Abstract
Spatial omics data are advancing the study of tissue organization and cellular communication at an unprecedented scale. Flexible tools are required to store, integrate and visualize the large diversity of spatial omics data. Here, we present Squidpy, a Python framework that brings together tools from omics and image analysis to enable scalable description of spatial molecular data, such as transcriptome or multivariate proteins. Squidpy provides efficient infrastructure and numerous analysis methods that allow to efficiently store, manipulate and interactively visualize spatial omics data. Squidpy is extensible and can be interfaced with a variety of already existing libraries for the scalable analysis of spatial omics data.
Similar content being viewed by others
Main
Dissociation-based single-cell technologies have enabled the deep characterization of cellular states and the creation of cell atlases of many organs and species1. However, how cellular diversity constitutes tissue organization and function is still an open question. Spatially resolved molecular technologies aim at bridging this gap by enabling the investigation of tissues in situ at cellular and subcellular resolution2,3,4. In contrast to the current state-of-the-art dissociation-based protocols, spatial molecular technologies acquire data in greatly diverse forms, in terms of resolution (few cells per observation to subcellular resolution), multiplexing (dozens of features to genome-wide expression profiles), modality (transcriptomics, proteomics and metabolomics) and often with an associated high-content image of the captured tissue2,3,4. Such diversity in generated data and corresponding formats currently represents an infrastructural hurdle that has hampered urgently needed development of interoperable analysis methods. The underlying computational challenges lie in efficient data representation as well as comprehensive analysis and visualization methods.
Existing analysis frameworks for spatial data focus either on preprocessing5,6,7,8 or on one particular aspect of spatial data analysis9,10,11,12,13. The combination of different analysis steps is still hampered by the lack of a unified data representation and of a modular application programming interface, for example loading processed data from Starfish5, combining stLearn’s11 integrative analysis of tissue images together with Giotto’s powerful spatial statistics13, BayesSpace spatial clustering14 or leveraging state-of-the-art deep-learning-based methods for image segmentation15,16 and visualization17. A comprehensive framework that enables community-driven scalable analyses of both spatial neighborhood graph and image, along with an interactive visualization module, is missing (Supplementary Table 1).
For this purpose we developed ‘Spatial Quantification of Molecular Data in Python’ (Squidpy), a Python-based framework for the analysis of spatially resolved omics data (Fig. 1). Squidpy aims to bring the diversity of spatial data in a common data representation and provide a common set of analysis and interactive visualization tools. Squidpy introduces two main data representations to manage and store spatial omics data in a technology-agnostic way: a neighborhood graph from spatial coordinates and large-source tissue images acquired in spatial omics data (Fig. 1b). Both data representations leverage sparse18 or memory-efficient19 approaches in Python for scalability and ease of use. They are also able to deal with both two-dimensional and three-dimensional (3D) information, thus laying the foundations for comprehensive molecular maps of tissues and organs. Such infrastructure is coupled with a wealth of tools that enable the identification of spatial patterns in tissue and the mining and integration of morphology data from large tissue images (Fig. 1c). Squidpy is built on top of Scanpy and Anndata20 and it relies on several scientific computing libraries in Python, such as Scikit-image21, Napari22 and Dask19. Its modularity makes it suitable to be interfaced with a variety of additional tools in the Python data science and machine-learning ecosystem (such as external segmentation methods and modern deep-learning frameworks), as well as several single-cell data analysis packages. It provides a rich documentation, with tutorials and example workflows, integrated in the continuous integration pipeline. It allows users to quickly explore spatial datasets and lays the foundations for both spatial omics data analysis as well as development of new methods. Squidpy is available at https://github.com/theislab/squidpy; documentation and extensive tutorials covering the presented results and more are available at https://squidpy.readthedocs.io/en/latest/.
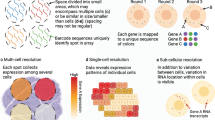
a, Squidpy supports inputs from diverse spatial molecular technologies with spot-based, single-cell or subcellular spatial resolution. b, Building upon the single-cell analysis software Scanpy20 and the Anndata format, Squidpy provides efficient data representations of these inputs, storing spatial distances between observations in a spatial graph and providing an efficient image representation for high-resolution tissue images that might be obtained together with the molecular data. c, Using these representations, several analysis functions are defined to quantitatively describe tissue organization at the cellular (spatial neighborhood) and gene level (spatial statistics, spatially variable genes and ligand–receptor interactions), to combine microscopy image information (image features and nuclei segmentation) with omics information and to interactively visualize high-resolution images.
Results
Squidpy provides infrastructure and analysis tools to identify spatial patterns in tissue
Spatial proximity is encoded in spatial graphs, which require flexibility to support the variety of neighborhood metrics that spatial data types and users may require. For instance, in Spatial Transcriptomics (ST23, Visium24 and DBit-seq25), a node is a spot and a neighborhood set can be defined by a fixed number of adjacent spots (square or hexagonal grid; Fig. 2a), whereas in imaging-based molecular data (seqFISH26, MERFISH27, Imaging Mass Cytometry28,29, CyCif30, 4i31 and Spatial Metabolomics32; Fig. 2a), a node can be defined as a cell (or pixel) and a neighborhood set can also be chosen based on a fixed radius (expressed in spatial units) from the centroid of each observation. Alternatively, other approaches, such as Euclidean distance or Delaunay triangulation, can be utilized to build the neighbor graph (Fig. 2a). Squidpy can compute all the aforementioned modalities thus making it technology-agnostic and providing the infrastructure for downstream analysis tools that aim at quantifying spatial organization of the tissue.
a, Example of nearest-neighbor graphs that can be built with Squidpy: grid-like and generic coordinates. b, Neighborhood enrichment analysis between cell clusters in spatial coordinates. Positive enrichment is found for the following cluster pairs: ‘Lateral plate mesoderm’ with ‘Allantois’ and ‘Intermediate mesoderm’ clusters, ‘Endothelium’ with ‘Hematoendothelial progenitors’, ‘Anterior somitic tissues’, ‘Sclerotome’ and ‘Cranial mesoderm’ clusters, ‘NMP’ with ‘Spinal cord’, ‘Allantois’ with ‘Mixed mesenchymal mesoderm’, ‘Erythroid’ with ‘Low quality’, ‘Presomitic mesoderm’ with ‘Dermomyotome’ and ‘Cardiomyocytes’ with ‘Mixed mesenchymal mesoderm’. These results were also reported by the original authors33. NMP, neuromesodermal progenitor. c, Visualization of selected clusters of the seqFISH mouse gastrulation dataset. d, Visualization in 3D coordinates of three selected clusters in the MERFISH dataset34. The ‘Pericytes’ are in pink, the ‘Endothelial 2’ are in red and the ‘Ependymal’ are in brown. The full dataset is visualized in Supplementary Fig. 2g. e, Results of the neighborhood enrichment analysis. The ‘Pericytes’ and ‘Endothelial 2’ clusters show a positive enrichment score. OD, oligodendrocytes. f, Visualization of subcellular molecular profiles in HeLa cells, plotted in spatial coordinates (approximately 270,000 observations/pixels). ER, endoplasmic reticulum. g, Cluster co-occurrence score computed for each cell, at increasing distance threshold across the tissue. The cluster ‘Nucleolus’ is found to be co-enriched at short distances with the ‘Nucleus’ and the ‘Nuclear envelope’ clusters. h, Visualization of SlideseqV2 dataset with cell-type annotations35. i, Cluster co-occurrence score computed for all clusters, conditioned on the presence of the ‘Ependymal’ cluster. At short distances, there is an increased colocalization between the ‘Endothelial_Tip’ cluster and the ‘Ependymal’ cluster. j, Ripley’s L statistics computed at increasing distances; clusters such as ‘CA1_CA2_CA3_Subiculum’ and ‘DentatePyramids’ show high Ripley’s L values across distances, providing quantitative evaluation of the ‘clustered’ spatial pattern across the slide. Clusters such as the ‘Endothelial_Stalk’, with a lower Ripley’s L value across increasing distances, have a more ‘random’ pattern. k, Expression of top three spatially variable genes (Ttr, Mbp and Hpca) as computed by Moran’s I spatial autocorrelation on the SlideseqV2 dataset. They seem to capture different patterning and specificity for cell types (‘Endothelial_Tip’, ‘Oligodendrocytes’ and ‘CA1_CA2_CA3_Subiculum’, respectively).
A key question in the analysis of spatial molecular data is the description and quantification of spatial patterns and cellular neighborhoods across the tissue. Squidpy provides several tools that leverage the spatial graph to address such questions. On a recently published seqFISH33 dataset we built a spatial nearest-neighbor graph based on Delaunay triangulation and computed a permutation-based neighborhood enrichment across cell-type annotations (Methods). Clusters of enriched cell types (such as ‘Lateral plate mesoderm’ with ‘Allantois’ and ‘Intermediate mesoderm’ clusters, ‘Endothelium’ with ‘Hematoendothelial progenitors’; Fig. 2b) are consistent with the original publication33 and the spatial proximity can be visualized in Fig. 2c. A similar analysis was performed on a MERFISH dataset34, where we could identify a neighborhood enrichment between ‘Endothelial 2’ and ‘Pericytes’ clusters, whereas the ‘Ependymal’ cluster shows a strong co-enrichment with itself but depleted enrichment with the other clusters (Fig. 2d,e shows selected clusters and Supplementary Fig. 2g shows the the full dataset). Furthermore, our implementation is scalable and ~tenfold faster than a similar implementation in Giotto13 (Supplementary Fig. 1a,b and Supplementary Table 2 show extensive comparisons), enabling analysis of large-scale spatial omics datasets. Sparse and scalable implementation in Squidpy enables working with subcellular-resolution spatial data such as 4i31. We considered ~270,000 pixels as subcellular resolution observations across 13 cells (Fig. 2f) and evaluated their cluster co-occurrence at increasing distances (Fig. 2g). As expected, the subcellular measurements annotated in the nucleus compartment co-occur together with the nucleus and the nuclear envelope, at short distances. The co-occurrence score represents an interpretable score to investigate patterns of spatial organization in tissue. When applied to a SlideseqV2 dataset35 (Fig. 2h), the co-occurrence score could provide a quantitative indication of a qualitative observation that the ‘Endothelial_Tip’ cluster shows a strong co-occurrence with the ‘Ependymal’ cluster (Fig. 2i). To obtain a global indication of the degree of clustering or dispersion of a cell-type annotation in the tissue area, the Ripley’s L can be computed. When applied to the same dataset (Fig. 2j), it highlighted how the ‘CA1 CA2 CA3 Subiculum’ and the ‘Dentate Pyramids’ annotations have a more ‘clustered’ spatial patterning than other annotations, such as the ‘Endothelial Stalk’. Squidpy implements three variations of the Ripley statistic (L, F and G; Supplementary Fig. 2b provides an additional example) that allows one to gain a global understanding of spatial patterning of discrete covariates. Finally, to identify genes that show strong spatial variability, we applied the Moran’s I spatial autocorrelation statistics (Methods) and visualized the three top genes (Fig. 2k; Ttr, Mbp and Hpca), which all show different spatial patterns and seem to largely colocalize with cell-type annotations (‘Endothelial Tip’, ‘Oligodendrocytes’ and ‘CA1 CA2 CA3 Subiculum’, respectively).
These statistics yield interpretable results across diverse experimental techniques, as demonstrated on an Imaging Mass Cytometry dataset36, where we showcase additional methods such as Ripley’s F function, average clustering and degree and closeness centrality (Supplementary Fig. 2). In conclusion, Squidpy provides a suite of orthogonal analysis tools that enable analysts to gain a quantitative understanding of the spatial organization of cellular and subcellular units.
Squidpy enables analysis and visualization of large images in spatial omics data
The high-resolution microscopy image additionally captured by spatial omics technologies represents a rich source of morphological information that can provide key biological insights into tissue structure and cellular variation. Squidpy introduces a new data object, the ImageContainer, which efficiently stores the image with an on-disk/in-memory switch based on xArray and Dask19,37. This object provides a general mapping between pixel coordinates and molecular profiles, enabling analysts to relate image-level observations to omics measurements (Fig. 3a). It provides seamless integration with napari22, thus enabling interactive visualization of analysis results stored in an Anndata object alongside the high-resolution image directly from a Jupyter notebook. It also enables interactive manual cropping of tissue areas and automatic annotation of observations in Anndata. As napari is an image viewer in Python, all the above-mentioned functionalities can be also interactively executed without additional requirements. Following standard image-based profiling techniques38, Squidpy implements a pipeline based on Dask Image19 and Scikit-image21 for preprocessing and segmenting images, extracting morphological, texture and deep-learning-powered features (Fig. 3a). To enable efficient processing of very large images, this pipeline utilizes lazy loading, image tiling and multiprocessing (Supplementary Fig. 1b). When using image tiling during processing, overlapping crops are used to mitigate border effects. Features can be extracted from a raw-tissue image crop or Squidpy’s segmentation module can be used to extract segmentation objects (nuclei or cells) counts, sizes or general image features at segmentation-mask level (Supplementary Fig. 2b).
a, Schematic drawing of the ImageContainer object and its relation to Anndata. The ImageContainer object stores multiple image layers with spatial dimensions x, y, z (left). An exemplary image-processing workflow consisting of preprocessing, segmentation and feature extraction is shown in the bottom. Using image features, pixel-level information is related to the molecular profile in Anndata (top right). Anndata and ImageContainer objects can be visualized interactively using napari (bottom right). DL, deep-learning. b, Fluorescence image with markers DAPI, anti-NeuN and anti-GFAP from a Visium mouse brain dataset (https://support.10xgenomics.com/spatial-gene-expression/datasets). The location of the inset in c is marked with a yellow box. c, Details of fluorescence image from b, showing from left to right the DAPI, anti-NeuN and anti-GFAP channels and nuclei segmentation of the DAPI stain using watershed segmentation. d, Image features per Visium spot computed from fluorescence image in b. From left to right are shown: number of nuclei in each Visium spot derived from the nuclei segmentation, the mean intensity of the anti-NeuN marker in each Visium spot and the mean intensity of the anti-GFAP marker in each Visium spot. e, Violin plot of log-normalized Gfap and Rbfox3 gene expression in Visium spots with low and high anti-GFAP and anti-NeuN marker intensity (lower and higher than median marker intensity), respectively. f, Calculation of per-cell features from a MIBI-TOF dataset41. Tissue image showing three markers CD45, CK and vimentin (left). Cell segmentation provided by the authors41 (center left). Mean intensity of CD45 per cell derived from the raw image using Squidpy (center right). Mean intensity of CK per cell derived from the raw image using Squidpy (right). For quantitative comparison see Supplementary Fig. 2. This example is part of the Squidpy documentation (https://squidpy.readthedocs.io/en/latest/auto_tutorials/tutorial_visium_fluo.html and https://squidpy.readthedocs.io/en/latest/auto_tutorials/tutorial_mibitof.html).
For segmentation, Squidpy provides a pipeline based on the watershed algorithm and provides an interface to state-of-the-art nuclei segmentation algorithms such as Cellpose16 and StarDist15 (Supplementary Fig. 5a). As an example for segmentation-based features, we computed nuclei segmentation using the 4,6-diamidino-2-phenylindole (DAPI) stain of a fluorescence mouse brain section (Fig. 3b,c) and showed the estimated number of nuclei per spot on the hippocampus (Fig. 3d, left). The cell-dense pyramidal layer can be easily distinguished with this view of the data, showcasing the richness and interpretability of information that can be extracted from tissue images when brought in a spot-based format. In addition, we can leverage segmented nuclei to inform cell-type deconvolution (or decomposition/mapping) methods such as Tangram39 or Cell2Location40. In Supplementary Fig. 4 we showcase how priors on nuclei densities derived from nuclei segmentation in Squidpy can be used both for inferring cell-type proportions as well as mapping cell types to segmentation objects with Tangram.
Image-based features contained in Squidpy include built-in summary, histogram and texture features and more-advanced features such as deep-learning-based (Supplementary Fig. 2a) or CellProfiler (Supplementary Fig. 5b) pipelines provided by external packages.
Using the anti-NeuN and anti-glial fibrillary acidic protein (GFAP) channels contained in the fluorescence mouse brain section, we calculated their mean intensity for each Visium spot using summary features (Fig. 3d center and right). This image-derived information relates well to molecular information: Visium spots with high marker intensity have a higher expression of Rbfox3 (for anti-NeuN marker) and Gfap (for anti-GFAP marker) than low-marker-intensity spots (Fig. 3e). Image features can also be calculated at the spot level, thus aggregating several cells or at an individual per-cell level. Using a multiplexed ion beam imaging by time of flight (MIBI-TOF) dataset41 with a previously calculated cell segmentation, we calculate mean intensity features of two markers contained in the original image (Fig. 3f). The calculated mean intensities have a high correlation with the associated mean intensity values contained in the associated molecular profile (Supplementary Fig. 3a). These results highlight how explicitly analyzing image-level information leads to insightful validation but also potentially new hypotheses.
Squidpy’s workflow enables the integrative analysis of spatial transcriptomics data
The feature extraction pipeline of Squidpy allows the comparison and joint analysis of spatial patterning of the tissue at the molecular and morphological level. Here, we show how Squidpy’s functionalities can be combined to analyze 10X Genomics Visium spatial transcriptomics data of a coronal mouse brain section.
As previously shown, we can apply spatially variable feature selection to identify genes that show a pronounced spatial pattern. Moran’s I spatial correlation statistics identifies Mobp and Nrgn (Fig. 4a,b) to be spatially variable; both genes show a distinct spatial expression pattern and seem to encompass the localization of several cell clusters (Fig. 4d; ‘Fiber tract’ and ‘Hypothalamus 2’ for Mobp and ‘Pyramidal layers’ and ‘Pyramidal layers/Dentate gyrus’ for Nrgn). An orthogonal method for the same task, Sepal42 ranks Krt18 as a top spatially variable gene, which shows a distinct expression in a subset of the ‘Lateral ventricle’ cluster (Fig. 4d and Supplementary Fig. 1f,g show a comparison with original implementation). The variety of tools for spatially variable gene identification provided by Squidpy enhances standard cluster-based gene expression signatures by providing insights into spatial distribution of genes. Ligand–receptor interaction analysis can be a useful approach to shortlist candidate genes driving cellular interactions. Squidpy provides a fast re-implementation of the CellphoneDB43 method (Supplementary Fig. 1b,d shows runtime comparison against original implementation and Giotto), which additionally leverages the Omnipath database for ligand–receptor annotations44 (Supplementary Fig. 1e shows a comparison with CellphoneDB). Applied to the same dataset, it highlighted different ligand–receptor pairs between the ‘Hippocampus’ cluster and the two ‘Pyramidal layer’ clusters. Whether permutation-based tests of ligand–receptor interaction identification are able to pinpoint cellular communication and pathway activity is an open question45. However, it is useful to inform such results with a quantitative understanding of cluster co-occurrence. Squidpy’s co-occurrence score is a simple but interpretable approach, which applied to the Visium dataset highlights an expected direct relationship between the previously described clusters (‘Hippocampus’ and the two ‘Pyramidal layer’ clusters; Fig. 4f).
a,b, Gene expression in spatial context of two spatially variable genes (Mobp and Nrgn) as identified by Moran’s I spatial autocorrelation statistic. c, Gene expression in spatial context of one spatially variable gene (Krt18) identified by the Sepal method42. d, Clustering of gene expression data plotted on spatial coordinates. e, Ligand–receptor interactions from the cluster ‘Hippocampus’ to clusters ‘Pyramidal layer’ and ‘Pyramidal layer dentate gyrus’. Shown are a subset of significant ligand–receptor pairs queried using the Omnipath database. Shown ligand–receptor pairs were filtered for visualization purposes, based on expression (mean expression > 13) and significant after false discovery rate (FDR) correction (P < 0.01). P values results from a permutation-based test with 1,000 permutations and were adjusted with the Benjamini–Hochberg method. f, Co-occurrence score between ‘Hippocampus’ and the rest of the clusters. As seen qualitatively by clusters in a spatial context in d, ‘Pyramidal layer’ and ‘Pyramidal layer dentate gyrus’ co-occur with the Hippocampus at short distances, given their proximity. g, H&E stain. h, Clustering of summary image features (channel intensity mean, s.d. and 0.1, 0.5, 0.9th quantiles) derived from the H&E stain at each spot location (for quantitative comparison to gene clusters from d see Supplementary Fig. 2e). i, Fraction of nuclei per Visium spot, computed using the cell segmentation algorithm StarDist15. j, Violin plot of fraction of nuclei per Visium spot (g) for the cortical clusters (d) plotted with P value annotation. The cluster Cortex_2 was omitted from this analysis because it entails a different region of the cortex (cortical subplate) for which the differential nuclei density score between isocortical layers is not relevant. Test performed was two-sided Mann–Whitney–Wilcoxon test with Bonferroni correction, P value annotation legend is the following: ****P ≤ 0.0001. Exact P values are the following: Cortex_5 versus Cortex_4, P = 1.691 × 10−36, U = 1,432; Cortex_5 versus Cortex_1, P = 2.060 × 10−54, U = 775; Cortex_5 versus Cortex_3, P = 5.274 × 10−51, U = 787. This example is part of the Squidpy documentation (https://squidpy.readthedocs.io/en/latest/auto_tutorials/tutorial_visium_hne.html).
Squidpy’s feature extraction pipeline enables direct comparison and joint analysis of image and omics data. The integrative analysis of gene expression and image data enhances pattern discovery and enables joint interpretation of the information obtained from morphology and molecular data. For instance, on the same mouse brain coronal section data, we compared clusters computed from gene expression profiles with clusters computed from summary statistics (mean, s.d., 0.1, 0.5 and 0.9th quantiles) of high-resolution hematoxylin and eosin (H&E) image channels (Fig. 4g,h). The image-based clusters recapitulate regions of image intensities with similar mean and standard variation, whereas the gene-based clusters are related to broad cell-type definition. We can see that several image-based clusters are highly overlapping with the gene-based clusters, especially in the cluster ‘Hippocampus’ (54% overlap with image feature cluster 10) and the cluster ‘Hypothalamus’ (72% overlap with image feature cluster 8). This shows how members of such clusters share a similar definition both at morphology and molecular level which allows further characterization of the cluster. In contrast, the image-based clusters provide a different view of the data in the cortex (no overlap >33% with any image feature clusters) (Supplementary Fig. 3e). Here, gene clusters identify broad cortical layers whereas the image-based clusters separate different regions of the cortex based on changing local image intensities, indicating changes in cell density, morphology or changes in the staining that are not captured by the gene expression data. For further examination of these image feature clusters, we calculated a nuclei segmentation using StarDist15 and extracted the number of nuclei per Visium spot (Fig. 4i). This nuclear count shows that image-based cluster 15 highlights an area in the bottom part of the cortex with low cell density that is not covered fully by the gene cluster ‘Cortex_5’. This example highlights how variation in interpretable image-based features can reveal higher variability within the same annotation and why the integrative tools available in Squidpy enables such analysis. In addition to explaining variation in the image-based clusters, the fraction of nuclei was combined with gene clusters to show that the nuclear density varies between the different cortical clusters (Fig. 4j,). This indicates that gene expression clusters represent a different grouping of the cortex than the one identified by the image-based clustering. Such regions of different nuclear densities and morphology in the brain are of broad interest to neuroscientists46,47,48 and low nuclei density in the outer cortical layer of the isocortex (corresponding to cluster ‘Cortex_5’) has been previously established48. Furthermore, Squidpy image-processing tools allow to quickly validate the robustness of such findings, by refining the selection of spots that fully overlap the detected tissue area to remove potential false positives (Supplementary Fig. 6). Therefore, nuclear density and morphological information represent valuable information to disentangle sources of variation in spatial transcriptomics data and allow scientists to generate additional insights for the biological system of interest. Similar tissue hallmarks that can be inferred from image data and may be used to explain gene expression variation, include blood vessels, tissue boundaries and fibrotic areas. Squidpy’s integrative analysis workflows leverage the spatial context and large microscopy images to generate new hypothesis classes in spatial transcriptomics data, thus bridging tissue-level characterizations of samples, which are typical in pathology, with the new high-resolution gene expression characterization yielded by spatial transcriptomics.
Discussion
In summary, Squidpy enables analysis of spatial molecular data by leveraging two data representations: the spatial graph and the tissue image. Squidpy infrastructure leverages sparse and memory-efficient implementations and its core spatial statistics and image analysis methods are fast and computationally efficient, making them suitable for the increasing size of modern spatial omics data. It interfaces with Scanpy and the Python data science ecosystem, providing a scalable and extendable framework for development of new methods in the field of biological spatial molecular data. Squidpy’s rich documentation in the form of functional application programming interface documentation, examples and tutorial workflows, is easy to navigate and is accessible to both experienced developers and beginner analysts. Furthermore, Squidpy is equipped with an extensive testing suite, implemented in a robust continuous integration pipeline. We foresee in the development roadmap support for GPU-accelerated workflows (specifically using Dask) and a tighter integration with the developing ecosystem of spatial omics methods, with explicit addition of an external module with methods and wrappers provided by contributors and additional tutorials and best practices of the nascent field of spatial omics data analysis. We hope that Squidpy will serve as a bridge between the molecular omics community and the image analysis and computer vision community to develop the next generation of computational methods for spatial omics technologies.
Methods
Infrastructure
Spatial graph
The spatial graph is a graph of spatial neighbors with cells (or spots in case of Visium) as nodes and neighborhood relations between spots as edges. We use spatial coordinates of spots to identify neighbors among them. Different approaches of defining a neighborhood relation among spots are used for different types of spatial datasets.
Visium spatial datasets have a hexagonal outline for their spots (each spot has up to eight spots situated around it). For this type of spatial dataset the parameter n_rings should be used. It specifies for each spot how many hexagonal rings of spots around it will be considered as neighbors.
sq.gr.spatial_neighbors(adata, coord_type=“grid”, n_neigh=6, n_rings=<int>)
It is also possible to create other types of grid-like graphs, such as squares, by changing the n_neigh argument. For a fixed number of the closest spots for each spot, it leverages the k-nearest neighbors search from Scikit-learn49 and n_neigh must be used to set the number of neighbors.
sq.gr.spatial_neighbors(adata, coord_type=“generic”, n_neigh=<int>)
To get all spots within a specified radius (in units of the spatial coordinates) from each spot as neighbors, the parameter radius should be used.
sq.gr.spatial_neighbors(adata, coord_type=“generic”, radius=<float>)
Finally, it is also possible to compute a neighbor graph based on Delaunay triangulation50.
sq.gr.spatial_neighbors(adata, coord_type=“generic”, delaunay=True)
The function builds a spatial graph and saves its adjacency and weighted adjacency matrices to adata.obsp[‘spatial_connectivities’] in either Numpy51 or Scipy sparse arrays18. The weights of the weighted adjacency matrix are distances in the case of coord_type = ‘generic’ and ordinal numbers of hexagonal rings in the case of coord_type = ‘grid’. Together with the connectivities, we also provide a sparse adjacency matrix of distances, saved in adata.obsp[‘spatial_distances’] We also provide spectral and cosine transformation of the adjacency matrix for uses in graph convolutional networks52.
ImageContainer
ImageContainer is an object for microscopy tissue images associated with spatial molecular datasets. The object is a thin wrapper of an xarray.Dataset37 and provides efficient access to in-memory and on-disk images. On-disk files are loaded lazily using dask19, meaning content is only read in memory when requested. The object can be saved as a zarr53 store. This allows handling of very large files that do not fit in the memory. The images represented by ImageContainer are required to have at least two dimensions, x and y, with an optional z dimension and a variable channels dimension.
ImageContainer is initialized with an in-memory array or a path to an image file on disk. Images are saved with the key layer. If lazy loading is desired, the lazy parameter needs to be specified.
sq.im.ImageContainer(PATH, layer=<str>, lazy=<bool>)
More image layers with the same spatial dimensions x, y and z such as segmentation masks can be added to an existing ImageContainer.
img.add_img(PATH, layer=<str>)
ImageContainer is able to interface with Anndata objects to relate any pixel-level information to the observations stored in Anndata (such as cells and spots). For instance, it is possible to create a lazy generator that yields image crops on-the-fly corresponding to locations of the spots in the image:
spot_generator = img.generate_spot_crops(adata) lambda x: (x for x in spot_generator) # yields crops at spots location
This of course works for both features computed at crop-level but also at segmentation-object level. For instance, it is possible to get centroid coordinates as well as several features of the segmentation object that overlap with the spot capture area.
Napari for interactive visualization
Napari is a fast, interactive, multi-dimensional image viewer in Python22. In Squidpy, it is possible to visualize the source image together with any Anndata annotation with napari. Such functionality is useful for fast and interactive exploration of analysis results saved in Anndata together with the high-resolution image. If multiple z dimensions are available, the individual z layers that can be interactively scrolled through. Furthermore, leveraging napari functionalities, it is possible to manually annotate tissue areas and assign underlying spots to annotations saved in the Anndata object. Such ability to relate manually defined tissue areas to observations in Anndata is particularly useful in settings where there is a pathologist annotation available and it needs to be integrated with analysis at gene expression or image level. All the steps described here are performed in Python, therefore available in the same environment where the analysis is performed (it does not require an additional tool).
img = sq.im.ImageContainer(PATH, layer=) img.interactive(adata)
Graph and spatial patterns analysis
Neighborhood enrichment test
The association between label pairs in the connectivity graph is estimated by counting the sum of nodes that belong to classes i and j (for example cluster annotation) and are proximal to each other, noted xij. To estimate the deviation of this number versus a random configuration of cluster labels in the same connectivity graph, we scramble the cluster labels while maintaining the connectivities and then recount the number of nodes recovered in each iteration (1,000 times by default). Using these estimates, we calculate expected means (µij) and standard deviations (σij) for each pair and a z score as,
The z score indicates if a cluster pair is over-represented or over-depleted for node–node interactions in the connectivity graph. This approach was described by Schapiro et al.54. The analysis and visualization can be performed with the analysis code shown below.
sq.gr.nhood_enrichment(adata, cluster_key=“<cluster_key>”) sq.pl.nhood_enrichment(adata, cluster_key=“<cluster_key>”)
Our implementation leverages just-in-time compilation with Numba55 to achieve greater performances in computation time (Supplementary Fig. 1).
Ligand–receptor interaction analysis
We provide a re-implementation of the popular CellphoneDB method for ligand–receptor interaction analysis43. In short, it is a permutation-based test of ligand–receptor expression across cell-type combinations. Given a list of annotated ligand–receptor pairs, the test computes the mean expression of the two molecules (ligand, receptor) between cell types and builds a null-distribution based on n permutations (default 1,000). A P value is computed based on the proportion of the permuted means against the true mean. In CellphoneDB, if a receptor or ligand is composed of several subunits, the minimum expression is considered for the test. In our implementation, we also include the option of taking the mean expression of all molecules in the complex. Our implementation also employs Omnipath44 as ligand–receptor interaction database annotation. A larger database that contains the original CellphoneDB database together with five other resources44. Finally, our implementation leverages just-in-time compilation with Numba55 to achieve greater performances in computation time (Supplementary Fig. 1).
Ripley’s spatial statistics
Ripley’s spatial statistics is a family of spatial analysis methods used to describe whether points with discrete annotation in space follow random, dispersed or clustered patterns. Ripley’s statistics can be used to describe the spatial patterning of cell clusters in the area of interest. In Squidpy, we re-implemented three of Ripley’s statistics: F, G and L functions. Ripley’s L function is a variance-stabilized transformation of Ripley’s K function, defined as
Where I(di,j < t) is the indicator function that sets whether the operand is 1 or 0 based on the (Euclidean) distance di,j evaluated at search radius t, A is the average density of point in the area of interest. Therefore, the Ripley’s L function is defined as:
The Ripley’s F and G functions are defined as:
Where di,j is the distance of the point to a random points for a spatial Poisson point process for F and the distance to any other point of the dataset for G. They can be easily computed with:
sq.gr.ripley(adata, cluster_key=“<cluster_key>”, mode=“<F|G|L>”) sq.pl.ripley(adata, cluster_key=“<cluster_key>”, mode=“<F|G|L>”)
Cluster co-occurrence ratio
Cluster co-occurrence ratio provides a score on the co-occurrence of clusters of interest across spatial dimensions. It is defined as
where cluster is the annotation of interest to be used as conditioning for the co-occurrence of all clusters. It is computed across n radius of size d across the tissue area. It was inspired by an analysis performed by Tosti et al. to investigate tissue organization in the human pancreas with spatial transcriptomics56.
sq.gr.co_occurrence(adata, cluster_key=“<cluster_key>”) sq.pl.co_occurrence(adata, cluster_key=“<cluster_key>”)
Spatial autocorrelation statistics
Spatial autocorrelation statistics are widely used in spatial data analysis tools to assess the spatial autocorrelation of continuous features. Given a feature (gene) and spatial location of observations, it evaluates whether the pattern expressed is clustered, dispersed or random57. In Squidpy, we implement two spatial autocorrelation statistics: Moran’s I and Geary’s C.
Moran’s I is defined as:
and Geary’s C is defined as:
where zi is the deviation of the feature from the mean \(\left( {x_i - \bar X} \right)\), wi,j is the spatial weight between observations, n is the number of spatial units and W is the sum of all wi,j. Test statistics and P values (computed from a permutation-based test or via analytic formulation, similar to libpysal58 and further FDR-corrected) are stored in adata.uns[‘moranI’] or adata.uns[‘gearyC’].
sq.gr.spatial_autocorr(adata, cluster_key=“<cluster_key>”,mode=“<moran|geary>”)
Sepal
Sepal is a recently developed method for spatially variable genes identification42. It simulates a diffusion process and evaluates the time it takes to reach a uniform state (convergence). It is a formulation of Fick’s second law to a regular graph (grid). It is defined as:
Where u(x,y,t) is the concentration (for example gene expression on a node in x,y coordinates), D is the diffusion coefficient, t is the update time and ∆(u(x,y,t)dt) is the laplacian on the graph (see elsewhere42 for an extended formulation). Convergence is reached if the change in entropy is below a given threshold:
The time t the gene takes to reach consensus is then used a ‘Sepal score’ and indicates the degree of spatial variability of the gene. It can be computed with:
sq.gr.sepal(adata)
Our re-implementation in Numba achieves greater computational efficiency (Supplementary Fig. 1).
Centrality scores
Centrality scores provide a numerical analysis on node patterns in the graph, which helps to better understand complex dependencies in large graphs. A centrality is a function C which assigns every vertex v in the graph a numeric value C(v) ∈ R. It therefore gives a ranking of the single components (cells) in the graph, which simplifies to identify key individuals. Group centrality measures have been introduced by Everett and Borgatti59. They provide a framework to assess clusters of cells in the graph (a specific cell type more central or more connected in the graph than others). Let G = (V,E) be a graph with nodes V and edges E. Additionally, let S be a group of nodes allocated to the same cluster cS. Then N(S) defines the neighborhood of all nodes in S. The following four (group) centrality measures are implemented. Group degree centrality is defined by the fraction of noncluster members that are connected to cluster members, so
Larger values indicate a more central cluster. Group degree centrality can help to identify essential clusters or cell types in the graph. Group closeness centrality measures how close the cluster is to other nodes in the graph and is calculated by the number of nongroup members divided by the sum of all distances from the cluster to all vertices outside the cluster, so
where dS,v = minu∈S du,v is the minimal distance of the group S from v. Hence, larger values indicate a greater centrality. Group betweenness centrality measures the proportion of shortest paths connecting pairs of nongroup members that pass through the group. Let S be a subset of a graph with vertex set VS. Let gu,v be the number of shortest paths connecting u to v and gu,v(S) be the number of shortest paths connecting u to v passing through S. The group betweenness centrality is then given by
The properties of this centrality score are fundamentally different from degree and closeness centrality scores, hence results often differ. The last measure described is the average clustering coefficient. It describes how well nodes in a graph tend to cluster together. Let n be the number of nodes in S. Then the average clustering coefficient is given by
with T(v) being the number of triangles through node v and deg(v) the degree of node v. The described centrality scores have been implemented using the NetworkX library in Python50.
sq.gr.centrality_scores(adata, cluster_key=“<cluster_key>”) sq.pl.centrality_scores(adata, cluster_key=“<cluster_key>”, selected_score=“<selected_score>”)
Interaction matrix represents the total number of edges that are shared between nodes with specific attributes (such as clusters or cell types).
sq.gr.interaction_matrix(adata, cluster_key=“<cluster_key>”, normalized=True) sq.pl.interaction_matrix(adata, cluster_key=“<cluster_key>”)
Python implementations rely on the NetworkX library50.
Image analysis and segmentation
Image processing
Before extracting features from microscopy images, the images can be preprocessed. Squidpy implements functions for commonly used preprocessing functions like conversion to grayscale or smoothing using a Gaussian kernel. In addition, custom processing functions can be used by passing a function to the method argument.
sq.im.process(img, method=“gray”) img.show()
Implementations are based on the Scikit-image package21 and allow lazy processing of very large images through tiling the image into smaller crops and processing these by using Dask. When using tiling, image crops are slightly overlapping, to reduce border effects.
Image segmentation
Nuclei segmentation is an important step when analyzing microscopy images. It allows the quantitative analysis of the number of nuclei, their areas and morphological features. There are a wide range of approaches for nuclei segmentation, from established techniques such as thresholding to modern deep-learning-based approaches.
A difficulty for nuclei segmentation is to distinguish between partially overlapping nuclei. Watershed is a classic algorithm used to separate overlapping objects by treating pixel values as local topology. For this, starting from points of lowest intensity, the image is flooded until basins from different starting points meet at the watershed ridge lines.
sq.im.segment(img, method=“watershed”) img.show()
Implementations in Squidpy are based on the original Scikit-image Python implementation21.
Custom approaches with deep-learning
Depending on the quality of the data, simple segmentation approaches like watershed might not be appropriate. Nowadays, many complex segmentation algorithms are provided as pretrained deep-learning models, such as Stardist15, Splinedist60 and Cellpose16. These models can be easily used within the segmentation function. We provide extensive tutorials https://squidpy.readthedocs.io/en/latest/tutorials.html#external-tutorials, where we show how Stardist15 and Cellpose16 can be easily interfaced with Squidpy to perform segmentation on both H&E and fluorescence images.
sq.im.segment(img, method=<pre-trained model>) img.show()
Image features
Tissue organization in microscopic images can be analyzed with different image features. This filters relevant information from the (high-dimensional) images, allowing for easy interpretation and comparison with other features obtained at the same spatial location. Image features are calculated from the tissue image at each location (x,y) where there is transcriptomics information available, resulting in an obs × features matrix similar to the obs × gene matrix. This image feature matrix can then be used in any single-cell analysis workflow, just like the gene matrix.
The scale and size of the image used to calculate features can be adjusted using the scale and spot_scale parameters. Feature extraction can be parallelized by providing n_jobs (see Supplementary Fig. 1). The calculated feature matrix is stored in adata[key].
sq.im.calculate_image_features(adata, img, features=<list>, spot_scale=<float>, scale=<float>, key_added=<str>)
Summary features calculate the mean, the s.d. or specific quantiles for a color channel. Similarly, histogram features scan the histogram of a color channel to calculate quantiles according a defined number of bins.
sq.im.calculate_image_features(adata, img, features=“summary”) sq.im.calculate_image_features(adata, img, features=“histogram”)
Textural features calculate statistics over a histogram that describes the signatures of textures. To grasp the concept of texture intuitively, the inextricable relationship between texture and tone is considered61; if a small-area patch of an image has little variation in its gray tone the dominant property of that area is tone. If the patch has a wide variation of gray-tone features, the dominant property of the area is texture. An image has a simple texture if it consists of recurring textural features. For a gray-level image I or for example a fluorescence color channel, a co-occurrence matrix C is computed. C is a histogram over pairs of pixels (i,j) with specific values (p,q) ∈ [0,1,…,255] (https://squidpy.readthedocs.io/en/latest/tutorials.html#external-tutorials) and a fixed pixel offset:
with Kronecker delta δ. The offset is a fixed pixel distance from i to j under a fixed direction angle. Based on the co-occurrence matrix different meaningful statistics (texture properties) can be calculated that summarize textural pattern characteristics of the image:
-
\(\mathop {\sum }\limits_{p,q} C_{p,q}\left( {p - q} \right)^2\) contrast
-
\(\mathop {\sum }\limits_{p,q} C_{p,q}\left| {p - q} \right|\) dissimilarity
-
\(\mathop {\sum }\limits_{p,q} \frac{{C_{p,q}}}{{1 + \left( {p - q} \right)^2}}\) homogeneity
-
\(\mathop {\sum }\limits_{p,q} C_{p,q}^2\) angular second moment
-
\(\mathop {\sum }\limits_{p,q} C_{p,q}\frac{{\left( {p - \mu _p} \right)\left( {q - \mu _q} \right)}}{{\sqrt {\sigma _p^2\sigma _q^2} }}\) correlation
sq.im.calculate_image_features(adata, img, features=“texture”)
All the above implementations rely on the Scikit-image Python package21.
Segmentation features
Similar to image features that are extracted from raw tissue images, segmentation features can be extracted from a segmentation object. These features allow to get statistics over the number, area and morphology of the nuclei in one image. To compute these features, the ImageContainer img needs to contain a segmented image at layer <segmented_img>
sq.im.calculate_image_features(adata, img,features=“segmentation”, features_kwargs={“label_layer”:<segmented_img>})
Custom features based on deep-learning models
Squidpy feature calculation function can also be used with custom user-defined features extraction functions. This enables the use of for example, pretrained deep-learning models as feature extractors. We provide tutorials https://squidpy.readthedocs.io/en/latest/tutorials.html#external-tutorials on how to interface popular deep-learning frameworks such as Tensorflow62 with ImageContainer, thus enabling users to perform an end-to-end deep-learning pipeline from Squidpy.
sq.im.calculate_image_features(adata, img, features=“custom”, features_kwargs={“func”:<pre-trained keras model>})
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The preprocessed datasets have been deposited at https://doi.org/10.6084/m9.figshare.c.5273297.v1 and they are all conveniently accessible in Python via the squidpy.dataset module. The datasets used in this article are the following: Imaging Mass Cytometry36, seqFISH33, 4i31, MERFISH34, SlideseqV2 (ref. 35), Mibi-tof41 and several Visium24 datasets available from https://support.10xgenomics.com/spatial-gene-expression/datasets. Information on preprocessing of such datasets can be found in Online Methods and code to reproduce it is at https://github.com/theislab/squidpy_reproducibility.
Code availability
Squidpy is a pip installable Python package and available at the following GitHub repository: https://github.com/theislab/squidpy, with documentation at: https://squidpy.readthedocs.io/en/latest/. All the code to reproduce the result of the analysis can be found at the following GitHub repository: https://github.com/theislab/squidpy_reproducibility.
References
Regev, A. et al. The human cell atlas. eLife 6, e27041 (2017).
Larsson, L., Frisén, J. & Lundeberg, J. Spatially resolved transcriptomics adds a new dimension to genomics. Nat. Methods 18, 15–18 (2021).
Zhuang, X. Spatially resolved single-cell genomics and transcriptomics by imaging. Nat. Methods 18, 18–22 (2021).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: Single cells, many features. Cell 165, 780–791 (2016).
Axelrod, S. et al. Starfish: open source image-based transcriptomics and proteomics tools. http://github.com/spacetx/starfish (2018).
Prabhakaran, S., Nawy, T. & Pe’er’, D. Sparcle: assigning transcripts to cells in multiplexed images. Preprint at BioRxiv https://doi.org/10.1101/2021.02.13.431099 (2021).
Park, J. et al. Cell segmentation-free inference of cell types from in situ transcriptomics data. Nat. Commun. 12, 3545 (2021).
Petukhov, V. et al. Cell segmentation in imaging-based spatial transcriptomics. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01044-w (2021).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Righelli, D. et al. SpatialExperiment: infrastructure for spatially resolved transcriptomics data in R using Bioconductor. Preprint at BioRxiv https://doi.org/10.1101/2021.01.27.428431 (2021).
Pham, D. et al. stLearn: integrating spatial location, tissue morphology and gene expression to find cell types, cell–cell interactions and spatial trajectories within undissociated tissues. Preprint at BioRxiv https://doi.org/10.1101/2020.05.31.125658 (2020).
Bergenstråhle, J., Larsson, L. & Lundeberg, J. Seamless integration of image and molecular analysis for spatial transcriptomics workflows. BMC Genomics 21, 482 (2020).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00935-2 (2021).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. in Medical Image Computing and Computer Assisted Intervention – MICCAI 2018 265–273 (Springer International Publishing, 2018).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Solorzano, L., Partel, G. & Wählby, C. TissUUmaps: interactive visualization of large-scale spatial gene expression and tissue morphology data. Bioinformatics 36, 4363–4365 (2020).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Dask Development Team. Dask: library for dynamic task scheduling. https://docs.dask.org/en/stable (2016).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Sofroniew, N. et al. napari/napari: 0.4.4rc0. https://doi.org/10.5281/zenodo.4470554 (2021).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
10X Genomics. Visium spatial gene expression reagent kits user guide. https://support.10xgenomics.com/spatial-gene-expression/library-prep/doc/user-guide-visium-spatial-gene-expression-reagent-kits-user-guide (2021).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Keren, L. et al. MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci. Adv. 5, eaax5851 (2019).
Lin, J.-R., Fallahi-Sichani, M., Chen, J.-Y. & Sorger, P. K. Cyclic immunofluorescence (CycIF), a highly multiplexed method for single-cell imaging. Curr. Protoc. Chem. Biol. 8, 251–264 (2016).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Alexandrov, T. Spatial metabolomics and imaging mass spectrometry in the age of artificial intelligence. Annu. Rev. Biomed. Data Sci. 3, 61–87 (2020).
Lohoff, T. et al. Integration of spatial and single-cell transcriptomic data elucidates mouse organogenesis. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01006-2 (2021).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0739-1 (2020).
Jackson, H. W. et al. The single-cell pathology landscape of breast cancer. Nature 578, 615–620 (2020).
Hoyer, S. & Hamman, J. J. xarray: N-D labeled arrays and datasets in Python. J. Open Res. Softw. https://doi.org/10.5334/jors.148 (2017).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Biancalani, T. et al. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat. Methods 18, 1352–1362 (2021).
Kleshchevnikov, V. et al. Comprehensive mapping of tissue cell architecture via integrated single cell and spatial transcriptomics. Preprint at bioRxiv https://doi.org/10.1101/2020.11.15.378125 (2020).
Hartmann, F. J. et al. Single-cell metabolic profiling of human cytotoxic T cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0651-8 (2020).
Anderson, A. & Lundeberg, J. sepal: identifying transcript profiles with spatial patterns by diffusion-based modeling. Bioinformatics https://doi.org/10.1093/bioinformatics/btab164 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Türei, D. et al. Integrated intra‐ and intercellular signaling knowledge for multicellular omics analysis. Mol. Syst. Biol. 17, e9923 (2021).
Dimitrov, D. et al. Comparison of resources and methods to infer cell-cell communication from single-cell RNA data. Preprint at bioRxiv https://doi.org/10.1101/2021.05.21.445160 (2021).
Amunts, K., Mohlberg, H., Bludau, S. & Zilles, K. Julich-Brain: a 3D probabilistic atlas of the human brain’s cytoarchitecture. Science 369, 988–992 (2020).
Ortiz, C., Carlén, M. & Meletis, K. Spatial transcriptomics: molecular maps of the mammalian brain. Annu. Rev. Neurosci. https://doi.org/10.1146/annurev-neuro-100520-082639 (2021).
Kandel, E., Koester, J. D., Mack, S. H. & Siegelbaum, S. Principles of Neural Science 6th edn (McGraw-Hill Education, 2021).
Pedregosa, F., Varoquaux, G. & Gramfort, A. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Hagberg, A., Swart, P. & S Chult, D. Exploring network structure, dynamics, and function using networkx. https://www.osti.gov/biblio/960616 (2008).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. Preprint at arXiv https://arxiv.org/abs/1609.02907 (2016).
Miles, A. et al. zarr-developers/zarr-python: v2.4.0. (2020). https://doi.org/10.5281/zenodo.3773450
Schapiro, D. et al. histoCAT: analysis of cell phenotypes and interactions in multiplex image cytometry data. Nat. Methods 14, 873–876 (2017).
Lam, S. K., Pitrou, A. & Seibert, S. Numba: a LLVM-based Python JIT compiler. In Proc. 2nd Workshop on the LLVM Compiler Infrastructure in HPC 1–6 (Association for Computing Machinery, 2015).
Tosti, L. et al. Single-nucleus and in situ RNA-sequencing reveal cell topographies in the human pancreas. Gastroenterology 160, 1330–1344 (2021).
Getis, A. & Ord, J. K. The analysis of spatial association by use of distance statistics. Geogr. Anal. 24, 189–206 (2010).
Rey, S. J. & Anselin, L. PySAL: a python library of spatial analytical methods. Rev. Reg. Stud. 37, 5–27 (2007).
Borgatti, S. P., Everett, M. G. & Johnson, J. C. Analyzing Social Networks (SAGE Publications, 2013).
Mandal, S. & Uhlmann, V. Splinedist: automated cell segmentation with spline curves. In 2021 IEEE 18th International Symposium on Biomedical Imaging (ISBI) 1082–1086 (IEEE, 2021).
Haralick, R. M., Shanmugam, K. & Dinstein, I. Textural features for image classification. Stud. Media Commun. SMC 3, 610–621 (1973).
Abadi, M. et al. TensorFlow: A system for large-scale machine learning. In 12th USENIX symposium on operating system design and implementation (OSDI 16), 265–283 (2016).
Acknowledgements
We acknowledge L. Zappia, M. Luecken and all members of the Theis laboratory for helpful discussion. We acknowledge G. Fornons for the Squidpy logo, Scanpy developers P. Angerer and F. Ramirez for useful discussion and code revision, A. Goeva and E. Macosko for cell-type annotation of the SlideseqV2 dataset, A. Andersson for general feedback and pointers to the Sepal implementation, L. Hetzel for draft of the spatial graph Delaunay method, M. Varrone for fixing a bug in the graph Delaunay method and G. Buckley for pointers to Dask image functionalities. We thank authors of original publications and 10X Genomics for making spatial omics datasets publicly available. S. Richter and G.P. are supported by the Helmholtz Association under the joint research school Munich School for Data Science. D.S.F. acknowledges support from a German Research Foundation fellowship through the Graduate School of Quantitative Biosciences Munich (GSC 1006 to D.S.F.) and by the Joachim Herz Foundation. A.C.S. has been funded by the German Federal Ministry of Education and Research under grant no. 01IS18036B. F.J.T. acknowledges support by the BMBF (grant nos. 01IS18036B, 01IS18053A and 031L0210A), the European Union’s Horizon 2020 Research and Innovation programme under grant agreement no. 874656, the Chan Zuckerberg Initiative DAF (advised fund of Silicon Valley Community Foundation, grant no. 2019-207271), the Bavarian Ministry of Science and the Arts in the framework of the Bavarian Research Association ‘ForInter’ (Interaction of human brain cells) and by the Helmholtz Association’s Initiative and Networking Fund through Helmholtz AI (grant no. ZT-I-PF-5-01) and by the Chan Zuckerberg foundation (grant no. 2019-002438, Human Lung Cell Atlas 1.0) and sparse2big (grant no. ZT-I-007).
Funding
Open access funding provided by Helmholtz Zentrum München - Deutsches Forschungszentrum für Gesundheit und Umwelt (GmbH).
Author information
Authors and Affiliations
Contributions
G.P., H.S., M.K., D.F. and F.J.T. designed the study. G.P., H.S., M.K., D.F., A.C.S., L.B.K., S. Rybakov, I.L.I., O.H., I.V., M.L. and S. Richter wrote the code. G.P., H.S. and M.K. performed the analysis. F.J.T. supervised the work. All authors read and corrected the final manuscript.
Corresponding author
Ethics declarations
Competing interests
"F.J.T. consults for Immunai Inc., Singularity Bio B.V., CytoReason Ltd, and Omniscope Ltd, and has ownership interest in Dermagnostix GmbH and Cellarity.
Peer review
Peer review information
Nature Methods thanks Raphael Gottardo and the other, anonymous, reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Lin Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 Supplementary methods.
Supplementary Table 1 and 2
File with two tables.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Palla, G., Spitzer, H., Klein, M. et al. Squidpy: a scalable framework for spatial omics analysis. Nat Methods 19, 171–178 (2022). https://doi.org/10.1038/s41592-021-01358-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01358-2
This article is cited by
-
Bento: a toolkit for subcellular analysis of spatial transcriptomics data
Genome Biology (2024)
-
DANCE: a deep learning library and benchmark platform for single-cell analysis
Genome Biology (2024)
-
SRT-Server: powering the analysis of spatial transcriptomic data
Genome Medicine (2024)
-
SpatialData: an open and universal data framework for spatial omics
Nature Methods (2024)
-
Streamlining spatial omics data analysis with Pysodb
Nature Protocols (2024)