Abstract
The dorsal horn of the spinal cord is critical to processing distinct modalities of noxious and innocuous sensation, but little is known of the neuronal subtypes involved, hampering efforts to deduce principles governing somatic sensation. Here we used single-cell RNA sequencing to classify sensory neurons in the mouse dorsal horn. We identified 15 inhibitory and 15 excitatory molecular subtypes of neurons, equaling the complexity in cerebral cortex. Validating our classification scheme in vivo and matching cell types to anatomy of the dorsal horn by spatial transcriptomics reveals laminar enrichment for each of the cell types. Neuron types, when combined, define a multilayered organization with like neurons layered together. Employing our scheme, we find that heat and cold stimuli activate discrete sets of both excitatory and inhibitory neuron types. This work provides a systematic and comprehensive molecular classification of spinal cord sensory neurons, enabling functional interrogation of sensory processing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lewin, G. R. & Moshourab, R. Mechanosensation and pain. J. Neurobiol. 61, 30–44 (2004).
Ma, Q. Population coding of somatic sensations. Neurosci. Bull. 28, 91–99 (2012).
Abraira, V. E. & Ginty, D. D. The sensory neurons of touch. Neuron 79, 618–639 (2013).
Lallemend, F. & Ernfors, P. Molecular interactions underlying the specification of sensory neurons. Trends Neurosci. 35, 373–381 (2012).
Li, C. L. et al. Somatosensory neuron types identified by high-coverage single-cell RNA-sequencing and functional heterogeneity. Cell Res. 26, 83–102 (2016).
Liu, Y. & Ma, Q. Generation of somatic sensory neuron diversity and implications on sensory coding. Curr. Opin. Neurobiol. 21, 52–60 (2011).
Prescott, S. A., Ma, Q. & De Koninck, Y. Normal and abnormal coding of somatosensory stimuli causing pain. Nat. Neurosci. 17, 183–191 (2014).
Usoskin, D. et al. Unbiased classification of sensory neuron types by large-scale single-cell RNA sequencing. Nat. Neurosci. 18, 145–153 (2015).
Duan, B., Cheng, L. & Ma, Q. Spinal circuits transmitting mechanical pain and itch. Neurosci. Bull. 34, 186–193 (2018).
Graham, B. A., Brichta, A. M. & Callister, R. J. Moving from an averaged to specific view of spinal cord pain processing circuits. J. Neurophysiol. 98, 1057–1063 (2007).
Peirs, C. & Seal, R. P. Neural circuits for pain: recent advances and current views. Science 354, 578–584 (2016).
Polgár, E., Fowler, J. H., McGill, M. M. & Todd, A. J. The types of neuron which contain protein kinase C gamma in rat spinal cord. Brain Res. 833, 71–80 (1999).
Todd, A. J. Identifying functional populations among the interneurons in laminae I-III of the spinal dorsal horn. Mol. Pain 13, 1744806917693003 (2017).
Bourane, S. et al. Gate control of mechanical itch by a subpopulation of spinal cord interneurons. Science 350, 550–554 (2015).
Bourane, S. et al. Identification of a spinal circuit for light touch and fine motor control. Cell 160, 503–515 (2015).
Duan, B. et al. Identification of spinal circuits transmitting and gating mechanical pain. Cell 159, 1417–1432 (2014).
Foster, E. et al. Targeted ablation, silencing, and activation establish glycinergic dorsal horn neurons as key components of a spinal gate for pain and itch. Neuron 85, 1289–1304 (2015).
Petitjean, H. et al. Dorsal horn parvalbumin neurons are gate-keepers of touch-evoked pain after nerve injury. Cell Rep. 13, 1246–1257 (2015).
Ross, S. E. et al. Loss of inhibitory interneurons in the dorsal spinal cord and elevated itch in Bhlhb5 mutant mice. Neuron 65, 886–898 (2010).
Ross, S. E. et al. Bhlhb5 and Prdm8 form a repressor complex involved in neuronal circuit assembly. Neuron 73, 292–303 (2012).
Sun, Y. G. et al. Cellular basis of itch sensation. Science 325, 1531–1534 (2009).
Todd, A. J. Neuronal circuitry for pain processing in the dorsal horn. Nat. Rev. Neurosci. 11, 823–836 (2010).
Abraira, V. E. et al. The cellular and synaptic architecture of the mechanosensory dorsal horn. Cell 168, 295–310.e219 (2017).
Rexed, B. The cytoarchitectonic organization of the spinal cord in the cat. J. Comp. Neurol. 96, 414–495 (1952).
Rexed, B. A cytoarchitectonic atlas of the spinal cord in the cat. J. Comp. Neurol. 100, 297–379 (1954).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Caldeira, V., Dougherty, K. J., Borgius, L. & Kiehn, O. Spinal Hb9:Cre-derived excitatory interneurons contribute to rhythm generation in the mouse. Sci. Rep. 7, 41369 (2017).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
van der Maaten, L. & Hinton, G. E. Visualizing high-dimensional data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Christensen, A. J. et al. In vivo interrogation of spinal mechanosensory circuits. Cell Rep. 17, 1699–1710 (2016).
Malmberg, A. B., Chen, C., Tonegawa, S. & Basbaum, A. I. Preserved acute pain and reduced neuropathic pain in mice lacking PKCγ. Science 278, 279–283 (1997).
Peirs, C. et al. Dorsal horn circuits for persistent mechanical pain. Neuron 87, 797–812 (2015).
Sun, Y. G. & Chen, Z. F. A gastrin-releasing peptide receptor mediates the itch sensation in the spinal cord. Nature 448, 700–703 (2007).
Lai, H. C., Seal, R. P. & Johnson, J. E. Making sense out of spinal cord somatosensory development. Development 143, 3434–3448 (2016).
Cameron, D. et al. The organisation of spinoparabrachial neurons in the mouse. Pain 156, 2061–2071 (2015).
Todd, A. J., McGill, M. M. & Shehab, S. A. Neurokinin 1 receptor expression by neurons in laminae I, III and IV of the rat spinal dorsal horn that project to the brainstem. Eur. J. Neurosci. 12, 689–700 (2000).
Al-Khater, K. M. & Todd, A. J. Collateral projections of neurons in laminae I, III, and IV of rat spinal cord to thalamus, periaqueductal gray matter, and lateral parabrachial area. J. Comp. Neurol. 515, 629–646 (2009).
Coggeshall, R. E. Fos, nociception and the dorsal horn. Prog. Neurobiol. 77, 299–352 (2005).
Sathyamurthy, A. et al. Massively parallel single nucleus transcriptional profiling defines spinal cord neurons and their activity during behavior. Cell Rep. 22, 2216–2225 (2018).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Aresh, B. et al. Spinal cord interneurons expressing the gastrin-releasing peptide receptor convey itch through VGLUT2-mediated signaling. Pain 158, 945–961 (2017).
Kardon, A. P. et al. Dynorphin acts as a neuromodulator to inhibit itch in the dorsal horn of the spinal cord. Neuron 82, 573–586 (2014).
Cui, L. et al. Identification of early RET+ deep dorsal spinal cord interneurons in gating pain. Neuron 91, 1413 (2016).
Huang, H. Y. et al. Expression of A-type K channel alpha subunits Kv 4.2 and Kv 4.3 in rat spinal lamina II excitatory interneurons and colocalization with pain-modulating molecules. Eur. J. Neurosci. 22, 1149–1157 (2005).
Hu, H. J. et al. The kv4.2 potassium channel subunit is required for pain plasticity. Neuron 50, 89–100 (2006).
Hunt, S. P., Pini, A. & Evan, G. Induction of c-fos-like protein in spinal cord neurons following sensory stimulation. Nature 328, 632–634 (1987).
Plath, N. et al. Arc/Arg3.1 is essential for the consolidation of synaptic plasticity and memories. Neuron 52, 437–444 (2006).
Zheng, J., Lu, Y. & Perl, E. R. Inhibitory neurones of the spinal substantia gelatinosa mediate interaction of signals from primary afferents. J. Physiol. (Lond.) 588, 2065–2075 (2010).
Borgius, L., Restrepo, C. E., Leao, R. N., Saleh, N. & Kiehn, O. A transgenic mouse line for molecular genetic analysis of excitatory glutamatergic neurons. Mol. Cell. Neurosci. 45, 245–257 (2010).
Romanov, R. A. et al. Molecular interrogation of hypothalamic organization reveals distinct dopamine neuronal subtypes. Nat. Neurosci. 20, 176–188 (2017).
La Manno, G. et al. Molecular diversity of midbrain development in mouse, human, and stem cells. Cell 167, 566–580.e519 (2016).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 14, 22–29 (2012).
Lin, D. et al. Functional identification of an aggression locus in the mouse hypothalamus. Nature 470, 221–226 (2011).
Acknowledgements
We thank Science for Life Laboratory, the National Genomics Infrastructure funded by the Swedish Research Council, and Uppsala Multidisciplinary Center for Advanced Computational Science for providing assistance in massively parallel sequencing and access to the UPPMAX computational infrastructure. This work was supported by the Swedish Medical Research Council, Knut and Alice Wallenbergs Foundation (Wallenberg Scholar and Wallenberg project grant), SFO grant (StratNeuro), Wellcome Trust (Pain Consortium), European Research Council advanced grant (PainCells 740491) and Karolinska Institutet (to P.E.); Swedish Foundation for Strategic Research, Knut and Alice Wallenberg Foundation (2015.0041) (to S.L.); Ragnar Söderberg Foundation and the Brain Foundation (to M.C.L.). A.Z. was supported by the Human Frontier Science Program.
Author information
Authors and Affiliations
Contributions
M.H., A.Z. and H.H. are equally contributing authors, P.E. and S.L. are co-corresponding authors and P.E. is the lead author. P.E. and S.L. supervised and designed the study with input from M.H., A.Z. and H.H. M.H. planned all experiments, performed cell preparations and planned sequencing, helped with analysis and generated figures, and together with N.S. performed in situ hybridization. A.Z. planned experiments, analyzed sequencing and in situ data and generated figures. H.H. planned experiments, performed single-cell sequencing experiments, and helped analyze data and generate figures. P.R. performed animal work for thermal and cold experiments, G.L.M. and P.L. analyzed data, J.E.T.J. and M.C.L. performed animal work for tracing experiments, and L.B. and O.K. carried out some animal experiments. P.E. wrote the paper with help from M.H., H.H., A.Z. and S.L. and input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Dissection and dissociation of the spinal cord dorsal horn (related to Fig. 1).
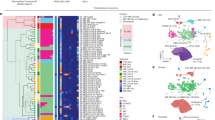
a) Upper row shows 3 representative sections from an intact hemisphere of spinal cord (from left to right: Cervical, thoracical, lumbar) The lower row shows 3 corresponding sections of only the dorsal horn after dissection. The dissection included the dorsal horn. Neurons ventral to the dorsal horn such as Clarke´s column was not part of the analyzed tissue. Scale bar = 200 µm for upper and 300 µm lower panels. b) Illustration of single cell suspension and the capturing of single cells in the C1 Fluidigm Chip. Scale bar = 50 µm for upper panel and 12 µm lower panels. c) Illustration of FACS from VglutGFP (left) and VgatTdTomato (right) dorsal horn suspensions. d) Heat map displaying the single cell expression profile of 1639 dorsal horn spinal cord neurons divided into glutamatergic (left) and GABAergic (right) neurons. Heat map is based on relative expression of each gene, dark green representing minimal and yellow maximal expression (“summer” color scheme). Neurons passed quality control, namely, containing at least 3000 molecules. Black vertical lines represent the 43 clusters defined by backspin. The color code below illustrates the final clustering after manual control. Clusters without distinguishable differences were fused (i.e. Cluster 4 and 6 of the glutamatergic neurons) and clusters without defining genes (thick black bars) were excluded resulting in a total number of 30 clusters.
Supplementary Figure 2 Determination of cell types by marker genes (related to Figs. 1, 2 and 3).
a) Molecule and gene count of the remaining 1545 cells. Black squares and error bars within each cluster represent the average values andstandart deviation). Clusters were hierarchically reordered to represent the relation between the 30 clusters (Number of cells for each cluster from left to right: 32, 34, 34, 34, 24, 75, 105, 77, 34, 129, 61, 35, 38, 34, 22, 18, 87, 145, 11, 19, 51, 39, 67, 85, 66, 48, 56, 34, 41, 11). b) Heatmaps showing the contribution of cells to each cluster by each of the donors (donors = different animals). Donors are named DH followed by a number that identifies the donor and the number of cells that were captured in each experiment is indicated in parenthesis. There are three heatmaps. The first is for VgattdTomato FACS sorted cells (n = 3 animals), the second for Vglut2GFP FACS sorted cells (n = 5 animals) and the bottom is unbiased dissociation of the dorsal horn and collection of cells (without FACS sorting) (n = 20 animals). This shows that cells are contributed to most clusters in each experiment and that this can be repeated over and over with similar results. The distribution of cells in each experiment is found by reading the heatmap horizontally. Some cell types are expected to be more abundant than others and will because of this appear stronger in the heatmap. Examining the heatmap vertically shows that there is no strong bias in contribution of cells between experiments. c) Contribution of donors (animals) to each cluster (neuron type) vs the cluster size. Large clusters are expected to have more donors than small clusters simply as the probability of capturing a cell belonging to that cluster is greater. If batch effects were strong, cell types should appear in certain batches (donors) but not in the other batches. Please note that even the smallest cluster with around 15 cells is composed of >7 donors (experiments). This can be taken as an indicator of the biological reproducibility of the clusters and the ability to identify all cell types across different experiments. d) Assessment of the predicted strength of clusters. Random forest classifier (max depth 30) was trained on 80% of the cells and testing its performance on the remaining 20% of the cells. The heatmap shows the predicted strength of the cluster identity. The rows are our “observed” cell types (i.e. the ones we assigned). The columns are the predictions the classifier makes. The color shows the probability of assigning each label, averaged across all the test cases. The mean probability along the diagonal is 69.8%.
Supplementary Figure 3 Relation of the identified neuron types to known literature (related to Figs. 1, 2 and 3).
a) Neurochemistry and morphology. Green indicates glutamatergic neurons, red GABAergic neurons. The lines depict predicted detection of the markers using immunohistochemistry. Immunohistochemistry is not as sensitive or quantitative as in situ hybridization. This figure therefore only depicts the high-expressing cell-types that we predict have been detected in the cited studies. The actual expression data of these markers is available as Supplementary Fig 4a. For morphology, CALB2+ and PVALB+ cells are often islet cells, PDYN+ and NPY+ are heterogenous but never Islet cells, PDYN+ cells are sometimes vertical cells. b) Predicted function of neuron types. Illustration of neuron types subjectively predicted to have been functionally analyzed. Note that mouse models used Cre driver lines and thus cells expressing the gene at any point in development will recombine and be included in the study and therefore, this summary has to be interpreted with this in mind. Due to transient developmental expression or low levels of expression, studies on Slc17a8, Ret, Bhlhb5, Slc6a5 were excluded. Stars indicate the gene also to be expressed in neurons belonging to the other neurotransmitter type which is not shown in the figure. For the actual expression profile of these genes in both neurotransmitter types, see Supplementary Fig. 4a.
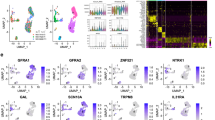
Supplementary Figure 4 Expression of previously used markers and new markers for identification of dorsal horn neuron types (related to Figs. 1, 2 and 3).
a) Violin plots of genes previously used as markers to characterize cell types involved in sensation. Note broad expression in many neuronal types, often also crossing neurotransmitter boundaries. b) Violin plots of minimal set of new markers to uniquely identify each neuronal type (colored by neuronal type). Vertical gives the proposed markers, horizontally the expression in other neuron types.
Supplementary Figure 5 Expression of transcription factors, neuropeptides/receptors and ion channels (related to Fig. 4).
Listed are those genes that show significant expression in at least one cluster (see Online Methods, Negative binomial generalized linear regression). Note that this list shows all significantly expressed genes (p < 0.001) while in the main Fig. 4, only significantly expressed with a minimum of 3 transcripts are shown. Thus, with the relaxed inclusion criteria, many more genes can be detected in this analysis than in main Fig. 4. (n = 1545 neurons).
Supplementary Figure 6 In situ mRNA staining using RNAScope (related to Fig. 5).
a) Cervical and lumbar section stained for Nmu (red), Tac2 (green) and Sst (blue) in order to detect cells representing the clusters Glut5-7. The celltypes are highlighted in the small panels on the right using arrows (Glut5 [red arrow], Glut6 [yellow arrow], Glut7 [white arrow]). The small panels represent the magnified region depicted by the yellow frame in the corresponding panel on the left. b) Cervical and lumbar section stained for Qrfpr (red), Crabp1 (green) and Npy (blue) in order to detect cells representing the clusters Gaba5-7. The cell types are highlighted in the small panels on the right using arrows (Gaba5 [red arrow], Gaba6 [yellow arrow], Gaba7 [white arrow]). The small panels represent the magnified region depicted by the yellow frame in the corresponding panel on the left. c) Two example cells for cluster Gaba1 expressing beside Gal both Slc17a6 and Gad1. d) Images of the lumbar and cervical section that were used as reference sections to the Allan Brain Reference Atlas. e) In order to align two sections, we defined in total 10 Reference points for each section (including the reference section; 5 for the left and right horn respectively, starting always at the central canal). Each image was then cropped and transformed so that they could be aligned using the reference points. Scale bars = 200 µm for overview picture and 50 µm for magnifications in panels a) and b), 20 µm in panel c) as well as 500 µm in panel d) and e).
Supplementary Figure 7 Spatial distribution of the neuronal types within the cervical spinal cord (related to Fig. 5).
a) Localization of all the neuronal types within the cervical dorsal spinal cord. Red dots represent single cells expressing the respective marker genes defining the indicated cell type. Lines represent Rexed laminae. Names of each neuronal type and number of identified cells (in parenthesis) are indicated on top of each spinal cord projection. n = 2 animals; Spatial information was collected and combined from 6 sections (3 sections per animal per staining) b) Quantification of distribution of each neuronal type within the six dorsal Rexed laminae of the cervical dorsal spinal cord. Size of the gray circle represents percentage of cells of a particular type found within a particular layer. c) Quantification of distribution of each neuronal type within the six dorsal Rexed laminae of the lumbar dorsal spinal cord. Size of the gray circle represents percentage of cells of a particular type found within a particular layer. d) Density heat map of glutamatergic and GABAergic neurons using Slc17a6 and Gad1 in lumbar or cervical spinal cord (n = 2 animals; Spatial information was collected and combined from total 60 section for Slc17a6 and 37 sections for Gad1) or e) Slc17a6 and Slc32a1 in the lumbar spinal cord (n = 3 animals Spatial information was collected and combined from total 28 section for Slc17a6 and 19 sections for Slc32a1). Scale bars = 1000 µm in panels a) and 500 µm in panel d) and e). Heat map uses “parula” color scheme, dark blue represent minimal and yellow maximal expression.
Supplementary Figure 8 Overview of the CTb488 injection area in the pons and traced projection neurons in the spinal cord (related to Fig. 5).
a) Coronal section of the pons displaying the Parabrachial nucleus (white dashed circles) as CTb488 injection site. CTb488 signal (bright yellow/white) can be seen in the Parabrachial nucleus and in the dorsal Inferior colliculus. b) Reference section from Allen mouse brain atlas (image 103/132). The Parabrachial nucleus is marked with black dashed circles. c) image of dorsal horn surface during dissection at cervical, lumbar and sacral levels. Rostral side is up and caudal side is down. Projection neurons are visible in green. Scale is not available due to variable power objective. Scale bar in a and b is 1000 µm and was constructed from the Allen mouse brain section.
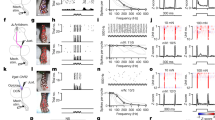
Supplementary Figure 9 Identification of heat- and cold-activated neurons (related to Fig. 6).
a) Localization the immediate early gene Arc within the lumbar dorsal spinal cord following noxious heat or cold as indicated using different thresholds. Red dots represent single cells detected to express Arc. With increased stringency (higher threshold) number of detected cells decreases and significance increases. The numbers of neurons detected in left and right dorsal horn and the predicted correctly identified positive cells (presented as %) are indicated at each threshold. b) Panel shows the relation between specificity and sensitivity (normalized). Red line depicted closest point to origin as simple optimal threshold criteria. Dash lines illustrate the threshold (>5) chosen for the further analysis for heat and threshold (>8) chosen for cold. c) Identification of neuronal types based on marker genes (marker genes>3 transcripts/cell) that become active by heat (Arc>5/cell). Red dots represent single neurons expressing the respective marker genes together with Arc. Below, quantification of the percent of neurons of each neuron types that were active (Arc+) and percent of Arc+ neurons belonging to the different neuron types as percent of all Arc+ neurons in the control (contralateral, left) and experimental (ipsilateral, right) side of the spinal cord following heat. d) Identification of neuronal types based on marker genes (marker genes>3 transcripts/cell) that become active by cold (Arc>9/cell). Red dots represent single neurons expressing the respective marker genes together with Arc. Below, quantification of the percent of neurons of each neuron types that were active (Arc+) and percent of Arc+ neurons belonging to the different neuron types as percent of all Arc+ neurons in the control (contralateral, left) and experimental (ipsilateral, right) side of the spinal cord following cold. n = 3 animals; analysis contains 3 lumbar sections per animal per staining. Scale bars = 1000 µm in panel a), c) and d).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Supplementary Table 1
A list of mainly IEGs (immediate early gene) that were excluded. In particular, immediate early genes were thought to form a bias due to the possibility of sensitive and faster responding cells during dissection.
Supplementary Table 2
Enrichment score for all genes and cell types.
Supplementary Table 3
The top markers that separate each junction of the dendrogram in Fig. 1c,e,f. For each junction, we searched for genes that best separate the two sides of the junction. The average of the log2(x + 1) expression (Sheet 1) or the fraction of positive cells (Sheet 2) in each group (expression > 0) were calculated, and genes were ranked by either of these scores. The tables show the top 50 genes found to be specific for each side of particular junctions (left and right) as mentioned above each sub-table. In addition to the score, the P value was calculated using rank sum (Sheet 1) or binomial distribution (Sheet 2). q values correspond to P values corrected for multiple one-sided testing since each gene was tested for all 29 junctions. The sample size is according to the dendrogram junction numbering and the cluster size as describe in Fig. 1c.
Supplementary Table 4
Cluster size and molecular counts for each cell type.
Supplementary Table 5
Marker genes and combinations that were used to define and detect each cell type.
Supplementary Table 6
Comparison of single-nuclei data of Sathyamurthy et al. to clusters identified in this publication. Pearson's correlation (after log2 + 1 transformation) was calculated on genes enriched in each cell type from both data sets. For each cluster in the single-nuclei data set we identified the best fit cluster as the cluster with the highest correlation to our dataset. 1 is total positive correlation, 0 is no correlation.
Supplementary Table 7
RNAscope probes used in this study
Supplementary Table 8
Sheet 1: Cell numbers obtained for each glutamtergic cell type using the automatic evaluation. Sheet 2: Cell numbers obtained for each gabaergic cell type using the automatic evaluation. Sheet 3: Numbers of Arc+ cells following heat or cold stimulation that colocalize with the respective cell type marker genes using the automatic evaluation. Sheet 4: Genes and common names for markers used in this study. Sheet 5: Minimal number of marker genes to unambiguously identify each neuron type.
Rights and permissions
About this article
Cite this article
Häring, M., Zeisel, A., Hochgerner, H. et al. Neuronal atlas of the dorsal horn defines its architecture and links sensory input to transcriptional cell types. Nat Neurosci 21, 869–880 (2018). https://doi.org/10.1038/s41593-018-0141-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0141-1
This article is cited by
-
Shank2 identifies a subset of glycinergic neurons involved in altered nociception in an autism model
Molecular Autism (2023)
-
The glycine receptor alpha 3 subunit mRNA expression shows sex-dependent differences in the adult mouse brain
BMC Neuroscience (2023)
-
Characterisation of NPFF-expressing neurons in the superficial dorsal horn of the mouse spinal cord
Scientific Reports (2023)
-
Single-cell transcriptomic landscape of the developing human spinal cord
Nature Neuroscience (2023)
-
Single-cell analysis reveals region-heterogeneous responses in rhesus monkey spinal cord with complete injury
Nature Communications (2023)