nat. Commun. 7, 11395 (2016).
Sakurai, T. et al. Input of orexin/hypocretin neurons revealed by a genetically encoded tracer in mice. Neuron 46, 297–308 (2005).
Tsujino, N. et al. Cholecystokinin activates orexin/hypocretin neurons through the cholecystokinin A receptor. J. Neurosci. 25, 7459–7469 (2005).
Winsky-Sommerer, R. et al. Interaction between the corticotropin-releasing factor system and hypocretins (orexins): a novel circuit mediating stress response. J. Neurosci. 24, 11439–11448 (2004).
Kim, S. Y. et al. Diverging neural pathways assemble a behavioural state from separable features in anxiety. Nature 496, 219–223 (2013).
Lebow, M. A. & Chen, A. Overshadowed by the amygdala: the bed nucleus of the stria terminalis emerges as key to psychiatric disorders. Mol. Psychiatry 21, 450–463 (2016).
Jennings, J. H. et al. Distinct extended amygdala circuits for divergent motivational states. Nature 496, 224–228 (2013).
Marcinkiewcz, C. A. et al. Serotonin engages an anxiety and fear-promoting circuit in the extended amygdala. Nature 537, 97–101 (2016).
Gunaydin, L. A. et al. Natural neural projection dynamics underlying social behavior. Cell 157, 1535–1551 (2014).
Zhang, F., Aravanis, A. M., Adamantidis, A., de Lecea, L. & Deisseroth, K. Circuit-breakers: optical technologies for probing neural signals and systems. Nat. Rev. Neurosci. 8, 577–581 (2007).
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc. Natl. Acad. Sci. USA 104, 5163–5168 (2007).
Wall, N. R., Wickersham, I. R., Cetin, A., De La Parra, M. & Callaway, E. M. Monosynaptic circuit tracing in vivo through Cre-dependent targeting and complementation of modified rabies virus. Proc. Natl. Acad. Sci. USA 107, 21848–21853 (2010).
Eban-Rothschild, A., Rothschild, G., Giardino, W. J., Jones, J. R. & de Lecea, L. VTA dopaminergic neurons regulate ethologically relevant sleep-wake behaviors. Nat. Neurosci. 19, 1356–1366 (2016).
Bonnavion, P., Jackson, A. C., Carter, M. E. & de Lecea, L. Antagonistic interplay between hypocretin and leptin in the lateral hypothalamus regulates stress responses. Nat. Commun. 6, 6266 (2015).
Winrow, C. J. et al. Pharmacological characterization of MK-6096 - a dual orexin receptor antagonist for insomnia. Neuropharmacology 62, 978–987 (2012).
Li, S.B., Nevárez, N., Giardino, W.J. & de Lecea, L. Optical Probing of Orexin/Hypocretin Receptor Antagonists. Sleep (Basel) (in the press).
Crestani, C. C. et al. Mechanisms in the bed nucleus of the stria terminalis involved in control of autonomic and neuroendocrine functions: a review. Curr. Neuropharmacol. 11, 141–159 (2013).
Ju, G. & Swanson, L. W. Studies on the cellular architecture of the bed nuclei of the stria terminalis in the rat: I. Cytoarchitecture. J. Comp. Neurol. 280, 587–602 (1989).
Krettek, J. E. & Price, J. L. Amygdaloid projections to subcortical structures within the basal forebrain and brainstem in the rat and cat. J. Comp. Neurol. 178, 225–254 (1978).
Radley, J. J., Gosselink, K. L. & Sawchenko, P. E. A discrete GABAergic relay mediates medial prefrontal cortical inhibition of the neuroendocrine stress response. J. Neurosci. 29, 7330–7340 (2009).
Weller, K. L. & Smith, D. A. Afferent connections to the bed nucleus of the stria terminalis. Brain Res. 232, 255–270 (1982).
Dong, H. W. & Swanson, L. W. Projections from bed nuclei of the stria terminalis, posterior division: implications for cerebral hemisphere regulation of defensive and reproductive behaviors. J. Comp. Neurol. 471, 396–433 (2004).
Pleil, K. E. et al. NPY signaling inhibits extended amygdala CRF neurons to suppress binge alcohol drinking. Nat. Neurosci. 18, 545–552 (2015).
McHenry, J. A. et al. Hormonal gain control of a medial preoptic area social reward circuit. Nat. Neurosci. 20, 449–458 (2017).
Füzesi, T., Daviu, N., Wamsteeker Cusulin, J. I., Bonin, R. P. & Bains, J. S. Hypothalamic CRH neurons orchestrate complex behaviours after stress. Nat. Commun. 7, 11937 (2016).
Mazzone, C. M. et al. Acute engagement of Gq-mediated signaling in the bed nucleus of the stria terminalis induces anxiety-like behavior. Mol. Psychiatry 23, 143–153 (2018).
Lin, C. W. et al. Genetically increased cell-intrinsic excitability enhances neuronal integration into adult brain circuits. Neuron 65, 32–39 (2010).
Lin, L. et al. The sleep disorder canine narcolepsy is caused by a mutation in the hypocretin (orexin) receptor 2 gene. Cell 98, 365–376 (1999).
Eban-Rothschild, A., Giardino, W. J. & de Lecea, L. To sleep or not to sleep: neuronal and ecological insights. Curr. Opin. Neurobiol. 44, 132–138 (2017).
González, J. A. et al. Inhibitory interplay between orexin neurons and eating. Curr. Biol. 26, 2486–2491 (2016).
Baimel, C., Lau, B. K., Qiao, M. & Borgland, S. L. Projection-target-defined effects of orexin and dynorphin on VTA dopamine neurons. Cell Rep. 18, 1346–1355 (2017).
Leinninger, G. M. et al. Leptin acts via leptin receptor-expressing lateral hypothalamic neurons to modulate the mesolimbic dopamine system and suppress feeding. Cell Metab. 10, 89–98 (2009).
Nguyen, A. Q., Dela Cruz, J. A., Sun, Y., Holmes, T. C. & Xu, X. Genetic cell targeting uncovers specific neuronal types and distinct subregions in the bed nucleus of the stria terminalis. J. Comp. Neurol. 524, 2379–2399 (2016).
Poulin, J. F., Arbour, D., Laforest, S. & Drolet, G. Neuroanatomical characterization of endogenous opioids in the bed nucleus of the stria terminalis. Prog. Neuropsychopharmacol. Biol. Psychiatry 33, 1356–1365 (2009).
McCarthy, E. A. et al. DREADD-induced silencing of the medial amygdala reduces the preference for male pheromones and the expression of lordosis in estrous female mice. Eur. J. Neurosci. 46, 2035–2046 (2017).
Shemesh, Y. et al. Ucn3 and CRF-R2 in the medial amygdala regulate complex social dynamics. Nat. Neurosci. 19, 1489–1496 (2016).
Unger, E. K. et al. Medial amygdalar aromatase neurons regulate aggression in both sexes. Cell Rep. 10, 453–462 (2015).
Bender, D., Holschbach, M. & Stöcklin, G. Synthesis of n.c.a. carbon-11 labelled clozapine and its major metabolite clozapine-N-oxide and comparison of their biodistribution in mice. Nucl. Med. Biol. 21, 921–925 (1994).
Guettier, J. M. et al. A chemical-genetic approach to study G protein regulation of beta cell function in vivo. Proc. Natl. Acad. Sci. USA 106, 19197–19202 (2009).
Giardino, W. J. et al. Control of chronic excessive alcohol drinking by genetic manipulation of the Edinger-Westphal nucleus urocortin-1 neuropeptide system. Transl. Psychiatry 7, e1021 (2017).
Giardino, W. J. & Ryabinin, A. E. CRF1 receptor signaling regulates food and fluid intake in the drinking-in-the-dark model of binge alcohol consumption. Alcohol. Clin. Exp. Res. 37, 1161–1170 (2013).
Acknowledgements
We acknowledge all de Lecea lab members, A.D. Abraham, and P.F. Apostolides for critical feedback. We thank A. Khan, T.A. Lucas, K. Malacon, M. Silvestre, D. Hoang, C. Liang, K. Choudhury, K. Cruz, and A. Yao for excellent technical assistance. This work was supported by National Institutes of Health grants F32 AA022832 (W.J.G.), R01 MH087592 (L.d.L.), R01 MH102638 (L.d.L.), and F32 MH106206 (D.J.C.). We also recognize support from A. Olson and the Stanford Neuroscience Microscopy Service, NIH NS069375.
Author information
Authors and Affiliations
Contributions
W.J.G. and L.d.L. conceived and designed the studies. W.J.G., A.E.-R., D.J.C., and S.-B. L. performed experiments. W.J.G. and D.J.C. analyzed data. R.C.M. provided equipment and resources, W.J.G. wrote the manuscript with contributions from A.E.-R., D.J.C., and L.d.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Hcrt-IRES-Cre validation and Hcrt-ChR2-LH RTPT behavioral and pharmacological analyses.
(a) Schematic of targeting vector for Hcrt-IRES-Cre knockin mouse line. (b) Knockin strategy for Hcrt-IRES-Cre. (c) Representative images showing co-expression of AAV-DIO-eYFP viral labeling (green) and Hcrt (top) or melanin-concentrating hormone (MCH; bottom) immunostaining (red) in LH of Hcrt-Cre mice, replicated independently with similar results in four mice. (d) Quantification of specificity and efficiency of Hcrt-LH infection with Cre-inducible viruses in Hcrt-Cre mice, n=4. (e) Raw Ca2+ activity traces from in vivo fiber photometry recordings of Hcrt-LH neurons in Hcrt-Cre mice injected with AAV-DIO-GCaMP6f virus. Traces show 60s baseline (left of dashed line), and 60s of Ca2+ activity data following exposure to salient odorant stimuli (right of dashed line) (f) Mean (± S.E.M.) fluorescent Ca2+ activity traces from in vivo fiber photometry recordings of Hcrt-LH neurons in control mice injected with AAV-DIO-GFP virus during baseline (left of dashed line), and following exposure to salient odorant stimuli (right of dashed line). Inset; mean (± S.E.M.) fluorescence levels during the 60s stimuli exposure phase (n=2 mice; 1-2 stimulus trials per subject). (g) Number of entries into the stimulation side for Hcrt-LH-ChR2 and control mice tested in the RTPT (n=7 eYFP, n=10 ChR2-eYFP, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,30 = 1.37, p = 0.27). Centre and error bars are mean ± S.E.M. (h) Mean time spent per entry to the stimulation side for Hcrt-LH-ChR2 and eYFP control mice tested in the RTPT (n=7 eYFP, n=10 ChR2-eYFP mice, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,30 = 2.28, p = 0.12, Bonferroni post-hoc comparison **p < 0.005 vs. Hcrt-LH-eYFP). Centre and error bars are mean ± S.E.M. (i) Hcrt-LH-ChR2 real-time place avoidance is significantly blocked by pharmacological treatment with double Hcrt-R antagonist MK-6096 (5mg/kg i.p., n=10 mice, two-tailed paired t-test; t9= 1.11, p = 0.297, n.s.). Centre and error bars are mean ± S.E.M. (j) Significant Hcrt-LH-ChR2 real-time place avoidance following pharmacological treatment with KOR antagonist norBNI (10 mg/kg i.p., n=6 mice, two-tailed paired t-test; t5 = 4.33, p = 0.0075, **p < 0.01 vs. No Light control). Centre and error bars are mean ± S.E.M. Scalebars: all 100 μm.
Supplementary Figure 2 LepRb-LH: GCaMP6 expression, RTPT behavioral analyses, and rabies mapping.
(a) Representative image of LH-AAV-DIO-GCaMP6 expression and fiberoptic placement in LepRb-Cre x LSL-Ai14 Cre-inducible tdTomato mouse. Merged image shows specific co-expression of viral DIO-GCaMP6f labeling (green) with LepRb-tdTomato reporter expression (red), replicated independently with similar results in three mice. (b) Number of entries into the stimulation side for LepRb-LH-ChR2 and eYFP control mice tested in the RTPT (n=8 eYFP, n=6 ChR2-eYFP, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,24 = 0.55, p = 0.58). Centre and error bars are mean ± S.E.M.). (c) Mean time spent per entry to the stimulation side for LepRb-LH-ChR2 and eYFP control mice tested in the RTPT (n=8 eYFP, n=6 ChR2-eYFP, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,24 = 4.47, p = 0.0224, Bonferroni post-hoc comparison *p < 0.05 ***p < 0.0005 vs. LepRb-LH-eYFP). Centre and error bars are mean ± S.E.M. (d) Representative image showing site of viral injection for Cre-defined rabies input mapping onto LH neurons of LepRb-Cre mice, replicated independently with similar results in three mice. (e) Representative images showing upstream inputs cells throughout the brain in LepRb-LH-RVdG-GFP mice, replicated independently with similar results in three mice. Scalebars: all 100 μm.
Supplementary Figure 3 Crf and Cck: non-overlapping BNST subpopulations, differential inputs onto LH neurons.
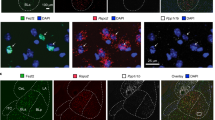
(a) Genetic screen of Cre-inducible Ai14 (tdTomato) reporter expression in adBNST neurons of wild-type mice, and mice from seven different BNST marker Cre driver lines, replicated independently with similar results in three mice. (b-c) Representative images of CRF and CCK immunostaining (red) in adBNST and pBNST and slices from LepRb-LH-RVdG-GFP mice, replicated independently with similar results in three mice. White arrowheads indicate neuropeptide co-expressing input cells. (d) Representative co-expression between Crf-Cre, Cck-Cre, Vgat-Cre, or Vglut2-Cre Ai14 (tdTomato, red) reporter expression and CRF or CCK (green) immunostaining in adBNST neurons, replicated independently with similar results in three mice. (e) Representative co-expression between CRF immunostaining & Drd1-Ai14 (top), and Tac1-Ai14 (bottom) in adBNST neurons, replicated independently with similar results in three mice. Scalebars: all 100 μm.
Supplementary Figure 4 Optogenetic stimulation of additional BNST and hypothalamus subpopulations.
(a) Representative image of AAV-DIO-eYFP expression in adlBNST and admBNST of Vgat-Cre mouse. (b) Vgat-BNST-ChR2 optogenetic stimulation is rewarding in the RTPT (n=4 eYFP, n=5 ChR2-eYFP, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,14 = 4.16, p = 0.038, Bonferroni post-hoc comparisons *p < 0.05, ***p < 0.0001 vs. Vgat-BNST-eYFP control group). Centre and error bars are mean ± S.E.M. (c) Representative image of AAV-DIO-eYFP expression in ovBNST of Drd1-Cre mouse. (d) Drd1-BNST-ChR2 optogenetic stimulation is aversive in the RTPT (n=4–8 mice per group, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,20 = 12.38, p = 0.0003, Bonferroni post-hoc comparisons ***p < 0.005 vs. Drd1-BNST-eYFP control group). Centre and error bars are mean ± S.E.M. (e) Representative image of AAV-DIO-ChR2-eYFP expression in admBNST of Six3-Cre mouse. (f) Six3-BNST-ChR2 optogenetic stimulation is rewarding in the RTPT (n=4 per group, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,12 = 8.79, p = 0.0045, Bonferroni post-hoc comparisons ***p < 0.001 vs. Six3-BNST-eYFP control group). Centre and error bars are mean ± S.E.M. (g) Representative image of AAV-DIO-eYFP expression in PVN of Crf-Cre mice. (h) Crf-PVN-ChR2 optogenetic stimulation is aversive in the RTPT (n=4 eYFP, n=6 ChR2-eYFP mice, two-way RM-ANOVA; ChR2 x stimulation interaction: F2,16 = 2.33, p = 0.13, Bonferroni post-hoc comparisons *p < 0.05 vs. Crf-PVN-eYFP control group). Centre and error bars are mean ± S.E.M. Scalebars: all 100 μm.
Supplementary Figure 5 Crf- and Cck-BNST neurons: anxiety-like behavior.
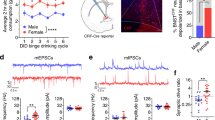
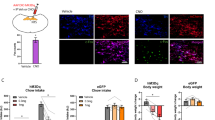
(a) Velocity and (b) percent time spent in the center in the open field test following Crf-BNST DREADD manipulation (n=19 mCherry, n=9 hM3Dq, n=7 hM4Di, two-way RM-ANOVA; DREADD x CNO interaction for velocity: F2,32 = 1.86, p = 0.172, DREADD x CNO interaction for center time: F2,32 = 6.03, p = 0.006, Bonferroni post-hoc comparisons *p < 0.05 vs. saline control). Centre and error bars are mean ± S.E.M. (c) Velocity and (d) percent time spent in the center in the open field test following Cck-BNST DREADD manipulation (n=15 mCherry, n=11 hM3Dq, n=4 hM4Di, two-way RM-ANOVA; DREADD x CNO interaction for velocity: F2,28 = 5.52, p = 0.010, Bonferroni post-hoc comparisons **p < 0.005 vs. saline control, DREADD x CNO interaction for center time: F2,28 = 4.99, p = 0.014, Bonferroni post-hoc comparisons *p < 0.05 vs. saline control). Centre and error bars are mean ± S.E.M. (e) Percent open arm time in the elevated plus maze (EPM) following Crf-BNST DREADD manipulation (n=21 mCherry, n=21 hM3Dq, n=13 hM4Di, two-way RM-ANOVA; DREADD x CNO interaction: F2,52 = 14.51, p < 0.0001, Bonferroni post-hoc comparisons ***p < 0.0001 vs. saline control). Centre and error bars are mean ± S.E.M. (f) Crf-BNST-ChR2 10Hz optogenetic stimulation increased anxiety-like behavior (n=11 mice, two-tailed paired t-test; t10 = 4.16, **p = 0.0020 vs. no-stimulation control). Centre and error bars are mean ± S.E.M. (g) Percent open arm time in the EPM following Cck-BNST DREADD manipulation (n=16 mCherry, n=11 hM3Dq, n=10 hM4Di, two-way RM-ANOVA; DREADD x CNO interaction: F2,34 = 9.431, p = 0.0006, Bonferroni post-hoc comparisons ***p < 0.0001 vs. saline control. Centre and error bars are mean ± S.E.M. (h) Cck-BNST-ChR2 10Hz optogenetic stimulation increased anxiety-like behavior (n=12 mice, two-tailed paired t-test; t11 = 5.53, ***p < 0.0005 vs. no-stimulation control). Centre and error bars are mean ± S.E.M.
Supplementary Figure 6 Crf- and Cck-BNST innervation of Hcrt-eGFP+ LH neurons.
(a) Representative images of BNST-AAV-EF1a-DIO-ChR2-mCherry axonal fibers innervating the Hcrt-LH field in Crf-Cre (left) and Cck-Cre (right) mice crossed with Hcrt-eGFP reporter mice, replicated independently with similar results in six mice per group. Scalebars: 100 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6
Rights and permissions
About this article
Cite this article
Giardino, W.J., Eban-Rothschild, A., Christoffel, D.J. et al. Parallel circuits from the bed nuclei of stria terminalis to the lateral hypothalamus drive opposing emotional states. Nat Neurosci 21, 1084–1095 (2018). https://doi.org/10.1038/s41593-018-0198-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0198-x
This article is cited by
-
Chemogenetic activation of corticotropin-releasing factor-expressing neurons in the anterior bed nucleus of the stria terminalis reduces effortful motivation behaviors
Neuropsychopharmacology (2024)
-
Role of corticotropin-releasing factor neurotransmission in the lateral hypothalamus on baroreflex impairment evoked by chronic variable stress in rats
Pflügers Archiv - European Journal of Physiology (2024)
-
CaMKIIa+ neurons in the bed nucleus of the stria terminalis modulate pace of natural reward seeking depending on internal state
Psychopharmacology (2024)
-
Optogenetic and pharmacological interventions link hypocretin neurons to impulsivity in mice
Communications Biology (2023)
-
Rewarding-unrewarding prediction signals under a bivalent context in the primate lateral hypothalamus
Scientific Reports (2023)