Abstract
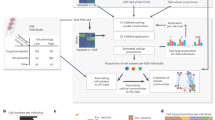
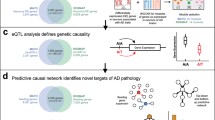
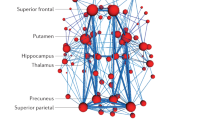
There is a need for new therapeutic targets with which to prevent Alzheimer’s disease (AD), a major contributor to aging-related cognitive decline. Here we report the construction and validation of a molecular network of the aging human frontal cortex. Using RNA sequence data from 478 individuals, we first build a molecular network using modules of coexpressed genes and then relate these modules to AD and its neuropathologic and cognitive endophenotypes. We confirm these associations in two independent AD datasets. We also illustrate the use of the network in prioritizing amyloid- and cognition-associated genes for in vitro validation in human neurons and astrocytes. These analyses based on unique cohorts enable us to resolve the role of distinct cortical modules that have a direct effect on the accumulation of AD pathology from those that have a direct effect on cognitive decline, exemplifying a network approach to complex diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hebert, L. E., Weuve, J., Scherr, P. A. & Evans, D. A. Alzheimer disease in the United States (2010–2050) estimated using the 2010 census. Neurology 80, 1778–1783 (2013).
Cummings, J. L., Morstorf, T. & Zhong, K. Alzheimer’s disease drug-development pipeline: few candidates, frequent failures. Alzheimers Res. Ther. 6, 37 (2014).
Schneider, J. A., Arvanitakis, Z., Leurgans, S. E. & Bennett, D. A. The neuropathology of probable Alzheimer disease and mild cognitive impairment. Ann. Neurol. 66, 200–208 (2009).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Gaiteri, C., Mostafavi, S., Honey, C. J., De Jager, P. L. & Bennett, D. A. Genetic variants in Alzheimer disease - molecular and brain network approaches. Nat. Rev. Neurol. 12, 413–427 (2016).
Mitra, K., Carvunis, A. R., Ramesh, S. K. & Ideker, T. Integrative approaches for finding modular structure in biological networks. Nat. Rev. Genet. 14, 719–732 (2013).
Parikshak, N. N., Gandal, M. J. & Geschwind, D. H. Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat. Rev. Genet. 16, 441–458 (2015).
Gustafsson, M. et al. Modules, networks and systems medicine for understanding disease and aiding diagnosis. Genome Med. 6, 82 (2014).
Geschwind, D. H. & Konopka, G. Neuroscience in the era of functional genomics and systems biology. Nature 461, 908–915 (2009).
Gaiteri, C., Ding, Y., French, B., Tseng, G. C. & Sibille, E. Beyond modules and hubs: the potential of gene coexpression networks for investigating molecular mechanisms of complex brain disorders. Genes Brain Behav. 13, 13–24 (2014).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, 17 (2005).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013).
Miller, J. A., Woltjer, R. L., Goodenbour, J. M., Horvath, S. & Geschwind, D. H. Genes and pathways underlying regional and cell type changes in Alzheimer’s disease. Genome Med. 5, 48 (2013).
Boyle, P. A. et al. Much of late life cognitive decline is not due to common neurodegenerative pathologies. Ann. Neurol. 74, 478–489 (2013).
Jack, C. R. Jr. et al. Age-specific population frequencies of cerebral β-amyloidosis and neurodegeneration among people with normal cognitive function aged 50–89 years: a cross-sectional study. Lancet Neurol. 13, 997–1005 (2014).
Bennett, D. A., Schneider, J. A., Arvanitakis, Z. & Wilson, R. S. Overview and findings from the Religious Orders Study. Curr. Alzheimer Res. 9, 628–645 (2012).
Bennett, D. A. et al. Overview and findings from the Rush Memory and Aging Project. Curr. Alzheimer Res. 9, 646–663 (2012).
Wilson, R. S. et al. Conscientiousness, dementia related pathology, and trajectories of cognitive aging. Psychol. Aging 30, 74–82 (2015).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Gaiteri, C. et al. Identifying robust communities and multi-community nodes by combining top-down and bottom-up approaches to clustering. Sci. Rep. 5, 16361 (2015).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible? PLOS Comput. Biol. 7, e1001057 (2011).
Yan, C. & Boyd, D. D. Histone H3 acetylation and H3 K4 methylation define distinct chromatin regions permissive for transgene expression. Mol. Cell. Biol. 26, 6357–6371 (2006).
Matarin, M. et al. A genome-wide gene-expression analysis and database in transgenic mice during development of amyloid or tau pathology. Cell Rep. 10, 633–644 (2015).
Mancarci, B. O. et al. Cross-laboratory analysis of brain cell type transcriptomes with applications to interpretation of bulk tissue data. eNeuro 4. ENEURO. 0212-17, 2017 (2017).
Koller, D. & Friedman, N. ProbabilisticGraphical Models: Principles and Techniques. (MIT Press, Cambridge, MA, USA, 2009).
Raber, J., Huang, Y. & Ashford, J. W. ApoE genotype accounts for the vast majority of AD risk and AD pathology. Neurobiol. Aging 25, 641–650 (2004).
Yu, L., Boyle, P. A., Leurgans, S., Schneider, J. A. & Bennett, D. A. Disentangling the effects of age and APOE on neuropathology and late life cognitive decline. Neurobiol. Aging 35, 819–826 (2014).
Beecham, G. W. et al. Genome-wide association meta-analysis of neuropathologic features of Alzheimer’s disease and related dementias. PLoS Genet. 10, e1004606 (2014).
Farfel, J. M. et al. Relation of genomic variants for Alzheimer disease dementia to common neuropathologies. Neurology 87, 489–496 (2016).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Lee, P. H., O’Dushlaine, C., Thomas, B. & Purcell, S. M. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics 28, 1797–1799 (2012).
Chibnik, L. B. et al. CR1 is associated with amyloid plaque burden and age-related cognitive decline. Ann. Neurol. 69, 560–569 (2011).
Perälä, N., Sariola, H. & Immonen, T. More than nervous: the emerging roles of plexins. Differentiation 83, 77–91 (2012).
Suwa, A., Kurama, T. & Shimokawa, T. SHIP2 and its involvement in various diseases. Expert Opin. Ther. Targets 14, 727–737 (2010).
Soeda, Y. et al. The inositol phosphatase SHIP2 negatively regulates insulin/IGF-I actions implicated in neuroprotection and memory function in mouse brain. Mol. Endocrinol. 24, 1965–1977 (2010).
Talbot, K. et al. Demonstrated brain insulin resistance in Alzheimer’s disease patients is associated with IGF-1 resistance, IRS-1 dysregulation, and cognitive decline. J. Clin. Invest. 122, 1316–1338 (2012).
Schneider, J. A. et al. Cognitive impairment, decline and fluctuations in older community-dwelling subjects with Lewy bodies. Brain 135, 3005–3014 (2012).
Bennett, D. A. et al. Neuropathology of older persons without cognitive impairment from two community-based studies. Neurology 66, 1837–1844 (2006).
Bennett, D. A., Wilson, R. S., Boyle, P. A., Buchman, A. S. & Schneider, J. A. Relation of neuropathology to cognition in persons without cognitive impairment. Ann. Neurol. 72, 599–609 (2012).
Wilson, R. S. et al. Individual differences in rates of change in cognitive abilities of older persons. Psychol. Aging 17, 179–193 (2002).
Wilson, R., Barnes, L. & Bennett, D. Assessment of lifetime participation in cognitively stimulating activities. J. Clin. Exp. Neuropsychol. 25, 634–642 (2003).
De Jager, P. L. et al. A genome-wide scan for common variants affecting the rate of age-related cognitive decline. Neurobiol. Aging 33, 1017.e1–1017.e15 (2012).
Hyman, B. T. & Trojanowski, J. Q. Consensus recommendations for the postmortem diagnosis of Alzheimer disease from the National Institute on Aging and the Reagan Institute Working Group on diagnostic criteria for the neuropathological assessment of Alzheimer disease. J. Neuropathol. Exp. Neurol. 56, 1095–1097 (1997).
McKhann, G. et al. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 34, 939–944 (1984).
Schneider, J. A., Arvanitakis, Z., Bang, W. & Bennett, D. A. Mixed brain pathologies account for most dementia cases in community-dwelling older persons. Neurology 69, 2197–2204 (2007).
Levin, J. Z. et al. Comprehensive comparative analysis of strand-specific RNA sequencing methods. Nat. Methods 7, 709–715 (2010).
Adiconis, X. et al. Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat. Methods 10, 623–629 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Leek, J. T. & Storey, J. D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007).
Stranger, B. E. et al. Patterns of cis regulatory variation in diverse human populations. PLoS Genet. 8, e1002639 (2012).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLOS Comput. Biol. 6, e1000770 (2010).
Mostafavi, S. et al. Normalizing RNA-sequencing data by modeling hidden covariates with prior knowledge. PLoS One 8, e68141 (2013).
Wu, J., Xiong, H. & Chen, J. in Proc. 15th ACM SIGKDD Intl. Conf. Knowledge Discovery and Data Mining 877–886 (ACM, 2009).
Sugino, K. et al. Molecular taxonomy of major neuronal classes in the adult mouse forebrain. Nat. Neurosci. 9, 99–107 (2006).
Sugino, K. et al. Cell-type-specific repression by methyl-CpG-binding protein 2 is biased toward long genes. J. Neurosci. 34, 12877–12883 (2014).
Doyle, J. P. et al. Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell 135, 749–762 (2008).
Perrone-Bizzozero, N. I., Tanner, D. C., Mounce, J. & Bolognani, F. Increased expression of axogenesis-related genes and mossy fibre length in dentate granule cells from adult HuD overexpressor mice. ASN Neuro. 3, 259–270 (2011).
Beckervordersandforth, R. et al. In vivo fate mapping and expression analysis reveals molecular hallmarks of prospectively isolated adult neural stem cells. Cell Stem Cell 7, 744–758 (2010).
Okaty, B. W., Miller, M. N., Sugino, K., Hempel, C. M. & Nelson, S. B. Transcriptional and electrophysiological maturation of neocortical fast-spiking GABAergic interneurons. J. Neurosci. 29, 7040–7052 (2009).
Maze, I. et al. G9a influences neuronal subtype specification in striatum. Nat. Neurosci. 17, 533–539 (2014).
Heiman, M. et al. Molecular adaptations of striatal spiny projection neurons during levodopa-induced dyskinesia. Proc. Natl. Acad. Sci. USA 111, 4578–4583 (2014).
Tan, C. L. et al. MicroRNA-128 governs neuronal excitability and motor behavior in mice. Science 342, 1254–1258 (2013).
Dalal, J. et al. Translational profiling of hypocretin neurons identifies candidate molecules for sleep regulation. Genes Dev. 27, 565–578 (2013).
Phani, S., Gonye, G. & Iacovitti, L. VTA neurons show a potentially protective transcriptional response to MPTP. Brain Res. 1343, 1–13 (2010).
Arlotta, P. et al. Neuronal subtype-specific genes that control corticospinal motor neuron development in vivo. Neuron 45, 207–221 (2005).
Schmidt, E. F. et al. Identification of the cortical neurons that mediate antidepressant responses. Cell 149, 1152–1163 (2012).
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008).
Zamanian, J. L. et al. Genomic analysis of reactive astrogliosis. J. Neurosci. 32, 6391–6410 (2012).
Fomchenko, E. I. et al. Recruited cells can become transformed and overtake PDGF-induced murine gliomas in vivo during tumor progression. PLoS One 6, e20605 (2011).
Paul, A., Cai, Y., Atwal, G. S. & Huang, Z. J. Developmental coordination of gene expression between synaptic partners during GABAergic circuit assembly in cerebellar cortex. Front. Neural Circuits 6, 37 (2012).
Galloway, J. N. et al. CGG repeats in RNA modulate expression of TDP-43 in mouse and fly models of fragile X tremor ataxia syndrome. Hum. Mol. Genet. 23, 5906–5915 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Tasaki, S. et al. Bayesian network reconstruction using systems genetics data: comparison of MCMC methods. Genetics 199, 973–989 (2015).
Grzegorczyk, M. & Husmeier, D. Improving the structure MCMC sampler for Bayesian networks by introducing a new edge reversal move. Mach. Learn. 71, 265–305 (2008).
Hoeting, J.A., Madigan, D., Raftery, A.E. & Volinsky, C.T. Bayesian model averaging: a tutorial. Stat. Sci. 14, 382–401 (1999).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
We thank the participants of ROS and MAP for their essential contributions and the gifts of their brains to these projects. All subjects gave informed consent. This work has been supported by many different NIH grants: U01AG046152, R01AG036836, P30AG10161, R01AG015819, R01AG017917, R01AG036547. This work was done as part of the National Institute of Aging’s Accelerating Medicines Partnership for AD (AMP-AD).
Author information
Authors and Affiliations
Contributions
D.A.B. and P.L.D.J. designed and funded the study. S.M. and C.G. designed the computational and statistical methods, performed the analysis, and generated the predictions. J.A.S. and D.A.B. collected the biological samples and phenotypic data. S.E.S., J.X., M.T., C.M., R.S., D.E.R., T.L.Y.-P. and P.L.D.J. contributed to the design and execution of data generation with the experimental pipeline. S.M., C.G., C.C.W., S.T., H.-U.K., E.P., V.K., L.Y., L.B.C., D.A.B. and P.L.D.J. contributed to designing and executing the analyses. S.M., C.G., E.M.B., A.R., T.L.Y.-P., D.A.B. and P.L.D.J. reviewed and interpreted results. All authors critically reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Effect of principal components adjustment on gene association with AD.
Figure shows the association strength (negative log10 pvalue) between 31 previously identified AD genes (in columns)(21 from GWAS and the remainder from smaller scale functional studies) and clinical AD. Rows report the results of the association analyses to clinical AD after removing the stated number of principal components (PCs) derived from the RNA-Seq data. The color scale reporting the level of significance is displayed to the right of the figure. This figure demonstrates that removing expression PCs typically does not improve the association strength between gene expression level and AD status.
Supplementary Figure 2 Overlap of association results for our five traits.
For each of the five AD-related traits shown in the figure, a univariate transcriptome-wide association study (TWAS) was conducted. To quantify the proportion of genes whose expression is associated with each pair of AD traits, the following procedure was used: for each trait i, we used the 0.05 FDR threshold to identify genes whose expression associate with trait i. In this case, trait i is the “discovery” sample. Then, we used the pi1 statistic to quantify the proportion of the 0.05 FDR genes that are deemed as true positives for trait j, and so trait j is the “replication” sample. In summary, element (i,j) in this figure represents the pi1 statistic when assessing the 0.05 FDR genes (identified in the discovery sample) from trait i (rows) in the replication sample (trait j)(columns).
Supplementary Figure 3 Coexpression pattern in our cortical RNA-seq data.
This heatmap illustrates the expression patterns of all measured genes (columns), grouped by module, in all analyzed DLPFC samples (rows). Module membership is shown at the bottom of the image. The color scale for relative expression is shown to be the right of the figure.
Supplementary Figure 4 Module preservation and replication.
(A) Module preservation (z-summary, x axis) as assessed in the 4 test datasets. Each row reports the preservation of a module in the 4 datasets. The red dashed line marks the “strongly preserved” threshold and the green dotted line marks the “moderately preserved” threshold (as defined empirically by Langfelder, PLOS Computational Biology 2011). (B) We assessed the replication of the trait to transcriptional measure associations, at the gene-level and module-level (see Supplementary material). To do so, we computed the correlation between vectors of module-to-trait association in our study and in the Zhang et al. study (shown by the red dotted line). We compared the observed correlation between module-trait vectors with gene level associations through resampling. The histogram shows the empirical distribution of correlation coefficient between vectors of gene-to-trait associations in the Zhang study and this study, where we select 47 random genes 10,000 times.
Supplementary Figure 5 Effect of cell population frequency in cortical tissue.
(A) This figure shows the correlation strength (signed, negative log10 p-value: positive denotes a direct correlation) between 7 known gene markers of common brain cell types and the AD-related traits analyzed in this study. As shown, GFAP, which is a marker of astrocytes, is most strongly associated with amyloid load. (B) This figure shows the correlation strength between expression level of all genes with an AD-related trait before (x-axis) and after (y-axis) adjusting for 7 standard cell type markers (shown in Figure S3A).
Supplementary Figure 6 Expression of module 109 meta-feature categorized in order of AD risk, by APOE genotype status.
Due to small sample sizes, genotypes e2/e2 and e2/e4 were collapsed into larger categories. APOE genotype status is defined as: e2 (rs7412-T, rs429358-T), e3 (rs7412-C, rs429358-T), e4 (rs7412-C, rs429358-C). Each dot represents one subject.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 12
Rights and permissions
About this article
Cite this article
Mostafavi, S., Gaiteri, C., Sullivan, S.E. et al. A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease. Nat Neurosci 21, 811–819 (2018). https://doi.org/10.1038/s41593-018-0154-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0154-9