Abstract
How learning enhances neural representations for behaviorally relevant stimuli via activity changes of cortical cell types remains unclear. We simultaneously imaged responses of pyramidal cells (PYR) along with parvalbumin (PV), somatostatin (SOM), and vasoactive intestinal peptide (VIP) inhibitory interneurons in primary visual cortex while mice learned to discriminate visual patterns. Learning increased selectivity for task-relevant stimuli of PYR, PV and SOM subsets but not VIP cells. Strikingly, PV neurons became as selective as PYR cells, and their functional interactions reorganized, leading to the emergence of stimulus-selective PYR–PV ensembles. Conversely, SOM activity became strongly decorrelated from the network, and PYR–SOM coupling before learning predicted selectivity increases in individual PYR cells. Thus, learning differentially shapes the activity and interactions of multiple cell classes: while SOM inhibition may gate selectivity changes, PV interneurons become recruited into stimulus-specific ensembles and provide more selective inhibition as the network becomes better at discriminating behaviorally relevant stimuli.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Recanzone, G. H., Schreiner, C. E. & Merzenich, M. M. Plasticity in the frequency representation of primary auditory cortex following discrimination training in adult owl monkeys. J. Neurosci. 13, 87–103 (1993).
Schoups, A., Vogels, R., Qian, N. & Orban, G. Practising orientation identification improves orientation coding in V1 neurons. Nature 412, 549–553 (2001).
Yang, T. & Maunsell, J. H. The effect of perceptual learning on neuronal responses in monkey visual area V4. J. Neurosci. 24, 1617–1626 (2004).
Yan, Y. et al. Perceptual training continuously refines neuronal population codes in primary visual cortex. Nat. Neurosci. 17, 1380–1387 (2014).
Poort, J. et al. Learning enhances sensory and multiple non-sensory representations in primary visual cortex. Neuron 86, 1478–1490 (2015).
Markram, H. et al. Interneurons of the neocortical inhibitory system. Nat. Rev. Neurosci. 5, 793–807 (2004).
Kepecs, A. & Fishell, G. Interneuron cell types are fit to function. Nature 505, 318–326 (2014).
Letzkus, J. J. et al. A disinhibitory microcircuit for associative fear learning in the auditory cortex. Nature 480, 331–335 (2011).
Donato, F., Rompani, S. B. & Caroni, P. Parvalbumin-expressing basket-cell network plasticity induced by experience regulates adult learning. Nature 504, 272–276 (2013).
Makino, H. & Komiyama, T. Learning enhances the relative impact of top-down processing in the visual cortex. Nat. Neurosci. 18, 1116–1122 (2015).
Chen, S. X., Kim, A. N., Peters, A. J. & Komiyama, T. Subtype-specific plasticity of inhibitory circuits in motor cortex during motor learning. Nat. Neurosci. 18, 1109–1115 (2015).
Kato, H. K., Gillet, S. N. & Isaacson, J. S. Flexible sensory representations in auditory cortex driven by behavioral relevance. Neuron 88, 1027–1039 (2015).
Sachidhanandam, S., Sermet, B. S. & Petersen, C. C. H. Parvalbumin-expressing GABAergic neurons in mouse barrel cortex contribute to gating a goal-directed sensorimotor transformation. Cell Rep. 15, 700–706 (2016).
Fagiolini, M. & Hensch, T. K. Inhibitory threshold for critical-period activation in primary visual cortex. Nature 404, 183–186 (2000).
Froemke, R. C., Merzenich, M. M. & Schreiner, C. E. A synaptic memory trace for cortical receptive field plasticity. Nature 450, 425–429 (2007).
Chen, J. L. et al. Structural basis for the role of inhibition in facilitating adult brain plasticity. Nat. Neurosci. 14, 587–594 (2011).
van Versendaal, D. et al. Elimination of inhibitory synapses is a major component of adult ocular dominance plasticity. Neuron 74, 374–383 (2012).
Kuhlman, S. J. et al. A disinhibitory microcircuit initiates critical-period plasticity in the visual cortex. Nature 501, 543–546 (2013).
Kuchibhotla, K. V. et al. Parallel processing by cortical inhibition enables context-dependent behavior. Nat. Neurosci. 20, 62–71 (2017).
Kvitsiani, D. et al. Distinct behavioural and network correlates of two interneuron types in prefrontal cortex. Nature 498, 363–366 (2013).
Pi, H.-J. et al. Cortical interneurons that specialize in disinhibitory control. Nature 503, 521–524 (2013).
Hangya, B., Pi, H.-J., Kvitsiani, D., Ranade, S. P. & Kepecs, A. From circuit motifs to computations: mapping the behavioral repertoire of cortical interneurons. Curr. Opin. Neurobiol. 26, 117–124 (2014).
Pinto, L. & Dan, Y. Cell-type-specific activity in prefrontal cortex during goal-directed behavior. Neuron 87, 437–450 (2015).
Karnani, M. M. et al. Cooperative subnetworks of molecularly similar interneurons in mouse neocortex. Neuron 90, 86–100 (2016).
Litwin-Kumar, A., Rosenbaum, R. & Doiron, B. Inhibitory stabilization and visual coding in cortical circuits with multiple interneuron subtypes. J. Neurophysiol. 115, 1399–1409 (2016).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Kerlin, A. M., Andermann, M. L., Berezovskii, V. K. & Reid, R. C. Broadly tuned response properties of diverse inhibitory neuron subtypes in mouse visual cortex. Neuron 67, 858–871 (2010).
Keller, A. J. & Martin, K. A. C. Local circuits for contrast normalization and adaptation investigated with two-photon imaging in cat primary visual cortex. J. Neurosci. 35, 10078–10087 (2015).
Wilson, D. E. et al. GABAergic neurons in ferret visual cortex participate in functionally specific networks. Neuron 93, 1058–1065.e4 (2017).
Rudy, B., Fishell, G., Lee, S. & Hjerling-Leffler, J. Three groups of interneurons account for nearly 100% of neocortical GABAergic neurons. Dev. Neurobiol. 71, 45–61 (2011).
Hu, H., Gan, J. & Jonas, P. Fast-spiking, parvalbumin+ GABAergic interneurons: from cellular design to microcircuit function. Science 345, 1255263 (2014).
Bock, D. D. et al. Network anatomy and in vivo physiology of visual cortical neurons. Nature 471, 177–182 (2011).
Hofer, S. B. et al. Differential connectivity and response dynamics of excitatory and inhibitory neurons in visual cortex. Nat. Neurosci. 14, 1045–1052 (2011).
Scholl, B., Pattadkal, J. J., Dilly, G. A., Priebe, N. J. & Zemelman, B. V. Local integration accounts for weak selectivity of mouse neocortical parvalbumin interneurons. Neuron 87, 424–436 (2015).
Ko, H. et al. Functional specificity of local synaptic connections in neocortical networks. Nature 473, 87–91 (2011).
Cohen, M. R. & Kohn, A. Measuring and interpreting neuronal correlations. Nat. Neurosci. 14, 811–819 (2011).
Cossell, L. et al. Functional organization of excitatory synaptic strength in primary visual cortex. Nature 518, 399–403 (2015).
Tsodyks, M. V., Skaggs, W. E., Sejnowski, T. J. & McNaughton, B. L. Paradoxical effects of external modulation of inhibitory interneurons. J. Neurosci. 17, 4382–4388 (1997).
Ozeki, H., Finn, I. M., Schaffer, E. S., Miller, K. D. & Ferster, D. Inhibitory stabilization of the cortical network underlies visual surround suppression. Neuron 62, 578–592 (2009).
Larkum, M. A cellular mechanism for cortical associations: an organizing principle for the cerebral cortex. Trends Neurosci. 36, 141–151 (2013).
Silberberg, G. & Markram, H. Disynaptic inhibition between neocortical pyramidal cells mediated by Martinotti cells. Neuron 53, 735–746 (2007).
Takahashi, N., Oertner, T. G., Hegemann, P. & Larkum, M. E. Active cortical dendrites modulate perception. Science 354, 1587–1590 (2016).
Runyan, C. A. & Sur, M. Response selectivity is correlated to dendritic structure in parvalbumin-expressing inhibitory neurons in visual cortex. J. Neurosci. 33, 11724–11733 (2013).
Jiang, X. et al. Principles of connectivity among morphologically defined cell types in adult neocortex. Science 350, aac9462 (2015).
Zhang, S. et al. Long-range and local circuits for top-down modulation of visual cortex processing. Science 345, 660–665 (2014).
Wertz, A. et al. Single-cell-initiated monosynaptic tracing reveals layer-specific cortical network modules. Science 349, 70–74 (2015).
Harris, K. D. & Mrsic-Flogel, T. D. Cortical connectivity and sensory coding. Nature 503, 51–58 (2013).
Pfeffer, C. K., Xue, M., He, M., Huang, Z. J. & Scanziani, M. Inhibition of inhibition in visual cortex: the logic of connections between molecularly distinct interneurons. Nat. Neurosci. 16, 1068–1076 (2013).
Lee, S., Kruglikov, I., Huang, Z. J., Fishell, G. & Rudy, B. A disinhibitory circuit mediates motor integration in the somatosensory cortex. Nat. Neurosci. 16, 1662–1670 (2013).
Cichon, J. & Gan, W.-B. Branch-specific dendritic Ca2+ spikes cause persistent synaptic plasticity. Nature 520, 180–185 (2015).
Hu, H., Cavendish, J. Z. & Agmon, A. Not all that glitters is gold: off-target recombination in the somatostatin-IRES-Cre mouse line labels a subset of fast-spiking interneurons. Front. Neural Circuits 7, 195 (2013).
Acknowledgements
We thank P. Znamenskiy and G. Keller for discussions of the results in this manuscript. We thank A. Keller for advice on the immunostaining protocol and M. Li for technical assistance. We thank the GENIE Program and Janelia Research Campus of the Howard Hughes Medical Institute for making GCaMP6 material available. This work was supported by the European Research Council (S.B.H., HigherVision 337797; T.D.M.-F., NeuroV1sion 616509), the Swiss National Science Foundation (SBH, 31003A_169525), the Marie Curie Actions of the European Union’s FP7 program (J.P., 332141; A.G.K., 301742), an Ambizione grant from the SNSF (A.G.K., PZ00P3_168046), the UCL Excellence fellowship (J.P.), an EMBO long-term postdoc fellowship (A.B., ALTF 74-2014), the Gatsby Charitable Foundation (M.S.), the Simons Foundation (M.S., SCGB 323228, 543039) and Biozentrum core funds (University of Basel).
Author information
Authors and Affiliations
Contributions
A.G.K., J.P., T.D.M.-F. and S.B.H. designed the experiments. A.G.K. and J.P. built the setup for imaging experiments in behaving mice, performed the experiments and analyzed the data. A.C. developed the MVAR model, performed the model-based data analysis and wrote the modeling results with supervision from M.S. and help from J.P. A.B. performed the immunostaining and did the simultaneous calcium imaging and loose patch recordings. A.B., A.G.K. and J.P. established and performed the post hoc cell matching procedure. A.G.K., J.P., T.D.M.-F. and S.B.H. wrote the paper, with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Co-registration of in vivo two-photon images and immunohistochemically stained brain sections.
(a, b) Top, example regions of in vivo images of GCaMP6f-labelled neurons. Bottom, same regions after post-hoc immunostaining for PV, SOM and VIP (red, blue and magenta respectively) and image registration to match the in vivo image. GCaMP6f label is shown in green. Corresponding interneurons are indicated by arrowheads. Scale bars, 20 μm. Image registration and cell matching was performed for each mouse (N = 8).
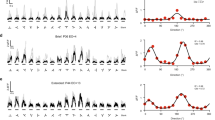
Supplementary Figure 2 Responses to the grating stimuli before and after learning.
(a, b) Average responses of all recorded neurons of each cell type before (a) and after learning (b) to the vertical (left) and angled grating (right). Calcium responses are baseline corrected (subtraction of baseline ΔF/F −0.5 to 0 s before stimulus onset), and aligned to grating onset (dashed line). Cells are sorted by their average response amplitude 0-1 s from stimulus onset. Number of cells included in each plot: 1249, 132, 58 and 175 for PYR, PV, SOM and VIP, respectively. Bottom, average responses of cells from the top, middle and bottom 10th percentiles of grating-evoked response amplitudes of each cell class. (125, 13, 6 and 18 cells in each 10th percentile, respectively) Shaded area represents SEM here and elsewhere. (c) Average response to the angled grating of all cells from each cell class after learning. (d) Similarity of response profiles to the angled grating of all pairs of cell classes attained with a random forest decoder to classify single cells to one of two classes based on the shape of their average baseline-subtracted response profiles (see Online Methods). Response profile similarity score = 2 × (1- classification accuracy). Scores near 0 and 1 indicate low and high response profile similarity between two cell classes respectively.
Supplementary Figure 3 Responses triggered by the onset of running, reward and odor.
(a) Top, average responses to the onset of running in the dark of all recorded neurons of each cell type as the mouse performed an odor discrimination task in the dark (see Online Methods). Calcium responses are baseline corrected (subtraction of baseline ΔF/F −0.5 to 0 s before running onset), and aligned to running onset (dashed line). Cells are sorted by their average response amplitude 0–1 sec from onset. Number of cells included in a,c,d: 2795, 185, 75 and 192 and in b 1064, 121, 52, 156 for PYR, PV, SOM and VIP, respectively. Bottom, average responses of cells from the top, middle and bottom 10th percentiles of running onset-aligned response amplitudes of each cell class (in a,c,d: 208, 19, 8 and 19 cells and in b: 106, 12, 5 and 16 cells in each 10th percentile, respectively). Shaded area represents SEM here and elsewhere. Running onsets were defined as times where running speed of the mouse crossed 6 cm/s after being stationary for at least 0.5 s. (b) Responses aligned to onset of running in the grey corridor during the visual discrimination task, similar to (a). Only periods when mice were in the grey corridor for at least 0.5 s before and 1 s after running onset were selected, to minimize optic-flow related responses. (c) Responses aligned to reward onset during the odor discrimination task (see Online Methods), similar to (a). (d) Responses aligned to the non-rewarded odor onset during the odor discrimination task (see Online Methods), similar to (a).
Supplementary Figure 4 Response changes during learning.
(a, d) Average response profiles in response to vertical (a) and angled grating stimulus (d) of all cells from each cell class before (dashed line) and after (solid line) learning. Vertical dashed line indicates grating onset. Shaded area represents SEM, here and elsewhere. N = 1249, 132 PV, 58 SOM and 175 VIP cells here and below. (b, e) Top: Difference between post-learning and pre-learning response profiles in response to the vertical (b) and angled (e) grating stimuli for all cells of different cell classes. Responses were baseline corrected before subtracting (baseline -.5 to 0 sec before stimulus onset), are shown aligned to grating onset (dashed line), and color coded for Δ(ΔF/F). Bottom: average responses of cells from the top, middle and bottom 10th percentiles of the response differences shown on top. (c, f) Fractions of cells in which the response amplitude to the vertical (c) and angled (f) grating stimuli increased, decreased, or showed no difference from pre-to post-learning in the period 0-1 s from stimulus onset.
Supplementary Figure 5 Effect of eye position, pupil size, eye movements, running and licking on selectivity of responses.
(a) Cumulative distribution of absolute selectivity of each cell class before and after learning. Sign test, **, P < 0.001; *, P < 0.05 here and below. (b) Distributions of average horizontal (nasal-temporal axis) and vertical (ventral-dorsal axis) eye positions and (c) pupil sizes of the contralateral eye in individual trials 0-1 s after onset of the vertical (V) and angled (A) grating, before and after learning. (d) Saccade rate 0-1 s after grating onset before and after learning. (e-i) Mean absolute selectivity of each cell class before and after learning (computed in the period of 0-1 s after grating onset) after equalizing the distributions of horizontal and vertical eye positions in all conditions (e), after equalizing the distributions of pupil sizes (f), when excluding all trials with eye movements (g), when excluding all trials with licks (h), and after equalizing the distributions of running speed (i). Error bars are SEM. Numbers of cells recorded both pre- and post-learning: 1249 PYR, 132 PV, 58 SOM and 175 VIP cells, N = 8 mice. (j) Response selectivity in the approach corridor before and after learning.
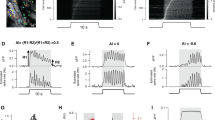
Supplementary Figure 6 Relationship between action potential firing rate and calcium transient size in simultaneous loose patch and GCaMP6f recordings from the three interneuron classes in visual cortex slices.
Black lines are individual cells, colored lines are averages across cells, plotted for firing rate bins including at least five cells. Error bars represent SEM, PV = 13, SOM = 17 and VIP = 11 cells.
Supplementary Figure 7 Changes in noise correlation during learning.
(a) Top: Mean noise correlation measured during the angled grating response (0-1 s from stimulus onset) between cell pairs of each combination of cell classes, before and after learning. The number of cell pairs here and below in each cell class combination (pre, post-learning) was: PYR-PYR 77599, 66633, VIP-VIP 984, 776, SOM-SOM 201, 131, PV-PV 1646, 1316, PV-VIP 818, 702, PV-PYR 17496, 15029, PYR-VIP 14485, 11893, SOM-PV 1176, 828, SOM-PYR 7121, 5545, SOM-VIP 476, 364. Error bars represent SEM. (b, c) Distributions of noise correlation between cell pairs of each combination of cell classes during the vertical (b) or angled (c) grating response before and after learning. (d, e) Fraction of cell pairs with significantly negative noise correlation during the vertical (d) or angled (e) grating response before and after learning. Only neurons with significant responses to the grating were included in the analysis.
Supplementary Figure 8 Changes in eye position, pupil size, eye movements, running, licking and reductions in response strength cannot account for reduction in noise correlations with learning.
Mean noise correlations measured during the vertical grating response (0-1 s from stimulus onset) after equalizing the distributions of eye position (a), pupil sizes (b), when excluding all trials with eye movements (c), when excluding all trials with licks (d), after equalizing the distributions of running speed (e), and after controlling for reductions in response strength (f). The number of cell pairs (pre, post-learning) was: PYR-PYR 77599, 66633, VIP-VIP 984, 776, SOM-SOM 201, 131, PV-PV 1646, 1316, PV-VIP 818, 702, PV-PYR 17496, 15029, PYR-VIP 14485, 11893, SOM-PV 1176, 828, SOM-PYR 7121, 5545, SOM-VIP 476, 364. Error bars represent SEM.
Supplementary Figure 9 Variability in running speed, visual flow and eye position did not make a large contribution to measured noise correlation.
(a, b) Variability in running speed and coupled visual flow was reduced by sampling trials only from the middle 50% of running speed distribution (25th to 75th percentile), and compared to an equal number of trials sampled randomly from the full distributions. Noise correlations obtained in these two conditions were very similar, both for individual cell pairs (a), and for average noise correlations between specific cell classes (b). (c, d) Same analysis for eye position. R indicates Pearson correlation coefficient. The number of cell pairs (pre, post-learning) was: PYR-PYR 77599, 66633, VIP-VIP 984, 776, SOM-SOM 201, 131, PV-PV 1646, 1316, PV-VIP 818, 702, PV-PYR 17496, 15029, PYR-VIP 14485, 11893, SOM-PV 1176, 828, SOM-PYR 7121, 5545, SOM-VIP 476, 364. Error bars represent SEM.
Supplementary Figure 10 Multivariate autoregressive (MVAR) linear dynamical system model.
(a) Root mean square (RMS) of the strength of the three types of inputs in the MVAR model for all cells of each class (N = 1249 PYR, 132 PV, 58 SOM and 175 VIP cells). Running speed has relatively small contributions. (b) Cross-validated R2 (mean over cells, N = 1614) for models of increasing complexity. The MVAR model performed better than the other models (all Ps < 10−150, sign test). The small increase in R2 when interactions were included indicates that the cross-validated prediction performance improved despite the considerable increase in model complexity (approximately 40,000 extra parameters per animal), which would be expected to lead to a large drop in performance due to overfitting if interactions were not informative. In contrast, the negative R2 values for the model that included time-varying interactions indicate that the inclusion of time-varying interaction weights leads to overfitting on the training data. (c) Histograms of the difference in R2 obtained from the MVAR model and each of the other models tested in (b) for each cell. In all comparisons the majority of cells performed better with the MVAR model (positive ΔR2 in 93%, 83% and 99% of cells compared to models with average response profiles only, MVAR without interactions and MVAR with time varying interactions respectively). (d) Average pre-learning noise correlations observed (grey), after setting interaction weights to zero (orange), and after shuffling residuals (white), similar to Fig. 4c. The number of cell pairs was: PYR-PYR 77599, VIP-VIP 984 SOM-SOM 201, PV-PV 1646, PV-VIP 818, PV-PYR 17496, PYR-VIP 14485, SOM-PV 1176, SOM-PYR 7121, SOM-VIP 476. (e) Mean interaction weights obtained from the MVAR model fit pre-and post-learning. Error bars represent SEM.
Supplementary Figure 11 Effect of deleting and constraining parameters in the MVAR model.
(a) Effect of deleting all interaction weights between cells in the MVAR model on selectivity in PYR (top) and VIP cells (bottom) before (left) and after (right) learning. N = 1249 PYR, 175 VIP, 132 PV and 58 SOM cells here and below. Bars indicate average absolute selectivity with weights intact and deleted. *, P < 10-3. Sign test comparing intact and deleted conditions, PYR cells pre-learning P < 10−11, post-learning P < 10−17, VIP cells pre-learning P < 10−3, post-learning P > 0.05. Error bars represent SEM. (b) Effect of deleting specific interaction weights onto PV, SOM and PYR cells on their selectivity pre- and post-learning (Δselectivity). Same data as in Fig. 4g–i, showing distributions. Vertical lines indicate mean. **, P < 10−3, *, P < 0.05 here and below. (c) Interaction weights in MVAR model before and after learning for cell pairs with the same or opposite stimulus-input preference. Error bars represent SEM. Wilcoxon rank-sum test, N interaction weights range from 50 to 41548. (d) Distribution of interaction weights in MVAR model before and after learning for PYR-PYR and PYR-PV cell pairs with the same or opposite stimulus-input preference, similar to Fig. 4j. Vertical lines indicate mean. N interaction weights pre- and post-learning for same or opposite preference pairs, PYR-PYR pre 20074, 16750 post 41548, 38056, PYR-PV pre 2132, 1513, post 4856, 4300 (e) MVAR jointly fit to the pre- and post-learning data, while constraining specific parameters (stimulus inputs or interaction weights) to remain fixed across learning and allowing others to vary. Holding all or none of the parameters fixed gave poor or good fits of selectivity changes during learning respectively (left, example R2 with all parameters free or restrained shown for PYR cells, values for all cell classes indicated by horizontal lines in right panel). When stimulus inputs to PYR or SOM cells were held fixed over learning, the model failed to fully capture their respective selectivity changes (right, inputs restrained, bootstrap test with resampling of residuals (see Online Methods) on the difference between model with all parameters free compared to model with parameters fixed, P-values < 0.002). In contrast, the relatively large changes in PV selectivity were not significantly disrupted when stimulus inputs to PV cells were held fixed (P = 0.4), indicating that changes in interaction weights and stimulus inputs to other cell types are sufficient to account for the selectivity changes observed in PV cells during learning. The model with VIP inputs fixed also fully captured VIP selectivity changes (P = 0.71), likely due to the selectivity changes in these cells being very small. Furthermore, simultaneously fixing both PV inputs and PYR to PV weights significantly impaired the model to fully capture PV selectivity changes (both relative to the full model (P = 0.03) and relative to the model with fixed PV inputs alone (P < 0.002). Error bars represent 90% confidence intervals from bootstrapping with resampling of cells.
Supplementary Figure 12 After correcting for neuropil contamination, learning-related changes in selectivity as well as changes in interactions remain largely unchanged.
We subtracted from each ROI the average fluorescence of all pixels within a neuropil mask surrounding the cell (see Online Methods) and repeated key analyses. Panels a and b reproduce Fig. 2d, g. 1249 PYR, 132 PV, 58 SOM and 175 VIP cells in a and b. R is Pearson correlation coefficient. Panel c reproduces Fig. 3c. *, P < 10−2, **, P < 10−9, sign test, error bars represent SEM. The number of cell pairs in each cell class combination (pre, post-learning) was: PYR-PYR 74581, 64921, VIP-VIP 1166, 907, SOM-SOM 215, 99, PV-PV 1731, 1369, PV-VIP 790, 718, PV-PYR 17792, 15238, PYR-VIP 14681, 12009, SOM-PV 1250, 690, SOM-PYR 7112, 4952, SOM-VIP 455, 377.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Khan, A.G., Poort, J., Chadwick, A. et al. Distinct learning-induced changes in stimulus selectivity and interactions of GABAergic interneuron classes in visual cortex. Nat Neurosci 21, 851–859 (2018). https://doi.org/10.1038/s41593-018-0143-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0143-z
This article is cited by
-
Synaptic configuration and reconfiguration in the neocortex are spatiotemporally selective
Anatomical Science International (2024)
-
The plasticitome of cortical interneurons
Nature Reviews Neuroscience (2023)
-
A cell-type-specific error-correction signal in the posterior parietal cortex
Nature (2023)
-
Opto-juxtacellular interrogation of neural circuits in freely moving mice
Nature Protocols (2023)
-
Distributing task-related neural activity across a cortical network through task-independent connections
Nature Communications (2023)