Abstract
In Huntington’s disease (HD), expansion of CAG codons in the huntingtin gene (HTT) leads to the aberrant formation of protein aggregates and the differential degeneration of striatal medium spiny neurons (MSNs). Modeling HD using patient-specific MSNs has been challenging, as neurons differentiated from induced pluripotent stem cells are free of aggregates and lack an overt cell death phenotype. Here we generated MSNs from HD patient fibroblasts through microRNA-based direct neuronal conversion, bypassing the induction of pluripotency and retaining age signatures of the original fibroblasts. We found that patient MSNs consistently exhibited mutant HTT (mHTT) aggregates, mHTT-dependent DNA damage, mitochondrial dysfunction and spontaneous degeneration in culture over time. We further provide evidence that erasure of age stored in starting fibroblasts or neuronal conversion of presymptomatic HD patient fibroblasts results in differential manifestation of cellular phenotypes associated with HD, highlighting the importance of age in modeling late-onset neurological disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
02 September 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Gusella, J. F. et al. A polymorphic DNA marker genetically linked to Huntington’s disease. Nature 306, 234–238 (1983).
The Huntington’s Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 (1993).
Ross, C. A. et al. Huntington disease: natural history, biomarkers and prospects for therapeutics. Nat. Rev. Neurol. 10, 204–216 (2014).
Kremer, B. et al. A worldwide study of the Huntington’s disease mutation. The sensitivity and specificity of measuring CAG repeats. N. Engl. J. Med. 330, 1401–1406 (1994).
Brinkman, R. R., Mezei, M. M., Theilmann, J., Almqvist, E. & Hayden, M. R. The likelihood of being affected with Huntington disease by a particular age, for a specific CAG size. Am. J. Hum. Genet. 60, 1202–1210 (1997).
Arrasate, M., Mitra, S., Schweitzer, E. S., Segal, M. R. & Finkbeiner, S. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004).
Vonsattel, J. P. & DiFiglia, M. Huntington disease. J. Neuropathol. Exp. Neurol. 57, 369–384 (1998).
Arber, C. et al. Activin A directs striatal projection neuron differentiation of human pluripotent stem cells. Development 142, 1375–1386 (2015).
Camnasio, S. et al. The first reported generation of several induced pluripotent stem cell lines from homozygous and heterozygous Huntington’s disease patients demonstrates mutation related enhanced lysosomal activity. Neurobiol. Dis. 46, 41–51 (2012).
An, M. C. et al. Genetic correction of Huntington’s disease phenotypes in induced pluripotent stem cells. Cell. Stem Cell. 11, 253–263 (2012).
HD iPSC Consortium. Induced pluripotent stem cells from patients with Huntington’s disease show CAG-repeat-expansion-associated phenotypes. Cell. Stem Cell. 11, 264–278 (2012).
Jeon, I. et al. Neuronal properties, in vivo effects, and pathology of a Huntington’s disease patient-derived induced pluripotent stem cells. Stem Cells 30, 2054–2062 (2012).
Zhang, N., An, M. C., Montoro, D. & Ellerby, L. M. Characterization of human Huntington’s disease cell model from induced pluripotent stem cells. PLoS. Curr. 2, RRN1193 (2010).
Nekrasov, E. D. et al. Manifestation of Huntington’s disease pathology in human induced pluripotent stem cell-derived neurons. Mol. Neurodegener. 11, 27 (2016).
Yoo, A. S., Staahl, B. T., Chen, L. & Crabtree, G. R. MicroRNA-mediated switching of chromatin-remodelling complexes in neural development. Nature 460, 642–646 (2009).
Abernathy, D. G. et al. MicroRNAs induce a permissive chromatin environment that enables neuronal subtype-specific reprogramming of adult human fibroblasts. Cell. Stem Cell. 21, 332–348 (2017).
Victor, M. B. et al. Generation of human striatal neurons by microRNA-dependent direct conversion of fibroblasts. Neuron 84, 311–323 (2014).
Miller, J. D. et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell. Stem Cell. 13, 691–705 (2013).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Huh, C. J. et al. Maintenance of age in human neurons generated by microRNA-based neuronal conversion of fibroblasts. Elife 5, e18648 (2016).
Mertens, J. et al. Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell. Stem Cell. 17, 705–718 (2015).
Silvestroni, A., Faull, R. L., Strand, A. D. & Möller, T. Distinct neuroinflammatory profile in post-mortem human Huntington’s disease. Neuroreport 20, 1098–1103 (2009).
Xue, M. et al. Contributions of multiple proteases to neurotoxicity in a mouse model of intracerebral haemorrhage. Brain 132, 26–36 (2009).
Li, S. H. et al. Lack of huntingtin-associated protein-1 causes neuronal death resembling hypothalamic degeneration in Huntington’s disease. J. Neurosci. 23, 6956–6964 (2003).
Lee, J. H. et al. Reinstating aberrant mTORC1 activity in Huntington’s disease mice improves disease phenotypes. Neuron 85, 303–315 (2015).
Valenza, M. et al. Dysfunction of the cholesterol biosynthetic pathway in Huntington’s disease. J. Neurosci. 25, 9932–9939 (2005).
Tomás-Zapico, C. et al. α-Synuclein accumulates in huntingtin inclusions but forms independent filaments and its deficiency attenuates early phenotype in a mouse model of Huntington’s disease. Hum. Mol. Genet. 21, 495–510 (2012).
Corrochano, S. et al. α-Synuclein levels modulate Huntington’s disease in mice. Hum. Mol. Genet. 21, 485–494 (2012).
Strand, A. D. et al. Expression profiling of Huntington’s disease models suggests that brain-derived neurotrophic factor depletion plays a major role in striatal degeneration. J. Neurosci. 27, 11758–11768 (2007).
Zuccato, C. & Cattaneo, E. Role of brain-derived neurotrophic factor in Huntington’s disease. Prog. Neurobiol. 81, 294–330 (2007).
Plotkin, J. L. et al. Impaired TrkB receptor signaling underlies corticostriatal dysfunction in Huntington’s disease. Neuron 83, 178–188 (2014).
Zhang, Q. et al. The zinc finger transcription factor Sp9 is required for the development of striatopallidal projection neurons. Cell. Rep. 16, 1431–1444 (2016).
Gutekunst, C. A. et al. Nuclear and neuropil aggregates in Huntington’s disease: relationship to neuropathology. J. Neurosci. 19, 2522–2534 (1999).
Ko, J., Ou, S. & Patterson, P. H. New anti-huntingtin monoclonal antibodies: implications for huntingtin conformation and its binding proteins. Brain Res. Bull. 56, 319–329 (2001).
Zheng, S. et al. Deletion of the huntingtin polyglutamine stretch enhances neuronal autophagy and longevity in mice. PLoS. Genet. 6, e1000838 (2010).
Taipale, M., Jarosz, D. F. & Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat. Rev. Mol. Cell. Biol. 11, 515–528 (2010).
Gomez-Pastor, R. et al. Abnormal degradation of the neuronal stress-protective transcription factor HSF1 in Huntington’s disease. Nat. Commun. 8, 14405 (2017).
Vilchez, D. et al. Increased proteasome activity in human embryonic stem cells is regulated by PSMD11. Nature 489, 304–308 (2012).
Lu, X. H. et al. Targeting ATM ameliorates mutant Huntingtin toxicity in cell and animal models of Huntington’s disease. Sci. Transl. Med. 6, 268ra178 (2014).
Goebel, H. H., Heipertz, R., Scholz, W., Iqbal, K. & Tellez-Nagel, I. Juvenile Huntington chorea: clinical, ultrastructural, and biochemical studies. Neurology 28, 23–31 (1978).
Kim, I., Rodriguez-Enriquez, S. & Lemasters, J. J. Selective degradation of mitochondria by mitophagy. Arch. Biochem. Biophys. 462, 245–253 (2007).
Liu, L. et al. Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160, 177–190 (2015).
Kumar, A. & Ratan, R. R. Oxidative stress and Huntington’s disease: the good, the bad, and the ugly. J. Huntingt. Dis. 5, 217–237 (2016).
Vonsattel, J. P. et al. Neuropathological classification of Huntington’s disease. J. Neuropathol. Exp. Neurol. 44, 559–577 (1985).
Yoo, A. S. et al. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature 476, 228–231 (2011).
Dragunow, M. et al. In situ evidence for DNA fragmentation in Huntington’s disease striatum and Alzheimer’s disease temporal lobes. Neuroreport 6, 1053–1057 (1995).
Richner, M., Victor, M. B., Liu, Y., Abernathy, D. & Yoo, A. S. MicroRNA-based conversion of human fibroblasts into striatal medium spiny neurons. Nat. Protoc. 10, 1543–1555 (2015).
Lobo, M. K., Karsten, S. L., Gray, M., Geschwind, D. H. & Yang, X. W. FACS-array profiling of striatal projection neuron subtypes in juvenile and adult mouse brains. Nat. Neurosci. 9, 443–452 (2006).
Acknowledgements
The authors thank A. Bowman and L. Solnica-Krezel for suggestions, B. Steger and J. Peyer for data quantification, the Washington University Center for Cellular Imaging (WUCCI) for their help in generating electron microscopy data, the Genome Technology Access Center (GTAC) for generating transcriptome datasets, and the Core Usage Funding Program from the Institute of Clinical and Translational Services (ICTS) and the Genome Engineering and iPSC Center (GEiC) at Washington University School of Medicine for their assistance in generating and characterizing iPSC lines. M.B.V. is supported by a National Science Foundation Graduate Research Fellowship (DGE-1143954) and a NIH/NIA dissertation award (1R36AG053444-01). A.S.Y. is supported by the Andrew B. and Virginia C. Craig Faculty Fellowship endowment, an NIH Director’s Innovator Award (DP2NS083372-01), a Seed Grant from Washington University Center of Regenerative Medicine, the Ellison Medical Foundation New Scholar in Aging Award, Cure Alzheimer’s Fund (CAF) and a Presidential Early Career Award for Scientists and Engineers (PECASE) (4DP2NS083372-02).
Author information
Authors and Affiliations
Contributions
M.B.V., M.R. and A.S.Y. designed experiments and wrote the manuscript. M.B.V., M.R. and H.E.O performed the experiments and analyzed data. H.E.O. edited the manuscript. C.M. and B.Z. aligned and analyzed the genomic data. C.J.H. contributed to the microarray data analysis and interpretation. S.W.L. contributed to the supplementary data. X.W.Y. contributed to data interpretation and conceived the experiments with the small-molecule ATM-kinase inhibitor. A.M.M. and B.L.D. made and pseudotyped AVV-HTT shRNA and control viruses and conceived the related experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Neuronal reprogramming independent of donor’s age or CAG-repeat size and DARPP-32 antibody validation.
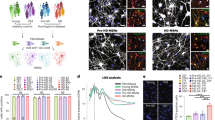
Additional HD lines carrying large number of CAG repeats can be reprogrammed into functional neurons by miR-9/9*-124+CDM, and DARPP-32 (Santa Cruz - H62 Clone) antibody shows specificity to striatum in both mouse and human striatal sections. (a), Electrophysiolocal properties were analyzed in monoculture free of rat or mouse primary glia/neurons. HD.59 was analyzed at PID 23 while HD.180 was analyzed at PID 30. Experiment was repeated independently over 3 times (b), RNA-seq analysis at PID 32 of adult control and HD patient fibroblasts reprogrammed with miR-9/9*-124+CDM reveals expression of the full length DARPP-32 transcript. Representative tracks from RNA-seq analysis of 7 HD-MSNs and 5 controls. (c), Immunostaining with an additional anti-DARPP-32 antibody (abcam; ab40801) also produced positive cells, although with less intensity (Cells depicted were reprogrammed from GM04855 fibroblasts). (d), We further validated the up-regulation of DARPP-32 by qPCR with DARPP-32 specific probes. n = 3 technical replicates from MSNs reprogrammed from a single HD patient. Mean and S.E.M. (e), DARPP-32 (Santa Cruz - H62 Clone) showed specificity to MSNs in the striatum of an adult mouse brain as seen by immunohistochemistry analysis, where only the striatum is labeled (shown in red) in a brain coronal section. Similarly, immunohistochemistry performed in a human postmortem brain section of globus pallidus with putamen of a 89 year old healthy female obtained from NIH NeuroBioBank shows antibody labeling specificity to the striatum. Experiments in C,D and E were only performed once.
Supplementary Figure 2 CAG-sizing of primary fibroblasts and microRNA-derived MSNs.
CAG repeat analysis confirmed HTT mutation and number of CAGs in cell lines mainly used in this study. After several passages in culture and subsequent reprogramming into MSNs by miR-9/9*-124+CDM for three weeks, CAG size was stable (for each group non-transduced fibroblasts are shown on the left and reprogrammed MSNs on the right).
Supplementary Figure 3 Reprogramming HD patient fibroblasts generates functional neurons.
Reprogramming HD patient fibroblasts generates functional neurons. (a), GFP labeled Ctrl-MSNs and tRFP labeled HD-MSNs (pseudo-colored gray) co-cultured and seeded atop rat primary neural cells for whole-cell recording. (b), Representative traces from current-clamp recordings of Ctrl-MSNs (green traces) at PID 35, and (c), HD-MSNs (gray traces); Ctrl-MSNs and HD-MSNs displayed similar firing patterns, with HD-MSNs having a greater percentage of cells that fired multiple action potentials. Inset display single trace at increasing stimulus steps and total number of cells that fired single or multiple action potentials. Voltage-clamp recordings demonstrate inward sodium and outward calcium currents typical of neurons. (d), HD-MSNs displayed spontaneous action potentials. (e), I-V curve. n = 11 from 3 control samples; n = 13 from 3 HD samples. Evoked response at 90 mV in HD-MSNs is significantly higher. One-way ANOVA (F39,440 = 24.21 P < 0.001) with post hoc Tukey’s test; ** p = 0.0068; Mean ± S.D. (f), Analysis of passive membrane properties. n = 3 averages for each line, totaling 18 controls and 20 HD-MSNs. Experiment was repeated independently twice. Mean ± s.e.m. Two-tailed unpaired t-test; p-value > 0.05 df = 4 Scale bar in a, 10 μm. (g), All recorded properties during electrophysiological analysis displayed for each reprogrammed line.
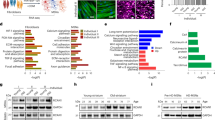
Supplementary Figure 4 Collection of Huntington’s disease–associated genes differentially expressed in HD-MSNs and validated by qPCR.
(a) Ingenuity pathway analysis (IPA) of differentially expressed genes identified in RNA-seq studies uncovered many genes that have been experimentally associated with HD. Genes shown in subcellular organization. Red genes are upregulated while green genes are downregulated in HD-MSNs, with darker colors representing higher expression levels. (b) qPCR Validation of Differentially Expressed Genes Identified in RNA-seq Analysis. Ctrl: MSNs converted from control fibroblasts. HD: MSNs converted from Huntington’s disease patient fibroblasts. A number of genes detected to be differentially expressed between HD and Ctrl MSNs were verified by qPCR. BDNF was not detected to be differentially expressed in RNA-seq analysis, and confirmed to be unchanged by qPCR, while MMP9 was detected to be differentially expressed between HD and Ctrls, but shows only a trend by qPCR analysis. Mean ± s.d.; n = 3 biological replicates per group. Two-tailed student’s t-test; df = 4; TRKB ***P = 7.3E-5; SP9 **P = 0.0025; HAP1 *P = 0.033; SCNA *P = 0.034; KCNA4 ***P = 0.0009; AEN **P = 0.0042; ITIH5 *P = 0.0468; BDNF n.s. = 0.68; MMP9 n.s. = 0.11; n.s. = not significant.
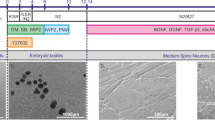
Supplementary Figure 5 HD fibroblasts do not exhibit inclusion bodies, even upon cellular insult.
HD fibroblasts do not exhibit inclusion bodies, even upon cellular insults. (a), Fibroblasts induced to age in vitro by serial passaging (18 times), forced to exit cell cycle by contact inhibition, and then cultured for an additional 7 weeks, do not exhibit inclusion bodies. (b), Ctrl (Ctrl.19) or HD fibroblasts (HD.40) challenged with 1 mM H2O2 to induce oxidative stress do not exhibit inclusion bodies. (c), HD.40 fibroblasts transduced with CDM and a non-specific microRNA (miR-N.S.) to mimic reprogramming conditions but not neuronal induction, do not form inclusion bodies. The formation of inclusion bodies is present in all three lines reprogrammed from HD patients. (d), All three HD MSNs lines examined exhibit aggregated HTT inclusions (IBs) at post-induction day 30 (PID) analyzed by MW8 immunostaining. (e), Quantification with EM48 yields similar number of cells displaying inclusions to MW8 staining; Two-tailed student’s t-test, n = 4 biological replicates per group; ***P = 0.0003 df = 6. (f), The appearance of aggregated cytoplasmic HTT protein in HD.40 MSNs is detected as early as PID 14. By PID 21 HTT inclusions are numerous and after PID 28, inclusions are bigger and more defined, with little to no granules. Quantification of inclusion formation shows significant changes by PID 14; n = 3 biological replicates with each sample containing approximately 100 cells; Mean ± s.e.m. One-Way ANOVA (F6,17 = 65.47, P< 0.0001) with post hoc Tukey’s test (from left, n.s. = 0.19, **P = 0.0028, ***P = 3.2E-10. (f), Time course analysis with Ctrl.20 and HD.42 for the appearance of oxidative DNA damage phenotype by 8OH-dG staining. Significant differences between controls and HD MSNs is detected as early as PID 20, and continue to augment with time in culture. One-Way ANOVA (F9,190 = 30.86, P < 0.0001) with post hoc Tukey’s test (from left, n.s. = 0.99, n.s. = 0.34, **P = 0.0012, ***P = 5.8E-11, ***P = 1.0E-14; ***P < 0.001; **P < 0.01; n.s. = not significant. n = 20 cells per time point for each line. Mean ± s.d. a-c experiments were repeated independently twice and d, over 3 times. f-g experiments have not been independently repeated.
Supplementary Figure 6 Ultrastructural analysis of HD-MSNs.
(a), Immunogold labeling of HTT in HD-MSNs shows fibrillar-like structures. (b), Immunogold labeling of HTT in HD.40-MSNs at PID 21 is prominent within single and double-membrane autophagosome-like structures (black arrowheads), as well as accumulated as non-membrane bound cytoplasmic structures (red arrowheads). In addition there is marked presence of lipofuscin granules which are known to accumulate with aging (labeled with an asterisk) and quantified in (c), for 3 independent control and HD lines. Two-tailed student’s t-test, n = average of all visible lipofuscins in 10 cells per line; * = p-value = 0.0172 t = 3.925 df = 4. (d), Greater magnification of red arrows in (b), where fibrilar-like structures can be seen. (e), Ultrastructural analysis in HD.40 was also marked by mitophagy (left), accumulation of lipid droplets (middle) and swollen mitochondria typical of apoptotic cells (right); N = nucleus. (f), Colocalization of HTT (EM48) and the autophagosome marker LC3 in additional HD lines. a-f Experiments were repeated in 3 pairs of HD and control MSNs and independently repeated twice.
Supplementary Figure 7 Biochemical analysis of mutant HTT expression in reprogrammed MSNs.
(a), Four samples used for western blotting, all reprogrammed with miR-9/9*-124+CDM and lysed for protein extraction at post-infection day (PID) 28. (b) Anti-huntingtin monoclonal antibody MW1 specifically binds to the polyglutamine domain of HTT exon 1 and therefore recognizes expanded polyglutamine while showing no detectable binding to normal HTT. Our analysis confirms the expression of soluble mutant HTT in MSNs reprogrammed from primary fibroblasts samples from HD patients. (c) Unlike MW1, the monoclonal anti-huntingtin antibody MW8 recognizes amino acids 83-90 near the c terminus of exon 1 of HTT and specifically recognizes aggregated forms of mutant HTT. Our analysis with MW8 reveals detectable levels of insoluble aggregated HTT in HD-MSNs. These experiments were repeated independently twice.
Supplementary Figure 8 Adult HD fibroblasts can be induced to pluripotency and rederived to human embryonic fibroblasts (HEFs).
(a), Schematic of HEF derivation. (b), Cells transduced with OCT4, SOX2, KLF4 and c-MYC (OSKM) express stem cell markers, (c), and retain a normal karyotype. (d), Induced pluripotent stem cells (iPSCs) can be differentiated into HEFs by addition of 20% fetal bovine serum (FBS) to culture media and passaging at least three times. (e), CAG sizing confirms that HEFs retain repeat number. (f), LAP2α levels are restored in HEFs; n = 1,000 cells from 3 independent experiments; Two-tailed student’s t-test p-value = 0.0230; df = 4 (g), MSNs directly converted from HD.40 FBs and HEFs express TUBB3, while heMSNs are nearly devoid of mHTT aggregates (MW8) at PID 21. (h), Quantification of mHTT aggregates in MSNs versus heMSNs of HD.50 in comparison to Ctrl.17 ; n = Average of 200 cells from 3 biological replicates; One-way ANOVA (F2,6 = 21.85 P = 0.0018) followed by Tukey’s test (P** = 0.0112, P* = 0.0016, n.s. = 0.16); Mean ± s.e.m; **P < 0.01; *P < 0.05. Experiments in b, c, e, were not repeatedly independently, while experiments shown in d, f and g were repeated independently over 3 times.
Supplementary Figure 9 Induction of pluripotency alters expression of ubiquitin–proteasome system (UPS)-related genes and also differs in MSNs reprogrammed from young or old fibroblast donors.
(a) HD.40 fibroblasts (HD-FB) and two iPSC clones derived from HD.40 fibroblasts were differentiated into embryonic fibroblasts (HD-HEF.1 and HD-HEF.2) and analyzed by qPCR for the expression of UPS-related genes. HD-HEFs have higher expression of many UPS genes. Mean ± s.e.m.; One-Way ANOVA (F50,102 = 20.17, P< 0.0001) with Holm-Sidak correction. n = 3 samples per group. PSMB6 *P = 0.017, UBE3A ***P = 1.2E-10, HSF1 *P = 0.038, HSPA8 ***P = 4.4E-14, DNAJB2 *P = 0.014, HSPA5 ***P = 5.1E-7, HSPB1 ***P = 7.5E-10. Only significant changes (P < 0.05) are marked, and represents p-values consistent for both HD-FB versus HD-HEF.1 or HD-HEF.2 tests. The expression of ATG12 was only significantly different in HEF2 (**P = 0.002). (b) Ubiquitin-proteosome system (UPS)-related genes differentially expressed from young (three days, five months and one year old) or old (aged 90, 92, and 92 years old) miR-9/9*-124+CDM reprogrammed MSNs by microarray analysis.
Supplementary Figure 10 Spontaneous degeneration in culture is associated with loss of DARPP-32-positive neurons and can be attenuated upon ATM inhibition.
Since major cell loss only occurs past PID 35, the levels of TUBB3-positive and DARPP-32-positive cells were quantified at PID 30 and PID 40 to determine extent to which DARPP-32-positive cells degenerate in culture. (a), Ctrl.17c and HD.42 fibroblasts were transduced with miR-9/9*-124+CDM concurrently and immunostained at PID 30 and at PID 40. At PID 35 cells were confirmed to have altered levels of cell death (data are quantified and shown in the Fig. 8 of the main text). Non-transduced fibroblasts were used as a negative control for immunostaining. (b), From PID 30 to PID 40, there are no changes observed in the number of TUBB3-positive cells in control samples, in contrast to HD samples in which the level of TUBB3-positive cells is dramatically reduced One-Way ANOVA (F3,8 = 32.25, P = 8.1E-5) with post hoc Tukey’s test (from left, n.s = 0.35, ***P = 0.0004, ***P = 0.0002). In addition, while the percent of DARPP-32-positive cells remains unchanged in control MSNs from PID 30 to PID 40, this percentage is reduced in HD samples at PID 40 in comparison to control. One-Way ANOVA (F3,8 = 5.211, P = 0.0276) with post hoc Tukey’s test (from left, n.s. = 0.52, *P = 0.022). n = averages from 100 cells from 3 random fields-of-view from 3 biological replicates for each time point. (c),Treatment with KU60019 reduces cell death levels at PID 35; One-Way ANOVA (F3,31 = 13.28, P = 9.5E-6) with post hoc Tukey’s test (***P = 6.5E-6, *p = 0.019, n.s. = 0.54). (d), KU60019 protects HD-MSNs against H2O2-induced oxidative stress; One-Way ANOVA (F3,31 = 12.58, P = 1.5E-5) with post hoc Tukey’s test (***P = 2.9E-6, *p = 0.048, n.s. = 0.14). n = averages from 1,000 cells from 8 or 9 biological replicates; Scale bars: 100 μm. ***P < 0.001; **P < 0.01; *P < 0.05; n.s. = not significant. Mean ± s.e.m.
Supplementary Figure 11 Evidence of increased mitophagy in HD-MSNs.
Evidence of increased mitophagy in HD-MSNs. (a), Additional HD and control lines used in this study stain positive for TUBB3 and successfully undergo direct conversion by miR-9/9*-124+CDM. Experiment has been repeated independently 3 times. (b) LC3-II staining at multiple intervals during direct conversion. At PID 20, analysis of 3 independent HD and control lines shows that 2 out of 3 HD-MSNs lines have a higher number of autophagosomes, however the overall change was not significantly different. Two-tailed student’s t-test; t = 1.31 df = 4 p = 0.2604; n = averages from 10 cells per group. (c) Co-staining of MitoTracker Red and LC3-II at PID 40 in two pairs of HD and control lines shows higher percentage of colocalization in HD-MSNs. One-Way ANOVA (F3,8 = 61.48, P = 7.2E-6) with post hoc Tukey’s test (from left, ***P = 2E-4, 5.7E-6, 8E-5, *P = 0.014); n = Averages from approximately 2,000 LC3-II puncta from 3 random field-of-views per group. Colocalization was measured using images acquired through confocal microscopy at 100x objective and analyzed post-acquisition by automated colocalization analysis. One-Way ANOVA with post hoc Tukey’s test; ***P < 0.001; **P < 0.01; *P < 0.05; n.s. = not significant. Mean ± s.e.m.
Supplementary Figure 12 Differential vulnerability to degeneration in distinct subtypes of HD neurons.
(a), HD.40 fibroblasts were reprogrammed into cortical-like neurons (CNs) with miR-9/9*-124+DAM (NeuroD2, ASCL1 and Myt1L) (b), HD.40 CNs immunostained with TUBB3 at PID 21. This experiment was repeated independently over 3 times. (c), qPCR analysis for the expression of cortical genes as well as MSN-marker DARPP-32 at PID 30; n = 3 samples per group; Two-tailed student’s t-test with Holm-Sidak correction (from left, ***P = 0.003, 0.001, 0.0016,*P = 0.037, 0.043, 0.043, df = 4). (d), Comet assay; One-Way ANOVA (F3,195 = 20.15, P = 2E-11) with post hoc Tukey’s test (from left, ***P = 4.4E-8, 7.1E-11, n.s. = 0.14); n = 50 cells per group. (e), Immunostaining and quantification of the DNA damage marker γH2AX; One-Way ANOVA (F3,8 = 18.23, P = 0.0006) with post hoc Tukey’s test (**P = 0.0074, ***P = 0.001, n.s. = 0.99); n = averages from 500 cells per group in 3 independent experiments. (f), SYTOX staining reveals differences in subtype-dependent cell death levels at PID 35; n = averages from 1,200 cells per group in 3 independent experiments; One-Way ANOVA (F3,8 = 29.2, P = 0.0001) with post hoc Tukey’s test (**P = 0.002, ***P = 0.0001, n.s. = 0.18). (g), HTT aggregation detected by MW8 antibody at PID 35 in HD-MSNs and HD-CNs and quantification of the percentage of cells with IBs in MSNs and CNs. n = averages from 100 cells per group in 3 independent experiments; One-Way ANOVA (F3,8 = 58.75, P = 8.5E-6) with post hoc Tukey’s test (from left, **P = 0.001 and 0.002, ***P = 2.1E-5). ***P < 0.001; **P < 0.01. Mean ± s.e.m. Scale bar in b,d,f, 100 μm; and in e, 20 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Victor, M.B., Richner, M., Olsen, H.E. et al. Striatal neurons directly converted from Huntington’s disease patient fibroblasts recapitulate age-associated disease phenotypes. Nat Neurosci 21, 341–352 (2018). https://doi.org/10.1038/s41593-018-0075-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0075-7
This article is cited by
-
Neuropathogenesis-on-chips for neurodegenerative diseases
Nature Communications (2024)
-
Cellular reprogramming as a tool to model human aging in a dish
Nature Communications (2024)
-
Molecular and cellular mechanisms of selective vulnerability in neurodegenerative diseases
Nature Reviews Neuroscience (2024)
-
Neural cell state shifts and fate loss in ageing and age-related diseases
Nature Reviews Neurology (2023)
-
Longitudinal modeling of human neuronal aging reveals the contribution of the RCAN1–TFEB pathway to Huntington’s disease neurodegeneration
Nature Aging (2023)