Abstract
Despite rapid progresses in the genome-editing field, in vivo simultaneous overexpression of multiple genes remains challenging. We generated a transgenic mouse using an improved dCas9 system that enables simultaneous and precise in vivo transcriptional activation of multiple genes and long noncoding RNAs in the nervous system. As proof of concept, we were able to use targeted activation of endogenous neurogenic genes in these transgenic mice to directly and efficiently convert astrocytes into functional neurons in vivo. This system provides a flexible and rapid screening platform for studying complex gene networks and gain-of-function phenotypes in the mammalian brain.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wu, Z., Yang, H. & Colosi, P. Effect of genome size on AAV vector packaging. Mol. Ther. 18, 80–86, https://doi.org/10.1038/mt.2009.255 (2010).
Shechner, D. M., Hacisuleyman, E., Younger, S. T. & Rinn, J. L. Multiplexable, locus-specific targeting of long RNAs with CRISPR-Display. Nat. Methods 12, 664–670, https://doi.org/10.1038/Nmeth.3433 (2015).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588, https://doi.org/10.1038/nature14136 (2015).
Zetsche, B., Volz, S. E. & Zhang, F. A split-Cas9 architecture for inducible genome editing and transcription modulation. Nat. Biotechnol. 33, 139–142 (2015).
Dahlman, J. E. et al. Orthogonal gene knockout and activation with a catalytically active Cas9 nuclease. Nat. Biotechnol. 34, 441–441 (2016).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183, https://doi.org/10.1016/j.cell.2013.02.022 (2013).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451, https://doi.org/10.1016/j.cell.2013.06.044 (2013).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646, https://doi.org/10.1016/j.cell.2014.09.039 (2014).
Perez-Pinera, P. et al. RNA-guided gene activation by CRISPR-Cas9-based transcription factors. Nat. Methods 10, 973–976, https://doi.org/10.1038/Nmeth.2600 (2013).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661, https://doi.org/10.1016/j.cell.2014.09.029 (2014).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350, https://doi.org/10.1016/j.cell.2014.11.052 (2015).
Polstein, L. R. & Gersbach, C. A. A light-inducible CRISPR-Cas9 system for control of endogenous gene activation. Nat. Chem. Biol. 11, 198–200, https://doi.org/10.1038/nchembio.1753 (2015).
Cheng, A. W. et al. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell. Res. 23, 1163–1171, https://doi.org/10.1038/cr.2013.122 (2013).
Gao, Y. et al. Complex transcriptional modulation with orthogonal and inducible dCas9 regulators. Nat. Methods 13, 1043–1049, https://doi.org/10.1038/nmeth.4042 (2016).
Maeder, M. L. et al. CRISPR RNA-guided activation of endogenous human genes. Nat. Methods 10, 977–979, https://doi.org/10.1038/Nmeth.2598 (2013).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucleic Acids Res. 41, 7429–7437, https://doi.org/10.1093/nar/gkt520 (2013).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838, https://doi.org/10.1038/nbt.2675 (2013).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328, https://doi.org/10.1038/Nmeth.3312 (2015).
Chavez, A. et al. Comparison of Cas9 activators in multiple species. Nat. Methods 13, 563–567, https://doi.org/10.1038/Nmeth.3871 (2016).
Black, J. B. et al. Targeted epigenetic remodeling of endogenous loci by CRISPR/Cas9-based transcriptional activators directly converts fibroblasts to neuronal cells. Cell. Stem Cell. 19, 406–414, https://doi.org/10.1016/j.stem.2016.07.001 (2016).
Nihongaki, Y. et al. CRISPR-Cas9-based photoactivatable transcription systems to induce neuronal differentiation. Nat. Methods 14, 963–966, https://doi.org/10.1038/nmeth.4430 (2017).
Liu, Y. et al. Ascl1 converts dorsal midbrain astrocytes into functional neurons in vivo. J. Neurosci. 35, 9336–9355, doi:https://doi.org/10.1523/Jneurosci.3975-14.2015 (2015).
Heinrich, C. et al. Directing astroglia from the cerebral cortex into subtype specific functional neurons. PLoS. Biol. 8, e1000373, https://doi.org/10.1371/journal.pbio.1000373 (2010).
Cheng, L. et al. Direct conversion of astrocytes into neuronal cells by drug cocktail. Cell. Res. 25, 1269–1272, https://doi.org/10.1038/cr.2015.120 (2015).
Davis, R. L., Weintraub, H. & Lassar, A. B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987–1000, doi:https://doi.org/10.1016/0092-8674(87)90585-X (1987).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676, https://doi.org/10.1016/j.cell.2006.07.024 (2006).
Mertens, J., Marchetto, M. C., Bardy, C. & Gage, F. H. Evaluating cell reprogramming, differentiation and conversion technologies in neuroscience. Nat. Rev. Neurosci. 17, 424–437, https://doi.org/10.1038/nrn.2016.46 (2016).
Hilton, I. B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517, https://doi.org/10.1038/nbt.3199 (2015).
Liu, X. S. et al. Editing DNA methylation in the mammalian genome. Cell 167, 233–247.e17, https://doi.org/10.1016/j.cell.2016.08.056 (2016).
Farzadfard, F., Perli, S. D. & Lu, T. K. Tunable and multifunctional eukaryotic transcription factors based on CRISPR/Cas. ACS Synth. Biol. 2, 604–613, https://doi.org/10.1021/sb400081r (2013).
Han, K. et al. SHANK3 overexpression causes manic-like behavior with unique pharmacogenetic properties. Nature 503, 72–77, https://doi.org/10.1038/nature12630 (2013).
Lombardi, L. M., Baker, S. A. & Zoghbi, H. Y. MECP2 disorders: from the clinic to mice and back. J. Clin. Invest. 125, 2914–2923, https://doi.org/10.1172/JCI78167 (2015).
Collins, A. L. et al. Mild overexpression of MeCP2 causes a progressive neurological disorder in mice. Hum. Mol. Genet. 13, 2679–2689, https://doi.org/10.1093/hmg/ddh282 (2004).
Joung, J. et al. Genome-scale activation screen identifies a lncRNA locus regulating a gene neighborhood. Nature 548, 343–346, https://doi.org/10.1038/nature23451 (2017).
Horlbeck, M. A. et al. Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. Elife 5, e19760, https://doi.org/10.7554/eLife.19760 (2016).
Simeonov, D. R. et al. Discovery of stimulation-responsive immune enhancers with CRISPR activation. Nature 549, 111–115, https://doi.org/10.1038/nature23875 (2017).
Chow, R. D. et al. AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma. Nat. Neurosci. 20, 1329–1341, https://doi.org/10.1038/nn.4620 (2017).
Ding, S. et al. Efficient transposition of the piggyBac (PB) transposon in mammalian cells and mice. Cell 122, 473–483, https://doi.org/10.1016/j.cell.2005.07.013 (2005).
McCarthy, K. D. & de Vellis, J. Preparation of separate astroglial and oligodendroglial cell cultures from rat cerebral tissue. J. Cell. Biol. 85, 890–902, https://doi.org/10.1083/Jcb.85.3.890 (1980).
Deverman, B. E. et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nat. Biotechnol. 34, 204–209, https://doi.org/10.1038/nbt.3440 (2016).
Zhou, H. et al. Cerebellar modules operate at different frequencies. Elife 3, e02536, https://doi.org/10.7554/eLife.02536 (2014).
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA.Gene Ther. 6, 1258–1266, https://doi.org/10.1038/sj.gt.3300947 (1999).
Huang, P. et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475, 386–389, https://doi.org/10.1038/nature10116 (2011).
Yang, S. et al. MANF regulates hypothalamic control of food intake and body weight. Nat. Commun. 8, 579, https://doi.org/10.1038/s41467-017-00750-x (2017).
Lundgaard, I. et al. Direct neuronal glucose uptake heralds activity-dependent increases in cerebral metabolism. Nat. Commun. 6, 6807, https://doi.org/10.1038/ncomms7807 (2015).
Greferath, U. et al. Inner retinal change in a novel rd1-FTL mouse model of retinal degeneration. Front. Cell. Neurosci. 9, 293, https://doi.org/10.3389/fncel.2015.00293 (2015).
Wang, P. et al. Dual role of Ski in pancreatic cancer cells: tumor-promoting versus metastasis-suppressive function. Carcinogenesis 30, 1497–1506, https://doi.org/10.1093/carcin/bgp154 (2009).
McCracken, J. M. et al. C57BL/6 substrains exhibit different responses to acute carbon tetrachloride exposure: implications for work involving transgenic mice. Gene Expr. 17, 187–205, https://doi.org/10.3727/105221617X695050 (2017).
Yao, X. et al. Homology-mediated end joining-based targeted integration using CRISPR/Cas9. Cell. Res. 27, 801–814, https://doi.org/10.1038/cr.2017.76 (2017).
Ivaniutsin, U., Chen, Y., Mason, J. O., Price, D. J. & Pratt, T. Adenomatous polyposis coli is required for early events in the normal growth and differentiation of the developing cerebral cortex. Neural Dev. 4, 3, https://doi.org/10.1186/1749-8104-4-3 (2009).
Ou, Z. et al. The combination of CRISPR/Cas9 and iPSC technologies in the gene therapy of human β-thalassemia in mice. Sci. Rep. 6, 32463, https://doi.org/10.1038/srep32463 (2016).
Huang, M. et al. Ptf1a, Lbx1 and Pax2 coordinate glycinergic and peptidergic transmitter phenotypes in dorsal spinal inhibitory neurons. Dev. Biol. 322, 394–405, https://doi.org/10.1016/j.ydbio.2008.06.031 (2008).
Schinko, J., Posnien, N., Kittelmann, S., Koniszewski, N. & Bucher, G. Single and double whole-mount in situ hybridization in red flour beetle (Tribolium) embryos. Cold Spring Harb. Protoc. 2009, t5258, https://doi.org/10.1101/pdb.prot5258 (2009).
Ramer, M. S. Anatomical and functional characterization of neuropil in the gracile fasciculus. J. Comp. Neurol. 510, 283–296, https://doi.org/10.1002/cne.21785 (2008).
Ting, J. T., Daigle, T. L., Chen, Q. & Feng, G. Acute brain slice methods for adult and aging animals: application of targeted patch clamp analysis and optogenetics. Methods Mol. Biol. 1183, 221–242, https://doi.org/10.1007/978-1-4939-1096-0_14 (2014).
Li, C. L. et al. Somatosensory neuron types identified by high-coverage single-cell RNA-sequencing and functional heterogeneity. Cell. Res. 26, 83–102, https://doi.org/10.1038/cr.2015.149 (2016).
Acknowledgements
We thank D. Li, E. Zuo, Y. Shi, Y. Liu, J. He, J. Pan, Y. Zhong, Y. Lu, Y. Zhang, J. Yang and X. Tang for technical assistance and valuable discussion. This work was supported by National Science and Technology Major Project (2017YFC1001302), CAS Strategic Priority Research Program (XDB02050007, XDA01010409), the MoST863 Program (2015AA020307), NSFC grants (31522037, 31500825, 31571509, 31522038), China Youth Thousand Talents Program (to H.Y.), Break through project of Chinese Academy of Sciences, Shanghai Sailing Plan for the Young Scientific Talents (15YF1414700), and The Ministry of Science and Technology of China (MOST; 2016YFA0100500).
Author information
Authors and Affiliations
Contributions
H.Z. designed and performed the experiments. J.L. designed and performed the in vivo gene activation in the liver and western blot. C.Z. designed the SPH activation system. N.G., H.L., X.H. and J.Z. cloned the vectors, and performed and analyzed the gene activation experiments in vitro, ex vivo and in vivo. X.S. and L.S. produced AAV8. Z.R., L.C. and F.L. designed and performed the in situ hybridization, ex vivo activation of Ascl1 and Neurog2 in astrocytes and assisted with the in vivo conversion experiments. C.L. and X.Z. performed single-cell RNA sequencing, Y.S. analyzed RNA sequencing and single-cell RNA-sequencing data. H.X. performed electrophysiology. Y.W., X.Y. and W.Y. generated the SPH transgenic mice. C.T. assisted with the immunofluorescence staining of brains and livers. P.H. designed and supervised the experiments of in vivo activation in SPH mice. H.Y. supervised the project and designed experiments. H.Z., P.H. and H.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Design of a fluorescence reporter system.
(a) mCherry reporter is driven by a minimal CMV promoter and activated by an upstream sgRNA target site. Note that miniCMV is a weak promoter, thus the basal fluorescence level is very low. See also18. (b) BFP was co-transfected to control the transfection efficiency (n = 2 cell cultures). Scale bar: 100 μm.
Supplementary Figure 2 Flow cytometry analysis of SPH-based activation.
(a) Schematic of the vectors used to evaluate SPH-based activation. Cre recombinase was used to remove the LSL cassette in the SPH vector. (b) Representative images show that co-transfection of sgRNA-miniCMV, SPH and Cre led to enhanced expression of mCherry (n = 4 cell cultures, 4 images). mCherry was activated in 93 ± 1% of the cells expressing SPH activator system. All values are presented as mean ± s.e.m.
Supplementary Figure 3 SPH shows more potent activation efficiency compared to SAM and VPR in primary astrocytes.
(a) Schematic illustration of all plasmids. The expressions of all functional elements were probed by the fluorescence reporter mCherry or GFP. (b) 2 days after transfection, cells were prepared for FACS. The mCherry and GFP double-positive cells were collected for qPCR. (c) SPH achieved the highest level of activation, which supports the observations in cell lines (n = 2 cell cultures).
Supplementary Figure 4 Characterization of SPH transgenic mice.
(a) Schematic of the integration sites of SPH in a SPH transgenic mouse genome. (b) Genotyping of SPH transgenic mice. “+” indicates positive mice. Percentage of positive progenies was calculated on 101 mice. Gel image was cropped. (c) Systemic evaluation of the Alb-Cre effectiveness in cleaving LSL cassettes specifically in the liver. Note that Alb is a liver-specific promoter (n = 1 mouse). Gel image was cropped. (d) Fluorescence images of the primary SPH fibroblasts infected by lentivirus containing sgRNA-EF1a-Cre-mCherry, confirming expression of dCas9 and activators were induced by Cre recombinase (n = 3 mice). (e) Expressions of dCas9 in Cre-expressing fibroblasts. Gel image was cropped. (f) Characterization of Cre effectiveness in the liver after hydrodynamic injection of the plasmids expressing Cre and mCherry (n = 1 mouse). (g) Stereotactic Injection of AAV-hSyn-mCherry-Cre to induce neuron-specific expression of EGFP (n = 1 mouse). Red arrowheads indicate primers. Scale Bars: d, f, g, 20 μm.
Supplementary Figure 5 Targeted activation of individual genes and lncRNA in the liver through tail vein injection of plasmids.
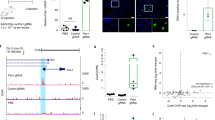
(a) Experimental design. Plasmids with Cre, sgRNA and mCherry expression were delivered into the liver by hydrodynamic injection via tail vein. Note that mRNA was extracted from the liver without isolating mCherry-positive cells and used for qPCR analysis (n = 1 mouse). Scale Bar: 100 μm. (b) Targeted activation of individual genes in the liver after injection of plasmids expressing indicated sgRNAs (number of mice denoted within graph). Increased amount of ASCL1 protein was confirmed by western blot (n = 1 mouse). Gel images were cropped. (c) Induction of lncRNAs in the liver (n = 3 mice). (d) Activation of multiple genes in the liver through tail vein injection of a mixture of sgRNA-expressing vectors (n = 2 mice). The sgLacZ serves as a control sgRNA.
Supplementary Figure 6 Remodeling of metabolic liver zonation by activating Dkk1 in vivo.
(a) Schematic of the plasmid used for sgDkk1 expression. (b,c) SPH mice were hyperdynamically injected with sgRNA-mCherry plasmids. Immunofluorescence staining were performed 4–6 days after plasmids injections on liver sections. mCherry (red) was stained to indicate cells receiving plasmids. Both GS and CYP2E1 (cyan) were β-catenin target genes locating inpericentral area of the liver lobule. Arrows indicate mCherry-positive cells showing downregulation of GS and CYP2E1. Note that expression of GS and CYP2E1 in pericentral hepatocytes receiving sgDkk1-Cre-mCherry was inhibited. Scale bar = 50 μm. (d) Increased Dkk1 transcripts were quantified by qPCR (n = 5 mice). (e, f) Percentage of cells showing suppressed expression of GS (88/439 VS 0/165, sgLacZ: n = 2 mice, sgDkk1: n = 2 mice) or CYP2E1 (127/829 VS 0/66, sgLacZ: n = 2 mice, sgDkk1: n = 5 mice) in plasmid-expressing hepatocytes. (g) Schematic illustration of DKK1-mediated suppression of Wnt/β-catenin signaling. Activation of Dkk1 leads to reduced expressions of two β-catenin target genes; green circles indicate mCherry-positive cells showing downregulation of GS and CYP2E1. The sgLacZ serves as a control sgRNA.
Supplementary Figure 7 Prescreen of the effective sgRNAs.
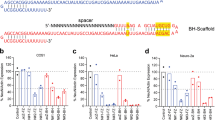
(a) Prescreening in primary fibroblasts derived from SPH mice (n = 2 cell cultures) and (b) N2a cells (n = 1 cell culture). The sgRNAs used in the sgRNA arrays are denoted as red. The sgLacZ serves as a control sgRNA.
Supplementary Figure 8 Simultaneous activation of 8 genes and 2 lncRNAs in mCherry-positive cells via hydrodynamic injection of all-in-one plasmids.
(a) Schematic of the all-in-one vector used for sgRNAs and Cre expression, and experimental flow of gene activations in the liver of SPH transgenic mice. Note that mRNA was extracted from mCherry-positive cells. (b) Simultaneous activation of 8 genes and 2 lncRNAs in the liver (n = 3 mice). The sgLacZ serves as a control sgRNA.
Supplementary Figure 9 Simultaneous activation of 10 genetic elements including 8 genes and 2 lncRNAs in the liver via hydrodynamic injection of dual-AAV vectors.
(a) Experimental flow of gene activations in the liver of SPH transgenic mice. (b) Schematic of the AAV-10-sgRNA-array and AAV-Alb-Cre. (c) Simultaneous activation of 8 genes and 2 lncRNAs (n = 2 mice). SPH mouse injected with vehicle serves as a control.
Supplementary Figure 10 Simultaneous activation of multiple genes in single-cell level.
Primary astrocytes derived from the SPH mice were transfected with an all-in-one vector expressing Cre, mCherry and an sgRNA array targeting the promoters of 8 genes. Due to detection limit, 3 (Ascl1, Slc6a4 and Slc7a11) out of 8 genes (Neurog2, Slc6a4, Dkk1, Hbb, Slc7a11, IL10, Ascl1, Neurod1) were detected in single-cell RNA-seq data. The average expression levels (FPKM values) of Ascl1, Slc6a4 and Slc7a11 were normalized to control group (n = 3 cells). 6 cells receiving the all-in-one vector were analyzed. Control cells were introduced with sgLacZ constructs.
Supplementary Figure 11 Enhance transcription of Hbb and Dkk1 was confirmed by in situ hybridization.
Enhance transcription of Hbb and Dkk1 was detected in the cerebral cortex after injecting AAV-hSyn-Cre and AAV-sgDkk1-sgHbb (n = 1 mouse). Scale Bar: 20 μm. The contralateral cerebral cortex serves as control.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhou, H., Liu, J., Zhou, C. et al. In vivo simultaneous transcriptional activation of multiple genes in the brain using CRISPR–dCas9-activator transgenic mice. Nat Neurosci 21, 440–446 (2018). https://doi.org/10.1038/s41593-017-0060-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-017-0060-6
This article is cited by
-
Proinflammatory polarization of engineered heat-inducible macrophages reprogram the tumor immune microenvironment during cancer immunotherapy
Nature Communications (2024)
-
Acoustically targeted noninvasive gene therapy in large brain volumes
Gene Therapy (2024)
-
CRISPR technologies for genome, epigenome and transcriptome editing
Nature Reviews Molecular Cell Biology (2024)
-
APOE4/4 is linked to damaging lipid droplets in Alzheimer’s disease microglia
Nature (2024)
-
Compact engineered human mechanosensitive transactivation modules enable potent and versatile synthetic transcriptional control
Nature Methods (2023)