Abstract
Amyotrophic lateral sclerosis–frontotemporal dementia (ALS-FTD) constitutes a devastating disease spectrum characterized by 43-kDa TAR DNA-binding protein (TDP-43) pathology. Understanding how TDP-43 contributes to neurodegeneration will help direct therapeutic efforts. Here we have created a TDP-43 knock-in mouse with a human-equivalent mutation in the endogenous mouse Tardbp gene. TDP-43Q331K mice demonstrate cognitive dysfunction and a paucity of parvalbumin interneurons. Critically, TDP-43 autoregulation is perturbed, leading to a gain of TDP-43 function and altered splicing of Mapt, another pivotal dementia-associated gene. Furthermore, a new approach to stratify transcriptomic data by phenotype in differentially affected mutant mice revealed 471 changes linked with improved behavior. These changes included downregulation of two known modifiers of neurodegeneration, Atxn2 and Arid4a, and upregulation of myelination and translation genes. With one base change in murine Tardbp, this study identifies TDP-43 misregulation as a pathogenic mechanism that may underpin ALS-FTD and exploits phenotypic heterogeneity to yield candidate suppressors of neurodegenerative disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
05 June 2018
In the version of this article initially published, the footnote number 17 was missing from the author list for the two authors who contributed equally. Also, the authors have added a middle initial for author Justin R. Fallon and an acknowledgement to the Babraham Institute Imaging Facility and Sequencing Core Facility. The errors have been corrected in the HTML and PDF versions of the article.
References
Burrell, J. R. et al. The frontotemporal dementia-motor neuron disease continuum. Lancet 388, 919–931 (2016).
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006).
Arai, T. et al. TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem. Biophys. Res. Commun. 351, 602–611 (2006).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Benajiba, L. et al. TARDBP mutations in motoneuron disease with frontotemporal lobar degeneration. Ann. Neurol. 65, 470–473 (2009).
Tollervey, J. R. et al. Characterizing the RNA targets and position-dependent splicing regulation by TDP-43. Nat. Neurosci. 14, 452–458 (2011).
Ayala, Y. M. et al. TDP-43 regulates its mRNA levels through a negative feedback loop. EMBO J 30, 277–288 (2011).
Philips, T. & Rothstein, J. D. Rodent models of amyotrophic lateral sclerosis. Curr. Protoc. Pharmacol. 69, 5.67, https://doi.org/10.1002/0471141755.ph0567s69 (2015).
Arnold, E. S. et al. ALS-linked TDP-43 mutations produce aberrant RNA splicing and adult-onset motor neuron disease without aggregation or loss of nuclear TDP-43. Proc. Natl Acad. Sci. USA 110, E736–E745 (2013).
Wu, L. S. et al. TDP-43, a neuro-pathosignature factor, is essential for early mouse embryogenesis. Genesis 48, 56–62 (2010).
Buratti, E. Functional significance of TDP-43 mutations in disease. Adv. Genet. 91, 1–53 (2015).
Sreedharan, J., Neukomm, L. J., Brown, R. H. Jr. & Freeman, M. R. Age-dependent TDP-43-mediated motor neuron degeneration requires GSK3, hat-trick, and xmas-2. Curr. Biol. 25, 2130–2136 (2015).
Johnson, B. S. et al. TDP-43 is intrinsically aggregation-prone, and amyotrophic lateral sclerosis-linked mutations accelerate aggregation and increase toxicity. J. Biol. Chem. 284, 20329–20339 (2009).
Jhuang, H. et al. Automated home-cage behavioural phenotyping of mice. Nat. Commun. 1, 68 (2010).
Borghero, G. et al. Genetic architecture of ALS in Sardinia. Neurobiol. Aging 35, 2882.e2887–2882.e2812 (2014).
Ahmed, R. M. et al. Assessment of eating behavior disturbance and associated neural networks in frontotemporal dementia. JAMA Neurol. 73, 282–290 (2016).
Burden, S. J., Yumoto, N. & Zhang, W. The role of MuSK in synapse formation and neuromuscular disease. Cold Spring Harb. Perspect. Biol 5, a009167 (2013).
Berger, R. et al. Analysis of aldehyde oxidase and xanthine dehydrogenase/oxidase as possible candidate genes for autosomal recessive familial amyotrophic lateral sclerosis. Somat. Cell Mol. Genet. 21, 121–131 (1995).
Garattini, E., Fratelli, M. & Terao, M. The mammalian aldehyde oxidase gene family. Hum. Genomics 4, 119–130 (2009).
Jiang, Y. M. et al. Gene expression profile of spinal motor neurons in sporadic amyotrophic lateral sclerosis. Ann. Neurol. 57, 236–251 (2005).
Kolarcik, C. L. & Bowser, R. Retinoid signaling alterations in amyotrophic lateral sclerosis. Am. J. Neurodegener. Dis. 1, 130–145 (2012).
Mar, A. C. et al. The touchscreen operant platform for assessing executive function in rats and mice. Nat. Protoc. 8, 1985–2005 (2013).
Thomas, A. et al. Marble burying reflects a repetitive and perseverative behavior more than novelty-induced anxiety. Psychopharmacology (Berl.) 204, 361–373 (2009).
Kenna, K. P. et al. NEK1 variants confer susceptibility to amyotrophic lateral sclerosis. Nat. Genet. 48, 1037–1042 (2016).
Brenner, D. et al. NEK1 mutations in familial amyotrophic lateral sclerosis. Brain 139, e28 (2016).
Nihei, K., McKee, A. C. & Kowall, N. W. Patterns of neuronal degeneration in the motor cortex of amyotrophic lateral sclerosis patients. Acta Neuropathol. 86, 55–64 (1993).
Kim, H., Ährlund-Richter, S., Wang, X., Deisseroth, K. & Carlén, M. Prefrontal parvalbumin neurons in control of attention. Cell 164, 208–218 (2016).
Polymenidou, M. et al. Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat. Neurosci. 14, 459–468 (2011).
Hutton, M. et al. Association of missense and 5′-splice-site mutations in tau with the inherited dementia FTDP-17. Nature 393, 702–705 (1998).
Swanson, E. et al. Extracellular tau oligomers induce invasion of endogenous tau into the somatodendritic compartment and axonal transport dysfunction. J. Alzheimers Dis. 58, 803–820 (2017).
Elden, A. C. et al. Ataxin-2 intermediate-length polyglutamine expansions are associated with increased risk for ALS. Nature 466, 1069–1075 (2010).
Becker, L. A. et al. Therapeutic reduction of ataxin-2 extends lifespan and reduces pathology in TDP-43 mice. Nature 544, 367–371 (2017).
Harauz, G. & Boggs, J. M. Myelin management by the 18.5-kDa and 21.5-kDa classic myelin basic protein isoforms. J. Neurochem. 125, 334–361 (2013).
Freischmidt, A. et al. Haploinsufficiency of TBK1 causes familial ALS and fronto-temporal dementia. Nat. Neurosci. 18, 631–636 (2015).
Cirulli, E. T. et al. Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways. Science 347, 1436–1441 (2015).
Skibinski, G. et al. Mutations in the endosomal ESCRTIII-complex subunit CHMP2B in frontotemporal dementia. Nat. Genet. 37, 806–808 (2005).
Takahashi, Y. et al. ERBB4 mutations that disrupt the neuregulin-ErbB4 pathway cause amyotrophic lateral sclerosis type 19. Am. J. Hum. Genet. 93, 900–905 (2013).
Van Hoecke, A. et al. EPHA4 is a disease modifier of amyotrophic lateral sclerosis in animal models and in humans. Nat. Med. 18, 1418–1422 (2012).
Landers, J. E. et al. Reduced expression of the Kinesin-Associated Protein 3 (KIFAP3) gene increases survival in sporadic amyotrophic lateral sclerosis. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.0812937106 (2009).
Nishimura, A. L. et al. Nuclear import impairment causes cytoplasmic trans-activation response DNA-binding protein accumulation and is associated with frontotemporal lobar degeneration. Brain 133, 1763–1771 (2010).
Johnson, J. O. et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat. Neurosci. 17, 664–666 (2014).
Fecto, F. et al. SQSTM1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch. Neurol. 68, 1440–1446 (2011).
Koyama, A. et al. Increased cytoplasmic TARDBP mRNA in affected spinal motor neurons in ALS caused by abnormal autoregulation of TDP-43. Nucleic Acids Res. 44, 5820–5836 (2016).
Egawa, N. et al. Drug screening for ALS using patient-specific induced pluripotent stem cells. Sci. Transl. Med. 4, 145ra104 (2012).
Liu, C., Song, X., Nisbet, R. & Götz, J. Co-immunoprecipitation with tau isoform-specific antibodies reveals distinct protein interactions and highlights a putative role for 2N tau in disease. J. Biol. Chem. 291, 8173–8188 (2016).
Trabzuni, D. et al. MAPT expression and splicing is differentially regulated by brain region: relation to genotype and implication for tauopathies. Hum. Mol. Genet 21, 4094–4103 (2012).
Borroni, B. et al. Association between tau H2 haplotype and age at onset in frontotemporal dementia. Arch. Neurol. 62, 1419–1422 (2005).
Behrouzi, R. et al. Pathological tau deposition in motor neurone disease and frontotemporal lobar degeneration associated with TDP-43 proteinopathy. Acta Neuropathol. Commun. 4, 33 (2016).
Moreno, J. A. et al. Oral treatment targeting the unfolded protein response prevents neurodegeneration and clinical disease in prion-infected mice. Sci. Transl. Med. 5, 206ra138 (2013).
Kang, S. H. et al. Degeneration and impaired regeneration of gray matter oligodendrocytes in amyotrophic lateral sclerosis. Nat. Neurosci. 16, 571–579 (2013).
Zhu, L. J., Holmes, B. R., Aronin, N. & Brodsky, M. H. CRISPRseek: a Bioconductor package to identify target-specific guide RNAs for CRISPR-Cas9 genome-editing systems. PLoS One 9, e108424 (2014).
Heath, C. J., Bussey, T. J. & Saksida, L. M. Motivational assessment of mice using the touchscreen operant testing system: effects of dopaminergic drugs. Psychopharmacology (Berl.) 232, 4043–4057 (2015).
Romberg, C. et al. Depletion of perineuronal nets enhances recognition memory and long-term depression in the perirhinal cortex. J. Neurosci. 33, 7057–7065 (2013).
Wu, L. S., Cheng, W. C. & Shen, C. K. Targeted depletion of TDP-43 expression in the spinal cord motor neurons leads to the development of amyotrophic lateral sclerosis-like phenotypes in mice. J. Biol. Chem. 287, 27335–27344 (2012).
Baghirova, S., Hughes, B. G., Hendzel, M. J. & Schulz, R. Sequential fractionation and isolation of subcellular proteins from tissue or cultured cells. MethodsX 2, 440–445 (2015).
Kalmar, B., Blanco, G. & Greensmith, L. Determination of muscle fiber type in rodents. Curr. Protoc. Mouse Biol. 2, 231–243 (2012).
Acknowledgements
We thank Babraham Institute Experimental Unit staff for technical assistance, the Babraham Institute Imaging Facility and Sequencing Core Facility, A. Weiss for technical assistance at UMMS, M. Brodsky for assistance with CRISPR mutagenesis, the DERC morphology core at UMMS for assistance with histological preparations, and S. Hilton for assistance with OR testing. We thank members of M.P.C.’s laboratory and J.-M. Gallo for discussions. E.K. is supported by a grant from the Korean Health Technology R&D Project, Korea-UK AD Collaborative Project (HI14C2173), Ministry of Health and Welfare, Republic of Korea. R.A is supported by MRC grant MR/L003813/1. S.Y. is supported by an ARUK grant (RF-2016A-1). M.P.C is supported by the van Geest Foundation. R.H.B. gratefully acknowledges support from the ALS Association, Project ALS, Target ALS, ALS-One, ALS Finding A Cure, the Max Rosenfeld fund for ALS Research, and NIH grants RO1NS088689, RO1FD004127, RO1NS065847 and RO1 NS073873. J. Sreedharan is funded by the Motor Neuron Disease Association, the Medical Research Council UK, the Lady Edith Wolfson Fellowship Fund, and the van Geest Foundation.
Author information
Authors and Affiliations
Contributions
J. Sreedharan, M.A.W., M.P.C., R.H.B., T.J.B., J.R.F., R.M. and L.M.S. designed experiments. M.A.W. and J. Sreedharan performed studies on cohort 1 mice including behavioral assessments, histology and transcriptomics. E.K. performed touchscreen studies on cohort 2 mice with assistance from B.U.P. A.D. collated ACBM data and quantified NMJ innervation. R.A. performed spinal cord dissections for laser capture and histology. O.M.P. and J.M. conducted histological studies and image analysis. J. Stephenson performed motor behavioral studies. S.Y. and E.K. performed the OR assay. F.M. quantified motor neurons and western blots. Z.L. performed sequencing to exclude off-target mutagenesis events. S.A. and A.S.-P. assisted with analysis of RNA-seq data and statistical analyses, respectively. R.R.R. performed neuromuscular electrophysiological studies. Y.B. and T.S. developed ACBM software and analyzed ACBM data. J. Sreedharan wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 ACBM walking phenotypes.
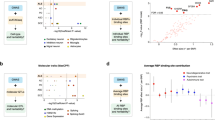
(a) Uncropped agarose gel of genotyping PCR used in figure 1b. (b) Automated continuous behavioral monitoring of walking behavior from 4 to 11.5 months of age separated by sex (n = 5 mice per genotype). Male mice; 4 months: interaction *P = 0.001; 6 months: interaction *P = 0.003; 7.5 months: interaction *P < 0.0001; 10 months: interaction *P < 0.0001; 11.5 months: interaction *P < 0.0001. Female mice; 4 months: interaction *P = 0.034; 6 months: n.s. interaction P = 0.138; 7.5 months: n.s. interaction P = 0.334; 10 months: interaction *P < 0.0001; 11.5 month: interaction *P < 0.0001; repeated measures two−way ANOVA. Error bars represent mean ± s.e.m.
Supplementary Figure 2 Neuromuscular investigations.
(a) Weights of psychology cohort 2 mice (n = 16 wild-type, 12 TDP−43Q331K/+, 14 TDP-43Q331K/Q331K mice). Comparison by repeated measures two−way ANOVA. (b) TDP−43 immunohistochemistry in motor neurons of 5−month−old mice (n = mice per genotype). Representative images shown. Scale bar, 10μm. (c) Representative examples of NMJ immunostaining in gastrocnemius muscles (scale bar, 20μm) with (d) quantification of innervation in wild−type and TDP43Q331K/Q331K mice at 5 months of age (n = 8 mice per genotype). Comparisons: Innervated: P = 0.528 (ns); denervated: P = 0.127 (ns). (e) Innervation/denervation analysis of gastrocnemius muscles in 18 to 23−month−old wild−type and TDP43Q331K/Q331K mice (n = 3 mice per genotype). Comparisons: Innervated: P = 0.678 (ns); denervated: P = 0.801 (ns)⊡ For (d) and (e) unpaired t test with correction using the Holm−Sidak method. (f) Succinate dehydrogenase enzymatic activity in gastrocnemius muscles from 5−month−old mice (n = 8 mice per genotype). Representative images shown. Scale bar, 200 μm. Error bars for (a,d,e) denote s.e.m. (g) Tetanic force responses to repetitive stimulation at 5, 10 and 20Hz from FDB-tibial nerve preparations. Representative wild−type and TDP43Q331K/Q331K mice are shown. (h) Motor unit number estimates in wild−type and TDP43Q331K/Q331K mice based on inspection (Insp) of traces such as those shown in main Fig. 2g, and extrapolation between average motor unit size of the first four units relative to maximum muscle twitch tension (MUNE) (n = 5 wild−type and 5 TDP-43Q331K/Q331K mice). Comparison: Kruskal−Wallis. (i) Maximum tetanic force in TDP43Q331K/Q331K mice compared with controls (50Hz stimulation) (n = 5 wild−τype and 5 TDP−43Q331K/Q331K mice). Comparison: P = 0.150 (ns); two-tailed Mann−Whitney. (j) Tetanus−twitch ratio (expressed as a percentage of maximum force) (n = 4 wild−type and 5 TDP−43Q331K/Q331K mice). Comparison: P > 0.999 (ns); two-tailed Mann−Whitney. (k,l) Fatigue profiles showing decline in maximum tetanic force with repeated stimulation (50Hz for 1 s every 5 s) in representative TDP43Q331K/Q331K and wild−type mouse FDB preparations. (m) Time constant of decay of tetanic force comparing the muscles tested (n = 4 wild−type and 5 TDP−43Q331K/Q331K mice). Comparison: P>0.999 (ns); two-tailed Mann−Whitney. (n,o) Contractile force measurements with progressively graded stimulation (arbitrary units) from representative wild−type and TDP−43Q331K/Q331K mice (n = 3 mice per genotype). Error bars for (h,i,j,m) represent median and interquartile range.
Supplementary Figure 3 Laser capture and RNA sequencing analysis of lumbar spinal motor neurons.
(a) Nissl staining of lumbar spinal cord showing anterior horn motor neurons before and after laser capture (n = 4 wild-type, 4 TDP−43Q331K/+, 4 TDP−43Q331K/Q331K mice). Scale bar, 300 μm. (b) Quality control measures of laser−capture RNA sequencing data. A mean of 49.9 million reads−per−mouse (range 43.7−57.3million) were obtained. Displayed is the percentage of reads that are mapped within genes and exons; reads from ribosomal or mitochondrial RNA; percentage of annotated genes measured and the percentage of reads that are mapped to the sense strand. Colored dots represent individual libraries prepared from each mouse sequenced.(c) Filtering of lumbar spinal cord DESeq2 alternative splice events that are significantly different between wild−type and TDP−43Q331K/Q331K mice. Non−expression hits reflect changes in splice junction usage that exceed a 1.5−fold change relative to expression of the gene from which they are derived. Log Reg hits include splice junctions whose usage changes relative to another junction with the same start or end position. (d) MA plot and (e) hierarchical clustering of alternative splice events in the lumbar spinal cord of wild−type and TDP−43Q331K/Q331K mice (n = 4 wild-type, 4 TDP−43Q331K/+, 4 TDP−43Q331K/Q331K mice). Comparison: DESeq2 wild−type v TDP−43Q331K/Q331K. (f) Immunohistochemistry for AOX1 in lumbar motor neurons of 5−month−old mice (n = 4 wild-type, 4 TDP−43Q331K/Q331K mice). Representative images shown. Scale bar, 100μm.
Supplementary Figure 4 Additional behavioral outcomes from five-choice serial reaction time tasks and marble burying assays.
(a−b) Response latencies. Genotype comparison: P = 0.448 in 6−month−old; P = 0.181 12−month−old mice. (c−d) Reward collection latencies. Genotype comparison: P = 0.662 in 6−month−old; P = 0.821 12−month−old mice. (e−f) Premature responses. Genotype comparison: P = 0.068 in 6−month−old; P = 0.194 12−month−old mice. (g−h) Perseverative responses after correct response. Genotype comparison: P = 0.312 in 6−month−old; P = 0.291 12−month−old mice. At 6 months of age, n = 15 wild−type, 16 TDP−43Q331K/+, 15 TDP−43Q331K/Q331K mice, and at 12 months, n = 15 wild−type, 16 TDP−43Q331K/+, 16 TDP−43Q331K/Q331K mice. For (a−h) mixed−effects model was used and error bars denote s.e.m. (i) Correlation between omissions at 6 and 12 months of age in individual mice (n = 15 wild−type, 16 TDP−43Q331K/+, 15 TDP−43Q331K/Q331K mice). Comparisons: wild-type: P = 0.600 (ns); TDP−43Q331K/+: P = 0.392 (ns); TDP−43Q331K/Q331K: P = 0.046 (*); Pearson’s analysis. (j) Progressive marble burying of cohort 1 mice. Pairwise comparisons: 5 months (n = 19 wild−type, 19 TDP−43Q331K/+, 17 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P = 0.03 (*); wild−type vs. TDP−43Q331K/Q331K: P = 0.013 (*); 8 months (n = 16 wild−type, 11 TDP−43Q331K/+, 15 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P>0.999 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.02 (*); 10 months (n = 16 wild−type, 14 TDP−43Q331K/+, 15 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P = 0.03 (*); wild−type vs. TDP−43Q331K/Q331K: P = 0.03 (*); 12 months (n = 16 wild−type, 13 TDP−43Q331K/+, 15 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P>0.999 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.001 (**); 14 months (n = 15 wild−type, 13 TDP−43Q331K/+, 14 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P = 0.396 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.003 (**); 18 months (n = 15 wild−type, 13 TDP−43Q331K/+, 14 TDP−43Q331K/Q331K mice); wild−type vs. TDP−43Q331K/+: P = 0.009 (**); wild−type vs. TDP−43Q331K/Q331K: P < 0.0001 (****); Kruskal−Wallis followed by Dunn’s test for pairwise comparisons. Error bars for (j) represent median and interquartile range.
Supplementary Figure 5 Frontal cortical histology.
(a) Nissl−stained coronal sections of frontal cortex. Representative images shown. Cortex scale bar, 1 mm; motor, cingulate and somatosensory cortex scale bars, 500 μm; motor cortex layer V scale bar, 500 μm.(b) Quantification of cells in cortical sub regions (n = 5 mice per genotype). Comparisons: Prefrontal: P = 0.477(ns); Motor: P = 0.931(ns); Motor layer V: P = 0.897(ns); Cingulate: P = 0.734(ns); Somatosensory: P = 0.150(ns); multiple t tests with correction using the Holm−Sidak method. (c) Quantification of frontal cortical area in TDP−43Q331K/Q331K mice relative to wild type (n = 6 mice per genotype). Comparison: P = 0.701 (ns); unpaired t test. Error bars denote s.e.m.(d) Uncropped western blots of frontal cortical tissue from four wild−type and four TDP−43Q331K/Q331K mice used in the quantification of TDP−43 in Fig. 4e.
Supplementary Figure 6 Frontal cortical RNA-seq, tau staining and validation in line 3 mice.
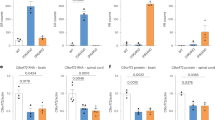
(a) Quality control measures of sequencing reads from 20 frontal cortical libraries from 5 month old mice (coloured dots represent libraries for individual mice). A mean of 58.7m reads−per−mouse (range 42.8−76.3m) were obtained. (b) Filtering of frontal cortex DESeq2 alternative splice events that are significantly different between 5−month−old wild−type and TDP−43Q331K/Q331K mice. Non−expression hits reflect changes in splice junction usage exceeding a 1.5−fold change relative to expression of the gene from which they are derived. Log Reg hits include splice junctions whose usage changes relative to another junction with the same start or end position. (c) Immunostaining for tau in the cortices of 20−month−old mice. Neuronal cells have been stained with NeuN. Representative images shown. Scale bar, 25 μm. (d) Marbles buried by line ♯3 mice (n = 4 wild−type, 8 TDP−43Q331K/+, 5 TDP−43Q331K/Q331K mice). Pairwise comparisons: wild−type vs. TDP−43Q331K/+: P = 0.691 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.020 (*); Kruskal−Wallis followed by Dunn’s test. Error bars represent median and interquartile range. (e) qPCR of expression and splicing changes in line ♯3 mice (n = 5 wild−type, 5 TDP−43Q331K/+, 5 TDP−43Q331K/Q331K mice). Pairwise comparisons: Tardbp expression: wild−type vs. TDP−43Q331K/+: P = 0.076 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.0016 (**); TDP−43Q331K/+ vs. TDP−43Q331K/Q331K: P = 0.036 (*); 0N: wild−type vs. TDP−43Q331K/+: P = 0.072 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.495 (ns); TDP−43Q331K/+ vs. TDP−43Q331K/Q331K: P = 0.03 (*); 2N/0N: wild−type vs. TDP−43Q331K/+: P = 0.877 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.002 (**); TDP−43Q331K/+ vs. TDP−43Q331K/Q331K: P = 0.002 (**); P < 0.0001 (****); one−way ANOVA followed by Holm−Sidak post−hoc tests for pairwise comparisons. Error bars denote s.e.m. (f) Immunohistochemistry for parvalbumin in cortices of line ♯3 mice. Representative images shown. Scale bar, 250μm and quantification of parvalbumin−positive neurons (n = 4 mice per genotype). Comparison: P = 0.006 (**); unpaired t test. Error bars denote s.e.m. (g) Filtering of 5−month−old frontal cortex DESeq2 alternative splice events that are significantly different between MB+ and MB− TDP−43Q331K/Q331K mice. Refer to subfigure (b). (h) Hierarchical clustering of alternative splice events in 5−month−old frontal cortices comparing MB+ and MB− TDP−43Q331K/Q331K mice (n = 6 wild−type, 4 MB+ TDP−43Q331K/Q331K and 4 MB− TDP−43Q331K/Q331K mice); Comparison: DESeq2 MB+ vs MB−.
Supplementary Figure 7 RNA-seq in aged mice.
(a) Quality control measures of sequencing reads from 28 frontal cortical libraries from 20−month−old mice (coloured dots represent libraries for individual mice). (b) Filtering of 20−month−old frontal cortex DESeq2 alternative splice events that are significantly different between wild−type and TDP−43Q331K/Q331K mice. Non−expression hits reflect changes in splice junction usage exceeding a 1.5−fold change relative to expression of the gene from which they are derived. Log Reg hits include splice junctions whose usage changes relative to another junction with the same start or end position. (c) Hierarchical clustering of differentially expressed genes in 20−month−old frontal cortices comparing MB+ and MB− TDP−43Q331K/+ mice (n = 8 wild−type, 5 MB+ TDP−43Q331K/+ and 5 MB− TDP−43Q331K/+ mice); Comparison: DESeq2 MB+ vs MB−. (d) Filtering of 20−month−old frontal cortex DESeq2 alternative splice events that are significantly different between MB+ and MB− TDP−43Q331K/+ mice. Refer to subfigure (b). (e) Hierarchical clustering of alternative splice events in 20−month−old frontal cortices comparing MB+ and MB− TDP−43Q331K/+ mice (n = 8 wild−type, 5 MB+ TDP−43Q331K/+ and 5 MB− TDP−43Q331K/+ mice); Comparison: DESeq2 MB+ vs MB−. (f) qPCR of splicing changes in Matr3 exon 14. Pairwise comparisons: wild−type vs. TDP−43Q331K/+: P = 0.07 (ns); wild−type vs. TDP−43Q331K/Q331K: P = 0.0014 (**); TDP−43Q331K/+ vs. TDP−43Q331K/Q331K: P = 0.033 (*). (g) qPCR of splicing changes in Sqstm1. Pairwise comparisons: wild−type vs. TDP−43Q331K/+: P = 0.003 (*); wild−type vs. TDP−43Q331K/Q331K: P < 0.0001 (****); TDP−43Q331K/+ vs. TDP−43Q331K/Q331K: P = 0.005 (**). (f−g) For qPCR, n = 5 mice per genotype; one−way ANOVA followed by Holm−Sidak post−hoc tests. Error bars denote s.e.m.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 2–6
Supplementary Table 1
RNA sequencing data
Supplementary Table 7
Additional statistical information
Rights and permissions
About this article
Cite this article
White, M.A., Kim, E., Duffy, A. et al. TDP-43 gains function due to perturbed autoregulation in a Tardbp knock-in mouse model of ALS-FTD. Nat Neurosci 21, 552–563 (2018). https://doi.org/10.1038/s41593-018-0113-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0113-5
This article is cited by
-
Reconsidering repurposing: long-term metformin treatment impairs cognition in Alzheimer’s model mice
Translational Psychiatry (2024)
-
Opinion: more mouse models and more translation needed for ALS
Molecular Neurodegeneration (2023)
-
Specific vulnerability of iPSC-derived motor neurons with TDP-43 gene mutation to oxidative stress
Molecular Brain (2023)
-
Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Nature Communications (2023)
-
Loss of TDP-43 function underlies hippocampal and cortical synaptic deficits in TDP-43 proteinopathies
Molecular Psychiatry (2023)