Abstract
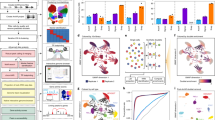
The recent development of experimental methods for measuring chromatin state at single-cell resolution has created a need for computational tools capable of analyzing these datasets. Here we developed Signac, a comprehensive toolkit for the analysis of single-cell chromatin data. Signac enables an end-to-end analysis of single-cell chromatin data, including peak calling, quantification, quality control, dimension reduction, clustering, integration with single-cell gene expression datasets, DNA motif analysis and interactive visualization. Through its seamless compatibility with the Seurat package, Signac facilitates the analysis of diverse multimodal single-cell chromatin data, including datasets that co-assay DNA accessibility with gene expression, protein abundance and mitochondrial genotype. We demonstrate scaling of the Signac framework to analyze datasets containing over 700,000 cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data used in the paper are publicly available. The PBMC multiomic dataset is available from 10X Genomics at https://support.10xgenomics.com/single-cell-multiome-atac-gex/datasets/1.0.0/pbmc_granulocyte_sorted_10k. The PBMC scATAC-seq datasets are available from 10X Genomics at https://support.10xgenomics.com/single-cell-atac/datasets. The synthetic scATAC-seq datasets are available from GitHub at https://github.com/pinellolab/scATAC-benchmarking. Data from the BICCN are available from the Neuroscience Multiomic Archive at https://nemoarchive.org/. Data for the CRC patient sample are available on NCBI Gene Expression Omnibus (GSE148509) and Zenodo (https://zenodo.org/record/3977808).
Code availability
Signac is available on CRAN (https://cloud.r-project.org/package=Signac) and on GitHub (https://github.com/timoast/signac), with documentation and tutorials available at https://satijalab.org/signac/. All code used in this paper is available on GitHub at https://github.com/timoast/signac-paper.
Change history
07 January 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41592-022-01393-7
References
Ai, S. et al. Profiling chromatin states using single-cell itChIP-seq. Nat. Cell Biol. 21, 1164–1172 (2019).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Carter, B. et al. Mapping histone modifications in low cell number and single cells using antibody-guided chromatin tagmentation (ACT-seq). Nat. Commun. 10, 3747 (2019).
Cusanovich, D. A. et al. Multiplex single cell profiling of chromatin accessibility by combinatorial cellular indexing. Science 348, 910–914 (2015).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Wang, Q. et al. CoBATCH for high-throughput single-cell epigenomic profiling. Mol. Cell https://doi.org/10.1016/j.molcel.2019.07.015 (2019).
Ku, W. L. et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nat. Methods 16, 323–325 (2019).
Lareau, C. A. et al. Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0147-6 (2019).
Luo, C. et al. Robust single-cell DNA methylome profiling with snmc-seq2. Nat. Commun. 9, 3824 (2018).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science https://doi.org/10.1126/science.aau0730 (2018).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0290-0 (2019).
Clark, S. J. et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 9, 781 (2018).
Ludwig, L. S. et al. Lineage tracing in humans enabled by mitochondrial mutations and Single-Cell genomics. Cell https://doi.org/10.1016/j.cell.2019.01.022 (2019).
Lareau, C. A. et al. Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0645-6 (2021).
Zhu, C. et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat. Struct. Mol. Biol. 26, 1063–1070 (2019).
Xing, Q. R. et al. Parallel bimodal single-cell sequencing of transcriptome and chromatin accessibility. Genome Res. 30, 1027–1039 (2020).
Liu, L. et al. Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity. Nat. Commun. 10, 470 (2019).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell https://doi.org/10.1016/j.cell.2020.09.056 (2020).
Mimitou, E. P. et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00927-2 (2021).
Fiskin, E., Lareau, C. A., Eraslan, G., Ludwig, L. S. & Regev, A. Single-cell multimodal profiling of proteins and chromatin accessibility using PHAGE-ATAC. Preprint at BioRxiv https://doi.org/10.1101/2020.10.01.322420 (2020).
Swanson, E. et al. Simultaneous trimodal single-cell measurement of transcripts, epitopes, and chromatin accessibility using TEA-seq. eLife 10, e63632 (2021).
Rubin, A. J. et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell 176, 361–376 (2019).
Pierce, S. E., Granja, J. M. & Greenleaf, W. J. High-throughput single-cell chromatin accessibility CRISPR screens enable unbiased identification of regulatory networks in cancer. Nat. Commun. 12, 2969 (2021).
Thornton, C. A. et al. Spatially mapped single-cell chromatin accessibility. Nat. Commun. 12, 1274 (2021).
Stuart, T. & Satija, R. Integrative single-cell analysis. Nat. Rev. Genet. https://doi.org/10.1038/s41576-019-0093-7 (2019).
Bravo González-Blas, C. et al. cistopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods https://doi.org/10.1038/s41592-019-0367-1 (2019).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324 (2018).
Xiong, L. et al. SCALE method for single-cell ATAC-seq analysis via latent feature extraction. Nat. Commun. 10, 4576 (2019).
Pliner, H. A. et al. Cicero predicts cis-regulatory DNA interactions from Single-Cell chromatin accessibility data. Mol. Cell 71, 858–871.e8 (2018).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Danese, A. et al. EpiScanpy: integrated single-cell epigenomic analysis. Nat.Commun. https://doi.org/10.1038/s41467-021-25131-3 (2021).
Fang, R. et al. Comprehensive analysis of single cell ATAC-seq data with SnapATAC. Nat. Commun. 12, 1337 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. https://doi.org/10.1038/s41588-021-00790-6 (2021).
Ji, Z., Zhou, W. & Ji, H. Single-cell regulome data analysis by SCRAT. Bioinformatics 33, 2930–2932 (2017).
Baker, S. M., Rogerson, C., Hayes, A., Sharrocks, A. D. & Rattray, M. Classifying cells with scasat, a single-cell ATAC-seq analysis tool. Nucleic Acids Res. 47, e10 (2019).
Zhao, C., Hu, S., Huo, X. & Zhang, Y. Dr.seq2: a quality control and analysis pipeline for parallel single cell transcriptome and epigenome data. PLoS ONE 12, e0180583 (2017).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. https://doi.org/10.1038/nbt.4096 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Xu, J. et al. Single-cell lineage tracing by endogenous mutations enriched in transposase accessible mitochondrial DNA. eLife https://doi.org/10.7554/eLife.45105 (2019).
Li, H. Tabix: fast retrieval of sequence features from generic TAB-delimited files. Bioinformatics 27, 718–719 (2011).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Deerwester, S., Dumais, S. T., Furnas, G. W., Landauer, T. K. & Harshman, R. Indexing by latent semantic analysis. J. Am. Soc. Inf. Sci. 41, 391–407 (1990).
McInnes, L. & Healy, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at arXiv https://arXiv.org/abs/1802.03426 (2018).
Pearce, E. L. et al. Control of effector CD8+ T cell function by the transcription factor eomesodermin. Science 302, 1041–1043 (2003).
Corces, M. R. et al. The chromatin accessibility landscape of primary human cancers. Science https://doi.org/10.1126/science.aav1898 (2018).
GTEx Consortium. The GTEx Consortium Atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Chen, H. et al. Assessment of computational methods for the analysis of single-cell ATAC-seq data. Genome Biol. 20, 241 (2019).
Li, Y. et al. An atlas of gene regulatory elements in adult mouse cerebrum. Preprint at bioRxiv https://doi.org/10.1101/2020.05.10.087585 (2020).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature https://doi.org/10.1038/s41586-019-0969-x (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with harmony. Nat. Methods https://doi.org/10.1038/s41592-019-0619-0 (2019).
Brenner, S. Sequences and consequences. Philos. Trans. R. Soc. Lond. B Biol. Sci. 365, 207–212 (2010).
Richmond, T. J. & Davey, C. A. The structure of DNA in the nucleosome core. Nature 423, 145–150 (2003).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Baglama, J. & Reichel, L. Augmented implicitly restarted Lanczos bidiagonalization methods. SIAM J. Sci. Comput. 27, 19–42 (2005).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 9354 (2019).
Waltman, L. & van Eck, N. J. A smart local moving algorithm for large-scale modularity-based community detection. Eur. Phys. J. B 86, 471 (2013).
Scrucca, L., Fop, M., Murphy, T. B. & Raftery, A. E. mclust 5: clustering, classification and density estimation using Gaussian finite mixture models. R. J. 8, 289–317 (2016).
Sing, T., Sander, O., Beerenwinkel, N. & Lengauer, T. ROCR: visualizing classifier performance in R. Bioinformatics 21, 3940–3941 (2005).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Hormozdiari, F., Kostem, E., Kang, E. Y., Pasaniuc, B. & Eskin, E. Identifying causal variants at loci with multiple signals of association. Genetics 198, 497–508 (2014).
Griffiths, J. A., Richard, A. C., Bach, K., Lun, A. T. L. & Marioni, J. C. Detection and removal of barcode swapping in single-cell RNA-seq data. Nat. Commun. 9, 2667 (2018).
Lun, A. T. L. et al. EmptyDrops: distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 20, 63 (2019).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at arXiv https://arxiv.org/abs/1303.3997 (2013).
Acknowledgements
This work was supported by the Chan Zuckerberg Initiative (EOSS-0000000082 and HCA-A-1704-01895 to R.S.) and the National Institutes of Health (DP2HG009623-01, RM1HG011014-01 and OT2OD026673-01 to R.S.; K99HG011489-01 to T.S.). C.A.L. was supported by a Stanford Science Fellowship. We are grateful to L. Ludwig (MDC Berlin) for insightful conversations about mtDNA lineage tracing. We thank B. Ren (UCSD) for assistance in accessing the BICCN mouse brain dataset. We thank the CRAN maintainers for their assistance in distributing the Signac R package and members of the Satija laboratory for feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
T.S. and A.S. developed the Signac package with guidance from R.S. R.S. supervised the research. T.S. and S.M. performed analyses. C.A.L. developed the mitochondrial lineage tracing methods and analysis. T.S. wrote the manuscript with input from all authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
In the past 3 years, R.S. has worked as a consultant for Bristol-Myers Squibb, Regeneron and Kallyope and served as an SAB member for ImmunAI, Resolve Biosciences, Nanostring, and the NYC Pandemic Response Lab. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks Junyue Cao and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling editor: Lin Tang, in collaboration with the Nature Methods team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Table 1
Comparison of single-cell chromatin analysis packages.
Rights and permissions
About this article
Cite this article
Stuart, T., Srivastava, A., Madad, S. et al. Single-cell chromatin state analysis with Signac. Nat Methods 18, 1333–1341 (2021). https://doi.org/10.1038/s41592-021-01282-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01282-5
This article is cited by
-
txci-ATAC-seq: a massive-scale single-cell technique to profile chromatin accessibility
Genome Biology (2024)
-
Mechanisms of action and resistance in histone methylation-targeted therapy
Nature (2024)
-
Single-cell multi-ome regression models identify functional and disease-associated enhancers and enable chromatin potential analysis
Nature Genetics (2024)
-
rworkflows: automating reproducible practices for the R community
Nature Communications (2024)
-
Uniform quantification of single-nucleus ATAC-seq data with Paired-Insertion Counting (PIC) and a model-based insertion rate estimator
Nature Methods (2024)