Abstract
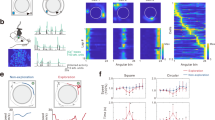
Natural environments are represented by local maps of grid cells and place cells that are stitched together. The manner by which transitions between map fragments are generated is unknown. We recorded grid cells while rats were trained in two rectangular compartments, A and B (each 1 m × 2 m), separated by a wall. Once distinct grid maps were established in each environment, we removed the partition and allowed the rat to explore the merged environment (2 m × 2 m). The grid patterns were largely retained along the distal walls of the box. Nearer the former partition line, individual grid fields changed location, resulting almost immediately in local spatial periodicity and continuity between the two original maps. Grid cells belonging to the same grid module retained phase relationships during the transformation. Thus, when environments are merged, grid fields reorganize rapidly to establish spatial periodicity in the area where the environments meet.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Moser, E. I. et al. Grid cells and cortical representation. Nat. Rev. Neurosci. 15, 466–481 (2014).
Rowland, D. C., Roudi, Y., Moser, M. B. & Moser, E. I. Ten years of grid cells. Annu. Rev. Neurosci. 39, 19–40 (2016).
Hafting, T., Fyhn, M., Molden, S., Moser, M. B. & Moser, E. I. Microstructure of a spatial map in the entorhinal cortex. Nature 436, 801–806 (2005).
McNaughton, B. L., Battaglia, F. P., Jensen, O., Moser, E. I. & Moser, M. B. Path integration and the neural basis of the ‘cognitive map’. Nat. Rev. Neurosci. 7, 663–678 (2006).
Stemmler, M., Mathis, A. & Herz, A. V. Connecting multiple spatial scales to decode the population activity of grid cells. Sci. Adv. 1, e1500816 (2015).
Sargolini, F. et al. Conjunctive representation of position, direction, and velocity in entorhinal cortex. Science 312, 758–762 (2006).
Kropff, E., Carmichael, J. E., Moser, M. B. & Moser, E. I. Speed cells in the medial entorhinal cortex. Nature 523, 419–424 (2015).
Solstad, T., Boccara, C. N., Kropff, E., Moser, M. B. & Moser, E. I. Representation of geometric borders in the entorhinal cortex. Science 322, 1865–1868 (2008).
Savelli, F., Yoganarasimha, D. & Knierim, J. J. Influence of boundary removal on the spatial representations of the medial entorhinal cortex. Hippocampus 18, 1270–1282 (2008).
Bonnevie, T. et al. Grid cells require excitatory drive from the hippocampus. Nat. Neurosci. 16, 309–317 (2013).
Zhang, S. J. et al. Optogenetic dissection of entorhinal-hippocampal functional connectivity. Science 340, 1232627 (2013).
Tolman, E. C. Cognitive maps in rats and men. Psychol. Rev. 55, 189–208 (1948).
Derdikman, D. et al. Fragmentation of grid cell maps in a multicompartment environment. Nat. Neurosci. 12, 1325–1332 (2009).
Kraus, B. J. et al. During running in place, grid cells integrate elapsed time and distance run. Neuron 88, 578–589 (2015).
Fuhs, M. C. & Touretzky, D. S. A spin glass model of path integration in rat medial entorhinal cortex. J. Neurosci. 26, 4266–4276 (2006).
Burak, Y. & Fiete, I. R. Accurate path integration in continuous attractor network models of grid cells. PLOS Comput. Biol. 5, e1000291 (2009).
Hardcastle, K., Ganguli, S. & Giocomo, L. M. Environmental boundaries as an error correction mechanism for grid cells. Neuron 86, 827–839 (2015).
Barry, C., Hayman, R., Burgess, N. & Jeffery, K. J. Experience-dependent rescaling of entorhinal grids. Nat. Neurosci. 10, 682–684 (2007).
Krupic, J., Bauza, M., Burton, S., Barry, C. & O’Keefe, J. Grid cell symmetry is shaped by environmental geometry. Nature 518, 232–235 (2015).
Stensola, H. et al. The entorhinal grid map is discretized. Nature 492, 72–78 (2012).
Stensola, T., Stensola, H., Moser, M. B. & Moser, E. I. Shearing-induced asymmetry in entorhinal grid cells. Nature 518, 207–212 (2015).
Fyhn, M., Hafting, T., Treves, A., Moser, M. B. & Moser, E. I. Hippocampal remapping and grid realignment in entorhinal cortex. Nature 446, 190–194 (2007).
Carpenter, F., Manson, D., Jeffery, K., Burgess, N. & Barry, C. Grid cells form a global representation of connected environments. Curr. Biol. 25, 1176–1182 (2015).
Skaggs, W. E. & McNaughton, B. L. Spatial firing properties of hippocampal CA1 populations in an environment containing two visually identical regions. J. Neurosci. 18, 8455–8466 (1998).
Paz-Villagrán, V., Save, E. & Poucet, B. Independent coding of connected environments by place cells. Eur. J. Neurosci. 20, 1379–1390 (2004).
Spiers, H. J., Hayman, R. M., Jovalekic, A., Marozzi, E. & Jeffery, K. J. Place field repetition and purely local remapping in a multicompartment environment. Cereb. Cortex 25, 10–25 (2015).
Couey, J. J. et al. Recurrent inhibitory circuitry as a mechanism for grid formation. Nat. Neurosci. 16, 318–324 (2013).
Yoon, K. et al. Specific evidence of low-dimensional continuous attractor dynamics in grid cells. Nat. Neurosci. 16, 1077–1084 (2013).
Kropff, E. & Treves, A. The emergence of grid cells: intelligent design or just adaptation? Hippocampus 18, 1256–1269 (2008).
Si, B. & Treves, A. A model for the differentiation between grid and conjunctive units in medial entorhinal cortex. Hippocampus 23, 1410–1424 (2013).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A. D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience 131, 1–11 (2005).
Langston, R. F. et al. Development of the spatial representation system in the rat. Science 328, 1576–1580 (2010).
Vincent, L. Morphological grayscale reconstruction in image analysis: applications and efficient algorithms. IEEE Trans. Image Process. 2, 176–201 (1993).
Acknowledgements
We thank S. Rosay and Y. Bitterman for discussion and M.P. Witter for advice on histology. We thank A.M. Amundsgård, K. Haugen, E. Kråkvik, B.B. Løfaldli, H. Waade and V. Frolov for technical assistance. The work was supported by the European Commission’s FP7 FET Proactive Programme on Neuro-Bio-Inspired Systems to E.I.M. (GRIDMAP, Grant Agreement 600725), an Advanced Investigator Grant from the European Research Council to E.I.M. (GRIDCODE, grant no. 338865), the Centre of Excellence scheme and the National Infrastructure Scheme of the Research Council of Norway (Centre for Neural Computation, grant number 223262 to M.-B.M. and E.I.M.; NORBRAIN1, grant number 197467 to E.I.M.), the Louis Jeantet Prize (M.-B.M. and E.I.M.), the Körber Prize (M.-B.M. and E.I.M.), and the Kavli Foundation (M.-B.M. and E.I.M.).
Author information
Authors and Affiliations
Contributions
T. Wernle, M.-B.M. and E.I.M. designed the experiment, T. Wernle and T. Waaga collected data, T. Wernle analyzed data, T. Waaga and M.M. contributed to analysis development, M.-B.M. and E.I.M. supervised the project, A.T. advised on analyses, and T. Wernle and E.I.M. wrote the paper with the input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Nissl-stained sagittal sections for 11 rats with grid cells in MEC layers II–V and 2 rats with head direction cells at the MEC-parasubiculum border (PaS).
Rat identity, brain hemisphere (L = left, R = right), and MEC or MEC-PaS layer are indicated at the top. Black arrowheads indicate the most dorsal and most ventral recording location along the tetrode track.
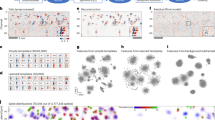
Supplementary Figure 2 Representative rate maps in the divided A|B and the merged AB environment and corresponding sliding correlation heat maps A|B x AB for all 10 rats.
(a) Left: Firing rate maps in A|B and AB. Cell number and rat identity are indicated at the top (t = tetrode, c = cell). Color coded from dark blue (0) to dark red (peak firing rate). Peak Firing rates are indicated at the top and bottom right. Scale bar, 50 cm. Right: Sliding correlation heat map for the cell to the left color-coded from dark blue to dark red (color bar). Scale bar, 50 cm. (b) Distribution of values for mean grid spacing in A, B, and AB across the entire cell population. A wide range of values, with no particular bias, is represented. (c) Ratemaps in A and B were in most cases represented by non-identical grid patterns, although the correlation between rate maps for A and B (shown in c) was higher than expected by chance (Pearson product-moment correlation A vs. B: 0.237 ± 0.023, one-sample two-sided Student’s t-test: t(127) = 10.4, P = 1.14 x 10−18; correlation A vs. 180-degree rotation of B: 0.002 ± 0.021; two-sample two-sided Student’s t-test for correlation A vs. B against correlation A vs. B rotated: t(254) = 7.6, P = 7.5 x 10−13). (d) Line plot showing change in mean correlation between A|B and AB in the central 10% bands and the distal 10% bands as a function of grid spacing. Grid spacing was sorted into bins of 10cm width (mean ± s.e.m., n = 128 cells, 10 rats). Note that, for cells with grid spacing < 90–100 cm, central correlations were lower than distal correlations across the entire range of values for grid spacing, suggesting that reorganization is stronger in the centre than the periphery across several modules. For cells with larger values for grid spacing, correlation values may be inaccurate since larger grids have fewer fields, which mostly touch the distal walls. This, and the frequent absence of fields in the center, may cause inaccuracy in estimates of central field translation.
Supplementary Figure 3 The grid pattern in the merged box AB was not a mere extension of the original map in A or B.
(a) The new map in AB is different from extended A and B maps. Top: Representative firing rate map in the divided A|B environment and the merged environment AB. Scale bar, 50 cm. Center: Simulated grid pattern A extended to the size of the merged map AB (200 x 200 cm) and the corresponding sliding correlation heat map between AB vs. extended pattern A, color-coded from dark blue to dark red (color bar). Scale bar, 50 cm. Bottom: same as center column but for the extended grid pattern of map B. (b) Left: Average sliding correlation heat map extended pattern A x AB for all cells and rats color coded as in a (n = 128 cells, 10 rats). Note that the correlation of the distal 10th percentile band of the AB map on the A side was significantly higher with extended map A (0.53 ± 0.02) than with extended map B (0.04 ± 0.02, mean ± s.e.m., n = 128 cells, 10 rats, t(254) = 17.4, P = 2.7 x 10−45, two-sided Student’s t-test). Right: same as to the left but for extended map B. The distal 10th percentile band of the AB map had a significantly higher correlation with extended map B (0.59 ± 0.01) than with extended map A (0.02 ± 0.02, mean ± s.e.m., n = 128 cells, 10 rats, t(254) = −16.7, P = 5.9 x 10−43, two-sided Student’s t-test). (c) When the central wall was reintroduced after successive trials in the merged environment, the original grid maps A and B were fully re-expressed. Left: correlation between map A on the last trial before AB and map A on the first trial after AB: 0.75 ± 0.02, mean ± s.e.m., n = 28 cells, 8 rats, t(27) = 33.7, P = 1.30 x 10−23 two-sided Student’s t-test; Right: same as to the left but for map B: 0.82 ± 0.01, mean ± s.e.m., n = 20 cells, 7 rats, t(19)=60.76, P = 13.13 x 10−23, two-sided Student’s t-test; up to 4 consecutive AB trials intervened between the two trials in the divided environment).
Supplementary Figure 4 Translocation of grid fields in the central transition zone of map AB.
(a) Left: Vector map for all cells in one rat. Vectors indicate field translocation and point from the original field position in A or B to the nearest field in AB within 60% of the cell’s grid spacing in A, B, and AB. Number of cells is indicated at the bottom right. Center: Distribution plots show translation vectors (in black) for blocks of 50 x 50 cm of the recording box. Mean resultant vectors (MVL) are added on top of the single vectors as thick red lines. Grey background shading indicates MVL P<0.001 with Rayleigh test for uniformity. Right: Average sliding cross-correlation heat map for the cells to the left, color coded from dark blue to dark red (color bar). Shifts were normalized to each cell’s average grid spacing in A, B, and AB (1 bin = 2 cm). (b)- (f) Further examples from other rats, displayed as in a. (g) As in a-f but the vector map is for all vectors from all cells and rats (n = 128, 10 rats). To compute average local field displacement, vectors were sorted into square windows of side length 16 cm and averaged (see Fig. 2c). (h) The displacement of firing fields in the centre of the box was accompanied by a change in peak firing rates of grid fields that could be matched before and after removal of the wall. The scatterplot shows that, after the wall was removed, the peak firing rate of grid fields in the central 10th percentile bands increased from 3.91 ± 0.15 Hz in A|B to 4.70 ± 0.18Hz in AB (t(548) = −3.32, P = 9.5 x 10−4, two-sided Student’s t-test; peak rate defined as the rate in the central bin of the grid field). In the distal 10th percentile bands, there was no significant change in firing rates (before: 3.61 ± 0.12 Hz; after: 3.44 ± 0.12 Hz; t(692) = 0.92, P = 0.35, two-sided Student’s t-test).
Supplementary Figure 5 Establishment of a locally continuous grid pattern in the merged environment.
(a) Standard deviation of grid field distances for all grid fields from all rats. Left: Before wall removal. Each dot represents one grid field and is color coded according to the standard deviation of distances to all neighboring fields within 130% of the cell’s average grid spacing in A, B, and AB (blue = minimum standard deviation in A|B and AB; red = maximum standard deviation in A|B and AB; n =128 cells, 1576 fields, 10 rats). Right: After wall removal (n =128 cells, 1447 fields, 10 rats). To compute average local standard deviations, standard deviations were sorted into square windows of side length 16 cm and averaged (see Fig. 3b, c). Scale bar, 50cm.(b) Local grid field offset as a function of distance from partition wall. Average field offset between A|B or AB and a template grid pattern determined from A (A|B vs. template in black; AB vs. template in red; offsets are binned into 10th percentile bands of 200 x 20 cm each, mean ± s.e.m., n = 128 cells). Grey stippled line indicates location of the former partition wall. (c) As in a but for a template pattern determined from B. Note, before wall removal phase offsets change abruptly in the center percentiles and approximate a step function. After wall removal, phase offsets change more linearly between A and B, indicating translocation of single grid fields into a pattern that is locally continuous throughout the environment. To quantify this transition, a linear regression was fit to the data from 10% bands spanning from the distal north wall in A to the distal south wall in B, as well as a step function with the step at the position of the partition wall. The residual mean square root (r.m.s.) for differences between data and step function increased when the wall was removed (r.m.s. expressed as percentage grid spacing for A|B vs. AB: 2.0% vs. 4.0%, square residuals A|B vs. AB: t(38) = 2.1, P = 0.04, two-sided Student’s t-test). The residual mean square root for differences between data and linear regression decreased (r.m.s. A|B vs. AB: 4.3% vs. 2.2%, square residuals A|B vs. AB: t(38) = 3.9, P = 3.3 x 10−4, two-sided Student’s t-test), suggesting that translocation of firing locations decreased gradually with distance from the partition wall. This trend was generally upheld when we analyzed data from subsets of grid cells with either small or large grid scales (small: 44 cm to 61 cm, n = 38 cells, r.m.s. step function A|B vs AB: 2.4 % vs. 3.6 %, r.m.s. linear regression A|B vs AB: 3.6% vs. 2.0%; large-scale grid cells range: 84 cm to 136 cm, n = 38 cells, r.m.s. step function A|B vs AB: 2.9% vs. 4.9%, r.m.s. linear regression A|B vs AB: 3.5% vs. 2.1%). The progression towards a linear change in offsets from the template indicates that local grid coherence increased throughout the environment.
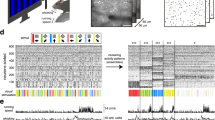
Supplementary Figure 6 Development of firing rate distribution during the first trial in the merged environment AB for all cells and rats.
Left column of each panel: correlation of interval rate maps for cumulative 1-min blocks in AB vs. the corresponding rate map for the entire trial in either A|B (light red line) or AB (dark red line). Only the central half of the environment is included in the correlation analysis (center half of A, center half of B; together they covered 200 cm x 100 cm of the box). Time intervals increased cumulatively by 60s (block 1 is for 0–1 min, block 2 for 0–2 min, block 3 for 0–3 min, etc). Rat number and cell identity are indicated at the top (T = tetrode, C = cell). Right column: same as to the left but for the distal half of A|B (light grey line) or AB (dark grey line). Also here the distal half of A and distal half of B covered together 200 cm x 100 cm.
Supplementary Figure 7 Grid cell patterns are stable across consecutive trials in AB.
(a) Example rate maps for one grid cell recorded across 5 consecutive trials in AB after the wall was removed for the third time (trial numbers are indicated on the top, trial numbers AB1 to AB5 correspond to the 8th to the 12th exposure in the merged box). Peak firing rates are indicated at the bottom right. Rat identity and cell number is indicated at the top (T = tetrode, C = cell). Scale bar, 50cm. (b) Average correlation between pairs of 5 consecutive trials in AB for 9 simultaneously recorded grid cells including the cell in a, color-coded from dark blue to dark red (color bar). Grid cells shared the same spacing and orientation. Note, all pairwise correlation values between trials were high, suggesting that in each cell, grid field locations showed little change between consecutive trials in AB.
Supplementary Figure 8 Phase relationships between pairs of grid cells were maintained after removal of the central partition wall.
(a) Left: Average difference in distance offset between same (S) and randomly (R) chosen cell pairs. Data are plotted as in Fig. 5b,c. Red line indicates median difference in offset of the displaced center peak, box edges indicate 25th and 75th percentiles, whiskers extend to the last data point that lies within 1.5 times the interquartile range, and data points larger than 1.5 times the interquartile range are considered outliers (black dots). Rat number and trial date are indicated at the top. Number of cells and cell pairs are indicated at the bottom right (***P < 0.001, two-sided Wilcoxon rank sum test). Center: Average angular difference of orientation offset between same and random cell pairs (***P < 0.001, Watson U2 test). Right: Pairs of grid cells share the same orientation and spacing. Average correlation of autocorrelograms for all cell pairs. Stippled red line indicates 90th percentile of all pairwise correlations from all cells and rats (n = 128 cells / 8128 pairs). (b) – (f) As in a but for different trials or animals (**P < 0.05, Watson U2 test).
Supplementary Figure 9 Representative border cells, nonperiodic spatially modulated cells, and head direction cells.
(a) Representative firing rate maps for 6 border cells from 3 rats for the divided A|B environment and the merged AB environment. Peak firing rates are indicated at the bottom right. Rat identity and cell number is indicated at the top (T = tetrode, C = cell). Scale bar, 50 cm. (b) As in a but for 7 non-periodic spatially modulated cells from 3 rats in A|B and AB. Scale bar, 50 cm. These cells had high spatial information content and stable firing locations (spatial information value of 0.82 ± 0.05 and spatial correlation between first and second half of trial of 0.49 ± 0.03; all trials and all parts of environment, mean ± s.e.m). (c) Polar plots of angular firing rate distribution for 10 representative head direction cells from 3 rats in A, B, and AB. Rat number and trial date are indicated at the top. Cell number is indicated at the left site (T = tetrode, C = cell). Mean resultant vector length, a measure for directional tuning, is indicated at the top in each plot. Peak firing rate is indicated at the bottom right.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9
Supplementary Table 1
Number of trials and number of functional cell types recorded from each individual animal in the study.
Rights and permissions
About this article
Cite this article
Wernle, T., Waaga, T., Mørreaunet, M. et al. Integration of grid maps in merged environments. Nat Neurosci 21, 92–101 (2018). https://doi.org/10.1038/s41593-017-0036-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-017-0036-6
This article is cited by
-
A consistent map in the medial entorhinal cortex supports spatial memory
Nature Communications (2024)
-
Modeling the grid cell activity based on cognitive space transformation
Cognitive Neurodynamics (2023)
-
The grid code for ordered experience
Nature Reviews Neuroscience (2021)
-
Effect of reward on electrophysiological signatures of grid cell population activity in human spatial navigation
Scientific Reports (2021)
-
Place cells and geometry lead to a flexible grid pattern
Journal of Computational Neuroscience (2021)