Abstract
A localized transcriptome at the synapse facilitates synapse-, stimulus- and transcript-specific local protein synthesis in response to neuronal activity. While enzyme-mediated mRNA modifications are known to regulate cellular mRNA turnover, the role of these modifications in regulating synaptic RNA has not been studied. We established low-input m6A-sequencing of synaptosomal RNA to determine the chemically modified local transcriptome in healthy adult mouse forebrains and identified 4,469 selectively enriched m6A sites in 2,921 genes as the synaptic m6A epitranscriptome (SME). The SME is functionally enriched in synthesis and modulation of tripartite synapses and in pathways implicated in neurodevelopmental and neuropsychiatric diseases. Interrupting m6A-mediated regulation via knockdown of readers in hippocampal neurons altered expression of SME member Apc, resulting in synaptic dysfunction including immature spine morphology and dampened excitatory synaptic transmission concomitant with decreased clusters of postsynaptic density-95 (PSD-95) and decreased surface expression of AMPA receptor subunit GluA1. Our findings indicate that chemical modifications of synaptic mRNAs critically contribute to synaptic function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
10 August 2018
In the version of this article initially published, a Supplementary Fig. 6f was cited in the last paragraph of the Results. No such panel exists; the citation has been deleted. The error has been corrected in the HTML and PDF versions of the article.
References
Kelsch, W., Lin, C.-W. & Lois, C. Sequential development of synapses in dendritic domains during adult neurogenesis. Proc. Natl. Acad. Sci. USA 105, 16803–16808 (2008).
Holtmaat, A. & Svoboda, K. Experience-dependent structural synaptic plasticity in the mammalian brain. Nat. Rev. Neurosci. 10, 647–658 (2009).
Citri, A. & Malenka, R. C. Synaptic plasticity: multiple forms, functions, and mechanisms. Neuropsychopharmacology 33, 18–41 (2008).
Sutton, M. A. & Schuman, E. M. Dendritic protein synthesis, synaptic plasticity, and memory. Cell 127, 49–58 (2006).
Kelleher, R. J. III, Govindarajan, A. & Tonegawa, S. Translational regulatory mechanisms in persistent forms of synaptic plasticity. Neuron 44, 59–73 (2004).
Bassell, G. J. & Warren, S. T. Fragile X syndrome: loss of local mRNA regulation alters synaptic development and function. Neuron 60, 201–214 (2008).
Kelleher, R. J. III & Bear, M. F. The autistic neuron: troubled translation? Cell 135, 401–406 (2008).
Santini, E. et al. Exaggerated translation causes synaptic and behavioural aberrations associated with autism. Nature 493, 411–415 (2013).
Cajigas, I. J. et al. The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 74, 453–466 (2012).
Perea, G., Navarrete, M. & Araque, A. Tripartite synapses: astrocytes process and control synaptic information. Trends Neurosci. 32, 421–431 (2009).
Eroglu, C. & Barres, B. A. Regulation of synaptic connectivity by glia. Nature 468, 223–231 (2010).
Sakers, K. et al. Astrocytes locallytranslate transcripts in their peripheral processes. Proc. Natl. Acad. Sci. USA 114, E3830–E3838 (2017).
Kiebler, M. A. & Bassell, G. J. Neuronal RNA granules: movers and makers. Neuron 51, 685–690 (2006).
Bramham, C. R. & Wells, D. G. Dendritic mRNA: transport, translation and function. Nat. Rev. Neurosci. 8, 776–789 (2007).
Martin, K. C. & Ephrussi, A. mRNA localization: gene expression in the spatial dimension. Cell 136, 719–730 (2009).
Wang, D. O., Martin, K. C. & Zukin, R. S. Spatially restricting gene expression by local translation at synapses. Trends Neurosci. 33, 173–182 (2010).
Fernandez-Moya, S. M., Bauer, K. E. & Kiebler, M. A. Meet the players: local translation at the synapse. Front. Mol. Neurosci. 7, 84 (2014).
Gilbert, W. V., Bell, T. A. & Schaening, C. Messenger RNA modifications: Form, distribution, and function. Science 352, 1408–1412 (2016).
Nainar, S., Marshall, P. R., Tyler, C. R., Spitale, R. C. & Bredy, T. W. Evolving insights into RNA modifications and their functional diversity in the brain. Nat. Neurosci. 19, 1292–1298 (2016).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Wang, Y. & Zhao, J. C. Update: mechanisms underlying N 6-methyladenosine modification of eukaryotic mRNA. Trends Genet. 32, 763–773 (2016).
Liu, N. et al. N 6-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Yoon, K. J. et al. Temporal control of mammalian cortical neurogenesis by m6A methylation. Cell 171, 877–889.e17 (2017).
Wang, Y. et al. N 6-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nat. Neurosci. 21, 195–206 (2018).
Lence, T. et al. m6A modulates neuronal functions and sex determination in Drosophila. Nature 540, 242–247 (2016).
Fustin, J.-M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013).
Hess, M. E. et al. The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry. Nat. Neurosci. 16, 1042–1048 (2013).
Widagdo, J. et al. Experience-dependent accumulation of N 6-methyladenosine in the prefrontal cortex is associated with memory processes in mice. J. Neurosci. 36, 6771–6777 (2016).
Walters, B. J. et al. The role of the RNA demethylase FTO (Fat Mass and Obesity-Associated) and mRNA methylation in hippocampal memory formation. Neuropsychopharmacology 42, 1502–1510 (2017).
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N 6-methyladenosine by m6A-seq based on immunocapturing and massively parallel sequencing. Nat. Protoc. 8, 176–189 (2013).
Ke, S. et al. A majority of m6A residues are in the last exons, allowing the potential for 3′ UTR regulation. Genes Dev. 29, 2037–2053 (2015).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep. 8, 284–296 (2014).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Wang, X. et al. N 6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Loh, K. H. et al. Proteomic analysis of unbounded cellular compartments: synaptic clefts. Cell 166, 1295–1307.e21 (2016).
Xiang, Y. et al. RNA m6A methylation regulates the ultraviolet-induced DNA damage response. Nature 543, 573–576 (2017).
Meyer, K. D. et al. 5′ UTR m6A promotes cap-independent translation. Cell 163, 999–1010 (2015).
Zhou, J. et al. Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Choi, J. et al. N 6-Methyladenosine in mRNA disrupts tRNA selection and translation-elongation dynamics. Nat. Struct. Mol. Biol. 23, 110–115 (2016).
Du, H. et al. YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat. Commun. 7, 12626 (2016).
Wang, X. et al. N 6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Shi, H. et al. YTHDF3 facilitates translation and decay of N 6-methyladenosine-modified RNA. Cell Res. 27, 315–328 (2017).
Li, A. et al. Cytoplasmic m6A reader YTHDF3 promotes mRNA translation. Cell Res. 27, 444–447 (2017).
Mohn, J. L. et al. Adenomatous polyposis coli protein deletion leads to cognitive and autism-like disabilities. Mol. Psychiatry 19, 1133–1142 (2014).
Liu, X. et al. Genome-wide association study of autism spectrum disorder in the East Asian populations. Autism Res. 9, 340–349 (2016).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Hussain, S. & Bashir, Z. I. The epitranscriptome in modulating spatiotemporal RNA translation in neuronal post-synaptic function. Front. Cell. Neurosci. 9, 420 (2015).
Abbasi-Moheb, L. et al. Mutations in NSUN2 cause autosomal-recessive intellectual disability. Am. J. Hum. Genet. 90, 847–855 (2012).
Dunkley, P. R., Jarvie, P. E. & Robinson, P. J. A rapid Percoll gradient procedure for preparation of synaptosomes. Nat. Protoc. 3, 1718–1728 (2008).
Westmark, P. R., Westmark, C. J., Jeevananthan, A. & Malter, J. S. Preparation of synaptoneurosomes from mouse cortex using a discontinuous Percoll-sucrose density gradient. J. Vis. Exp. 55, 1–9 (2011).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 12, 323 (2011).
Leng, N. et al. EBSeq: an empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29, 1035–1043 (2013).
Meng, J. et al. A protocol for RNA methylation differential analysis with MeRIP-Seq data and exomePeak R/Bioconductor package. Methods 69, 274–281 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Cui, X. et al. Guitar: An R/Bioconductor package for gene annotation guided transcriptomic analysis of RNA-related genomic features. Biomed. Res. Int. 2016, 8367534 (2016).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Tripathi, S. et al. Meta- and orthogonal integration of influenza “omics” data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44 (W1), W90–W97 (2016).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Kaech, S. & Banker, G. Culturing hippocampal neurons. Nat. Protoc. 1, 2406–2415 (2006).
Kanda, Y. Investigation of the freely available easy-to-use software ‘EZR’ for medical statistics. Bone Marrow Transplant 48, 452–458 (2013).
Hayashi, N., Oohira, A. & Miyata, S. Synaptic localization of receptor-type protein tyrosine phosphatase ζ/β in the cerebral and hippocampal neurons of adult rats. Brain Res. 1050, 163–169 (2005).
Buxbaum, J. D. et al. Molecular dissection of NRG1-ERBB4 signaling implicates PTPRZ1 as a potential schizophrenia susceptibility gene. Mol. Psychiatry 13, 162–172 (2008).
Takahashi, N. et al. Increased expression of receptor phosphotyrosine phosphatase-β/ζ is associated with molecular, cellular, behavioral and cognitive schizophrenia phenotypes. Transl. Psychiatry 1, e8 (2011).
Johnston, I. G., Paladino, T., Gurd, J. W. & Brown, I. R. Molecular cloning of SC1: a putative brain extracellular matrix glycoprotein showing partial similarity to osteonectin/BM40/SPARC. Neuron 4, 165–176 (1990).
Purcell, A. E., Jeon, O. H., Zimmerman, A. W., Blue, M. E. & Pevsner, J. Postmortem brain abnormalities of the glutamate neurotransmitter system in autism. Neurology 57, 1618–1628 (2001).
Zhurov, V. et al. Molecular pathway reconstruction and analysis of disturbed gene expression in depressed individuals who died by suicide. PLoS One 7, e47581 (2012).
Risher, W. C. et al. Astrocytes refine cortical connectivity at dendritic spines. Elife 3, 1–24 (2014).
Singh, S. K. et al. Astrocytes assemble thalamocortical synapses by bridging NRX1α and NL1 via hevin. Cell 164, 183–196 (2016).
Rothstein, J. D. et al. Knockout of glutamate transporters reveals a major role for astroglial transport in excitotoxicity and clearance of glutamate. Neuron 16, 675–686 (1996).
Valtcheva, S. & Venance, L. Astrocytes gate Hebbian synaptic plasticity in the striatum. Nat. Commun. 7, 13845 (2016).
Woolley, C. S. & McEwen, B. S. Estradiol regulates hippocampal dendritic spine density via an N-methyl-d-aspartate receptor-dependent mechanism. J. Neurosci. 14, 7680–7687 (1994).
Brinton, R. D. et al. Progesterone receptors: form and function in brain. Front. Neuroendocrinol. 29, 313–339 (2008).
Schumacher, M., Sitruk-Ware, R. & De Nicola, A. F. Progesterone and progestins: neuroprotection and myelin repair. Curr. Opin. Pharmacol. 8, 740–746 (2008).
Rosenberg, M. M. et al. Adenomatous polyposis coli plays a key role, in vivo, in coordinating assembly of the neuronal nicotinic postsynaptic complex. Mol. Cell. Neurosci. 38, 138–152 (2008).
Preitner, N. et al. APC is an RNA-binding protein, and its interactome provides a link to neural development and microtubule assembly. Cell 158, 368–382 (2014).
Raedle, J. et al. A de novo deletion of chromosome 5q causing familial adenomatous polyposis, dysmorphic features, and mild mental retardation. Am. J. Gastroenterol. 96, 3016–3020 (2001).
Zhou, X.-L. et al. Association of adenomatous polyposis coli (APC) gene polymorphisms with autism spectrum disorder (ASD). Am. J. Med. Genet. B. Neuropsychiatr. Genet. 144B, 351–354 (2007).
Yamanaka, H. et al. Expression of Apc2 during mouse development. Brain Res. Gene Expr. Patterns 1, 107–114 (2002).
Shintani, T. et al. APC2 plays an essential role in axonal projections through the regulation of microtubule stability. J. Neurosci. 29, 11628–11640 (2009).
Shintani, T., Takeuchi, Y., Fujikawa, A. & Noda, M. Directional neuronal migration is impaired in mice lacking adenomatous polyposis coli 2. J. Neurosci. 32, 6468–6484 (2012).
Almuriekhi, M. et al. Loss-of-function mutation in APC2 causes Sotos syndrome features. Cell Rep. 10, 1585–1598 (2015).
Richter, K. et al. Presynaptic cytomatrix protein bassoon is localized at both excitatory and inhibitory synapses of rat brain. J. Comp. Neurol. 448, 437–448 (1999).
Garner, C. C., Kindler, S. & Gundelfinger, E. D. Molecular determinants of presynaptic active zones. Curr. Opin. Neurobiol. 10, 321–327 (2000).
Goldstein, A. Y., Wang, X. & Schwarz, T. L. Axonal transport and the delivery of pre-synaptic components. Curr. Opin. Neurobiol. 18, 495–503 (2008).
Zhai, R. G. et al. Assembling the presynaptic active zone: a characterization of an active one precursor vesicle. Neuron 29, 131–143 (2001).
Dick, O. et al. The presynaptic active zone protein bassoon is essential for photoreceptor ribbon synapse formation in the retina. Neuron 37, 775–786 (2003).
Chun, J. et al. Neurobiology of receptor-mediated lysophospholipid signaling. From the first lysophospholipid receptor to roles in nervous system function and development. Ann. NY Acad. Sci. 905, 110–117 (2000).
Kimura, A. et al. Essential roles of sphingosine 1-phosphate/S1P1 receptor axis in the migration of neural stem cells toward a site of spinal cord injury. Stem Cells 25, 115–124 (2007).
Kappos, L. et al. A placebo-controlled trial of oral fingolimod in relapsing multiple sclerosis. N. Engl. J. Med. 362, 387–401 (2010).
Gatchel, J. R. et al. The insulin-like growth factor pathway is altered in spinocerebellar ataxia type 1 and type 7. Proc. Natl. Acad. Sci. USA 105, 1291–1296 (2008).
Shcheglovitov, A. et al. SHANK3 and IGF1 restore synaptic deficits in neurons from 22q13 deletion syndrome patients. Nature 503, 267–271 (2013).
Otowa, T. et al. Genome-wide and gene-based association studies of anxiety disorders in European and African American samples. PLoS One 9, e112559 (2014).
Kang, B. et al. Evaluation of hepatic-metastasis risk of colorectal cancer upon the protein signature of PI3K/AKT pathway. J. Proteome Res. 7, 3507–3515 (2008).
Morrow, E. M. et al. Identifying autism loci and genes by tracing recent shared ancestry. Science 321, 218–223 (2008).
Hirano, S., Yan, Q. & Suzuki, S. T. Expression of a novel protocadherin, OL-protocadherin, in a subset of functional systems of the developing mouse brain. J. Neurosci. 19, 995–1005 (1999).
Uemura, M., Nakao, S., Suzuki, S. T., Takeichi, M. & Hirano, S. OL-Protocadherin is essential for growth of striatal axons and thalamocortical projections. Nat. Neurosci. 10, 1151–1159 (2007).
Tsai, N. P. et al. Multiple autism-linked genes mediate synapse elimination via proteasomal degradation of a synaptic scaffold PSD-95. Cell 151, 1581–1594 (2012).
Pribiag, H., Peng, H., Shah, W. A., Stellwagen, D. & Carbonetto, S. Dystroglycan mediates homeostatic synaptic plasticity at GABAergic synapses. Proc. Natl. Acad. Sci. USA 111, 6810–6815 (2014).
Früh, S. et al. Neuronal dystroglycan is necessary for formation and maintenance of functional CCK-positive basket cell terminals on pyramidal cells. J. Neurosci. 36, 10296–10313 (2016).
Mitsui, K., Nakajima, D., Ohara, O. & Nakayama, M. Mammalianfat3: a large protein that contains multiple cadherin and EGF-like motifs. Biochem. Biophys. Res. Commun. 290, 1260–1266 (2002).
Deans, M. R. et al. Control of neuronal morphology by the atypical cadherin Fat3. Neuron 71, 820–832 (2011).
Nagae, S., Tanoue, T. & Takeichi, M. Temporal and spatial expression profiles of the Fat3 protein, a giant cadherin molecule, during mouse development. Dev. Dyn. 236, 534–543 (2007).
Chang, B. H., Mukherji, S. & Soderling, T. R. Calcium/calmodulin-dependent protein kinase II inhibitor protein: localization of isoforms in rat brain. Neuroscience 102, 767–777 (2001).
Feng, J. et al. Spinophilin regulates the formation and function of dendritic spines. Proc. Natl. Acad. Sci. USA 97, 9287–9292 (2000).
Law, A. J., Weickert, C. S., Hyde, T. M., Kleinman, J. E. & Harrison, P. J. Reduced spinophilin but not microtubule-associated protein 2 expression in the hippocampal formation in schizophrenia and mood disorders: molecular evidence for a pathology of dendritic spines. Am. J. Psychiatry 161, 1848–1855 (2004).
Acknowledgements
We thank Y. Hayashi for critical reading of this manuscript, CeMI imaging center at iCeMS, and the supporting facility at the medical school of Kyoto University for technical support. This work was supported by grants KAKENHI17H03546, KAKENHI26702038 and KAKENHI26115515, and grants from the Hirose Foundation and Astellas Foundation to D.O.W. I.O and B.J.G are supported by Japan Society for the Promotion of Science fellowships.
Author information
Authors and Affiliations
Contributions
D.O.W. conceived and designed the project. W.T.-H. purified synaptosomes and performed RNA extractions, m6A dot-blotting and data mining. D.O.W. and T.O. constructed libraries; D.M., K.I., B.J.G, H.Y. and M.P. performed bioinformatics analysis; T.O. performed immunostaining; I.O. constructed shRNA and performed KD, FISH and spine analysis in cultured hippocampal neurons. S.-y.K. performed electrophysiology. D.O.W and B.J.G. wrote the manuscript. K.C.M. and T.H. supervised parts of the project. All authors participated in data analysis and interpretation and made indispensable contributions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Cross-database comparison and experimental setup for preparation and characterization of m6A synaptic RNA in synaptosomes.
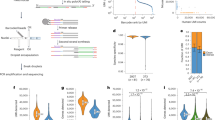
a. Summary of cross-database comparison of m6A mRNA and localized mRNA lists. b. Synaptosome purification from healthy adult mouse forebrains using percoll-sucrose discontinuous gradients. F3 and F4 fractions were pooled as synaptosomal fraction (SYN). Homogenate lysate without fractionation was used as control for comparison (HOM); c. Relative (de)enrichment of a panel of synaptic and somatic markers probed using western blot (n=3 independent experiments); d. Full scan of blots in c (n=3 independent experiments); e. Separation of RNA populations was confirmed by qRT-PCR examination of nuclear lncRNA Malat1, somatic RNA beta-3 tubulin and known synaptically localized transcript CaMKIIα (mean value ±s.e.m., n=3 independent experiments). f. Bioanalyzer analysis of RNA integrity in prepared fractions. HOM, homogenate; Cyto, cytoplasm, F3 and F4 (SYN), synaptosomes (n=3 independent experiments). g. A biological replicate of m6A dot-blot presented in Fig. 1b shows highly similar results (n=2 independent experiments).
Supplementary Figure 2 Validation of low-input m6A-seq and characterization of HOM and SYN peaks.
a. Sequencing summary in this study; b. Pairwise comparison of biological replicate sequencing libraries demonstrates strong, linear correlation between replicates (n=2 independent experiments); c. reads sorted and removed from further analysis; d. Mapping statistics for QC reads mapped with STAR to mm9 refseq.GTF. e. hypergeometric tests on peak overlapping to three previously published databases (n=2 independent experiments); f. Frequency plot of motif per 5nt per peak (y axis) against distance from peak summit (x axis) in HOM; g. Enriched human phenotypes among genes with START codon-associated and STOP codon-associated SYN peaks (ToppGene); h. Left, synapse-enriched peak in Ckap5 transcript; Right, synapse-depleted peak in Lysmd4. Red, IP tracks; blue, INPUT tracks; black, peak location track.
Supplementary Figure 3 Negative impact of methylation on synaptic mRNA stability and synaptic localization.
a. (left) Box plots depicting synaptic concentrations of genes in four groups with increasing methylation level. For each gene, methylation level (ML) was calculated as IP/INPUT reads ratios in each designated RNA region (full length, 5’UTR, CDS, and 3’UTR) and divided into four groups from the least methylated to the most methylated (a; ML < −2, b; −2 <= ML < 0, c; 0 <= ML < 2, d; 2 <= ML). S.TMP: transcripts per million at SYN. (right) Scatter plot and regression model labeled with slope, p-value testing the quality of model fitting, and Pearson’s correlation coefficient r; (n=2 independent experiments showing similar results). b. Analogous plots representing transcript enrichment at synapse compared to whole cell. Y-axis and X-axis represent relative expression values and relative methylation values averaged from two independent experiments. For box plots, genes were classified into one of the four groups as follows: e; rel. ML < −1, f; −1 <= rel. ML < 0, g; 0 <= rel. ML < 1, h; 1 <= rel. ML. Genes with TPM less than 1 were excluded from the analysis. number of data points included in this figure: (a) Total; a: 790, b: 5034, c: 4976, d: 1107, 5’UTR; a: 596, b: 2787, c: 4238, d: 1508, CDS; a: 1540, b: 5129, c: 3550, d: 1192, 3’UTR; a: 1062, b: 3590, c: 4011, d: 1808, (b) Total; e: 133, f: 7861, g: 3126, h: 29, 5’UTR; e: 730, f: 78 3367, g: 3306, h: 691, CDS; e: 216, f: 7311, g: 3193, h: 41. 3’UTR; e: 482, f: 5824, g: 3156, h: 256.
Supplementary Figure 4 m6A regulatory proteins in dendritic processes of dissociated hippocampal neuronal cultures and brain slices.
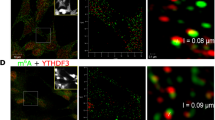
a. Confocal images of m6A regulatory proteins (magenta) and counter-stained with phalloidin (green) to label F actin-rich spines. Scale bar, 5 μm. b. m6A reader YTHDF1 in perfused mouse brain slices. Top, cortical cortex; Bottom, hippocampus and CA1 and CA3 regions of hippocampus. Scale bars, 100 μm and 50 μm. (n= 3 independent experiments).
Supplementary Figure 5 Knocking down METTL3 in dissociated hippocampal neuronal cultures causes cell death and knocking down YTHDF1 causes synapse malfunction.
a. Group quantification of METTL3 protein expression measured by immunofluorescence staining at Day2 after transfection; b. Immunofluorescence images of hippocampal neuronal cultures transfected with shMETTL3 vectors. DAPI (blue), GFP (green), METTL3 (red), (n=3 independent experiments); c. Confocal images of DIV19 dissociated hippocampal neurons expressing shScramble-GFP (left panel) or shYTHDF1-GFP (right panel). DAPI (blue), GFP (green), YTHDF1 (red). Arrowheads point to GFP(+) shScram or shYTHDF1 transfected neurons. Scale bar, 10 μm (n=3 independent experiments); d. Normalized YTHDF1 mRNA expression in shScramble, shYTHDF1-1, and shYTHDF1-2 cells (normalized to beta actin mRNA, n=3 independent experiments); e. Western-blots of YTHDF1 proteins in shScramble, YTHDF1-sh1, and shYTHDF1-2 cells lysates. β-actin was blotted on the same membrane as loading control (n=3 independent experiments). f. Confocal images of dendritic processes of DIV19 dissociated hippocampal neurons expressing shYTHDF1-2-GFP. Top, GFP labels morphology of dendritic shaft and spines; Bottom, PSD-95 staining in the same samples to label post-synaptic density (n=3 independent experiments). g. Group quantification of spine head width in DIV19 dissociated hippocampal neurons expressing shScramble-GFP or shYTHDF1-GFP (n=3 independent experiments). h. Group quantification of PSD-95 intensity in DIV19 dissociated hippocampal neurons expressing shScramble-GFP or shYTHDF1-GFP. i. GFP fluorescence images of control cells (expressing pCAG-EGFP and pX330) and YTHDF1-KD cells (expressing pCAG-EGFP and pX330-guide RNA sequences g1 or g2). Scale bar, 5 μm; j. Group quantifications of spine neck length showed similar phenotypes to YTHDF1-KD using shRNA vectors. k. Group quantification of spine head width showed similar phenotypes to YTHDF1-KD using shRNA vectors (n=2 independent experiments. ***p<1x10-3; Kruskal-Wallis’s multiple comparison test).
Supplementary Figure 6 Reducing YTHDF3 expression in hippocampal neurons causes excessive dendritic filopodia, in place of mature spines, and APC protein expression in YTHDF1-knockdown neurons is reduced.
a. Confocal images of DIV17 dissociated hippocampal neurons expressing shScram-GFP (top) or shYTHDF3-GFP (bottom). DAPI (blue), GFP (green), YTHDF3 (red). Arrowheads point to GFP(+) shScram or shYTHDF3 transfected neurons. Scale bar, 10 μm (n=3 independent experiments). b. Confocal images on dendritic shaft of GFP (green) and YTHDF3 (red). Scale bar, 10 μm (n=3 independent experiments); c. Quantification of YTHDF3 protein expression by immuno-staining in shScram and shYTHDF3 cells (Scram, sh1, and sh2). YTHDF3 protein in sh1 and sh2 samples decreased over a 4-day time course after transfection (n=3 independent experiments). d. Confocal images of dendritic shaft and spines of shScram and shYTHDF3-2 cells (n=3 independent experiments). Left, GFP labels morphology of dendritic shaft and spines; Right, PSD-95 staining in the same sample (magenta). Contour of affected dendrites was traced as the white lines through GFP expression. Fluorescence signals outside of the contour were masked to restrict quantification to the affected neurons. Scale bar, 5 μm (n=3 independent experiments). e. Confocal images of APC immunostaining using APC N-terminus antibody (top) or APC C-terminus antibody (bottom). GFP labels neurons expressing the shRNAs (shScram or shYTHDF1). Both antibodies consistently detected decreased APC protein expression in YTHDF1-KD neurons; Arrowheads point to GFP-positive neurons. DAPI (blue), GFP (green), APC protein (magenta). Scale bar, 10 μm (n=3 independent experiments).
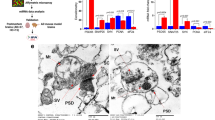
Supplementary Figure 7 Mapping of synaptic cleft proteins and other synaptic proteins encoded by SYN-methylated mRNA and by mRNAs carrying the most abundant methylation sites in SME.
Proteins mapped to excitatory and inhibitory synapses. Dark brown: proteins encoded by top SME genes; light brown: synaptic cleft proteins encoded by synaptically methylated transcripts.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 7 and 8
Supplementary Table 1
Synaptic transcriptome
Supplementary Table 2
HOM peaks and SYN peaks
Supplementary Table 3
Genes associated with peaks around start or stop codon
Supplementary Table 4
GO analysis using expressed genes as background
Supplementary Table 5
Synaptic m6A epitranscriptome (SME)
Supplementary Table 6
qRT-PCR primers
Supplementary Table 9
GO analysis of synaptically hypo- and hypermethylated genes
Supplementary Table 10
Overlap with astrocytic perisynaptic and soma-enriched gene lists
Supplementary Table 11
Values associated with all box plots
Rights and permissions
About this article
Cite this article
Merkurjev, D., Hong, WT., Iida, K. et al. Synaptic N6-methyladenosine (m6A) epitranscriptome reveals functional partitioning of localized transcripts. Nat Neurosci 21, 1004–1014 (2018). https://doi.org/10.1038/s41593-018-0173-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0173-6
This article is cited by
-
FMRP-mediated spatial regulation of physiologic NMD targets in neuronal cells
Genome Biology (2024)
-
Demethylase FTO-Mediated m6A Modification of lncRNA MEG3 Activates Neuronal Pyroptosis via NLRP3 Signaling in Cerebral Ischemic Stroke
Molecular Neurobiology (2024)
-
Captive ERVWE1 triggers impairment of 5-HT neuronal plasticity in the first-episode schizophrenia by post-transcriptional activation of HTR1B in ALKBH5-m6A dependent epigenetic mechanisms
Cell & Bioscience (2023)
-
Examining parent-of-origin effects on transcription and RNA methylation in mediating aggressive behavior in honey bees (Apis mellifera)
BMC Genomics (2023)
-
Comprehensive analysis of N6-methyladenosine-related RNA methylation in the mouse hippocampus after acquired hearing loss
BMC Genomics (2023)