Abstract
Cytokines are critical for intercellular communication in human health and disease, but the investigation of cytokine signaling activity has remained challenging due to the short half-lives of cytokines and the complexity/redundancy of cytokine functions. To address these challenges, we developed the Cytokine Signaling Analyzer (CytoSig; https://cytosig.ccr.cancer.gov/), providing both a database of target genes modulated by cytokines and a predictive model of cytokine signaling cascades from transcriptomic profiles. We collected 20,591 transcriptome profiles for human cytokine, chemokine and growth factor responses. This atlas of transcriptional patterns induced by cytokines enabled the reliable prediction of signaling activities in distinct cell populations in infectious diseases, chronic inflammation and cancer using bulk and single-cell transcriptomic data. CytoSig revealed previously unidentified roles of many cytokines, such as BMP6 as an anti-inflammatory factor, and identified candidate therapeutic targets in human inflammatory diseases, such as CXCL8 for severe coronavirus disease 2019.
Similar content being viewed by others
Main
Cytokines are a broad category of intercellular signaling proteins that act in almost every aspect of human immunology, from anti-pathogen immune responses to tissue-damaging inflammation1,2. However, the precise characterization of cytokine signaling activities has proven difficult due to two vexing properties of cytokine activity: redundancy and pleiotropy. Many cytokines, especially those with similar cell surface receptors and downstream pathways, have cellular effects that appear redundant within a specific cellular context3. At the same time, cytokines often have pleiotropic functions within an organism that depend heavily on cell-type-specific receptor usage and the presence of other signaling components3.
This apparent redundancy and pleiotropy in cytokine activities are poorly captured by most immunological assays such as the enzyme-linked immunosorbent assay (ELISA) and Luminex xMAP, which directly measure cytokine release. Cytokine release can be transient, unlike the longer-lasting and more functionally relevant measurement of target activities4. Recognizing this limitation, researchers have attempted to create databases of cytokine signaling targets. For example, the ‘Interferome’ database identifies interferon target genes in humans and mice through the collection and analysis of microarray data5. Gene-set enrichment analysis (GSEA) also annotates response genes for selected cytokines based on previous knowledge6. However, these databases and approaches cover a small fraction of cytokines, leaving most cytokine-induced target changes unexplored.
The need for systematic profiling approaches that allow modeling of cytokine target activity is urgent because cytokines can trigger life-threatening symptoms in many diseases. For example, coronavirus disease 2019 (COVID-19) mortality has been attributed mainly to a virus-induced cytokine storm, defined by excessive production of pro-inflammatory cytokines that lead to acute respiratory distress and widespread tissue damage7. Although pro-inflammatory cytokines help activate the immune response, there does not appear to be a strong relationship between cytokine storm severity and pathogen clearance. For example, successfully recovering patients with COVID-19 may not have any inflammatory symptoms8. Cytokine release syndrome also causes severe side effects in many cancer treatments, such as immunotherapies9 and chimeric antigen receptor T cell therapies10. Similarly to the disconnect between the severity of immune-related symptoms and disease outcomes in COVID-19, complete tumor remission can occur in patients without cytokine release syndrome11. While the immunological mechanisms of these observations remain unclear, they imply that if properly modulated, the benefits of cytokine signaling can be realized without substantial pathological effects.
With this goal in mind, and to model cytokine activity generally, we developed CytoSig (https://cytosig.ccr.cancer.gov/), a data-driven infrastructure hosted by the National Cancer Institute (NCI). CytoSig includes both a database of target genes modulated by cytokines and a predictive model of cytokine signaling activity and regulatory cascade from transcriptomic profiles. To build the CytoSig platform, we first created the Framework for Data Curation (FDC) to assist expert annotations on metadata deposited on databases through natural language processing functions (https://curate.ccr.cancer.gov/). Using the FDC, we analyzed 9,271 published studies and curated 20,591 transcriptomic profiles for human cytokine, chemokine and growth factor responses to create the CytoSig database and predictive model. We validated CytoSig by showing that it can reliably predict cytokine target activities in both human clinical studies and our in vivo experiments. Further, CytoSig identified CXCL8 signaling as a potential COVID-19 therapeutic target that may alleviate adverse inflammation without undermining protective immunity.
Results
The Framework for Data Curation on public repositories
We hypothesized that the large number of cytokine treatment datasets available publicly could serve as a knowledge base to model signaling activities in diverse biological contexts. However, two hurdles must be overcome to transform this body of data into a useful model. First, the experimental design behind each published dataset is unique, requiring labor-intensive expert interpretation of the metadata and standardization of the data into a format suitable for automated analysis. Second, one must identify and exclude experiments that involve cell models, stimuli, doses or time intervals that are not physiologically relevant. More broadly, such challenges exist for many other biological topics that could be addressed by data aggregation. To overcome these hurdles, we established the FDC, which couples large-scale automatic data processing with natural language processing functions to assist expert annotation of experimental design (Methods and Fig. 1a).
a, The FDC can automatically process RNA-seq and MicroArray transcriptomic data from public repositories. Then, with the FDC’s natural language processing functions, curators read the metadata of each sample to annotate experimental conditions, including cytokine treatment, cell model, dose and duration. The output is differential logFC upon treatment. b, Two uses of the CytoSig framework: (1) query a gene name to view upstream cytokine regulators or downstream target genes (if the query is a cytokine); (2) predict cytokine signaling activities through transcriptomic profiles using a linear regression model. Input, the input transcriptomic profile of the sample as the response variable in regression; signature, the average response signature of cytokines as explanatory covariates; activity, the regression coefficients reflecting signaling activities. c, Count of treatment response profiles with biological replicates for different molecule types. d, Example correlation between the expression of IL-10 target genes and its ligand or receptor. Each dot represents a TCGA lung adenocarcinoma sample (n = 513). The x axis shows the expression of IL10 or receptor (IL-10RA + IL-10RB). The y axis presents the expression scores of IL-10 targets from a monocyte treatment experiment. Pearson correlation (r) indicates the human physiological relevance of the current data. e, Distribution of target score correlations. Correlations were computed using all cytokine-response profiles and TCGA or GTEx expression matrices. Distributions of correlations are shown by violin plots, smoothed by a kernel density estimator, for both real and randomized data through gene label permutations. The P value, calculated with the one-sided Wilcoxon signed-rank test, represents the statistical significance of correlations being higher than zero (n = 112 ligands and n = 111 receptors). f, Similarity of signaling response profiles. We created a composite signature for each cytokine that consisted of the median logFC across all experiments and then calculated pairwise correlations between composite signatures for the hierarchical clustering. Red branches highlight the clusters of similar cytokines discussed in the paper.
The FDC automatically extracts RNA-sequencing (RNA-seq) data from the Sequence Read Archive (SRA)12 and the European Nucleotide Archive (ENA)13, along with automatically extracting MicroArray data from the Gene Expression Omnibus (GEO)14 and ArrayExpress (AE)15. For metadata annotation, the FDC interacts with curators in iterative cycles. If the metadata structure and experimental designs differ drastically across studies, as was the case for cytokine-response data, the initial cycle of curation relies heavily on human expertise. However, based on the initial curations, the curators may specify automatic annotation rules, including highlighting text patterns that drive annotation decisions, translating aliases to standard names, and implementing controlled vocabularies. These natural language processing functions will dramatically reduce the human effort required after iterative cycles. The FDC is suitable for a wide range of data collection projects and is available at https://curate.ccr.cancer.gov/.
Generating the CytoSig database of cytokine-modulated genes
CytoSig aims to provide both a database of target genes modulated by cytokines and a predictive model of cytokine signaling activities from a sample’s transcriptomic profile (Fig. 1b). Both goals depend on an extensive data collection of cytokine-induced target genes. We first queried the AE and GEO databases with names and aliases of human cytokines, chemokines and growth factors. Note that, for brevity, we use the term ‘cytokine’ at times in this paper to refer to these three types of signaling molecules generally. The cytokine name search yielded 9,271 candidate studies. Of 9,271 candidates, 5,186 studies had genome-wide expression matrices and could be automatically processed by the FDC.
After automatic data extraction, Ph.D. scientists with immunology training conducted a curation of the 5,186 selected experiments (Fig. 1a). Each dataset was assigned two curators, such that the secondary curator would proofread annotations of the primary curator and correct any errors. This initial manual curation was time intensive because the metadata structure and experimental designs differed drastically across studies. However, based on the rules learned from the initial curation, the natural language processing functions from the FDC accelerated the annotation process such that minimal human effort was subsequently required. This semiautomated extraction system ensures that CytoSig will remain updated and relevant as new datasets are released.
Of the 5,186 experiments examined, 962 experiments were designated as cytokine-response studies, which comprised 20,591 nonredundant individual samples (Supplementary Table 1). Curators then labeled each sample with the treatment cytokine, the cell model, treatment dose and duration. We combined these human annotations with automatically parsed matrices of gene expression values and merged biological replicates, which generated 2,056 differential expression signatures between cytokine treatments and controls (Fig. 1c). Certain cell models and experimental conditions tended to be more frequently used than others (Extended Data Fig. 1a–f).
For target genes, a differential signature presents the direction of the expression change (up or down) and the magnitude of that change, expressed as log2 fold change (logFC), under each experimental condition. These differential signatures have continuous magnitude values. So, rather than using cutoffs to define cytokine targets, the differential magnitudes were used in our further analysis.
CytoSig data reflect signaling activity in human physiology
Because our datasets are generated through treatment experiments in cell cultures, we evaluated whether our collected cytokine targets are target genes under human physiological conditions. We measured the Pearson correlation between expression levels of the cytokine and its candidate targets in independent human tissue data. For example, we defined interleukin (IL)-10 targets based on an IL-10 treatment profile conducted in monocytes16 and then measured the correlation between the IL-10 expression and average expression scores of its candidate targets across tumors in a lung adenocarcinoma cohort17, which we found to be 0.68 (Fig. 1d). We also found that the expression correlation between IL-10 targets and IL-10 receptors (IL-10RA + IL-10RB) is 0.62 (Fig. 1d). As a control to evaluate correlations expected by random, permutation of gene identities of the IL-10 treatment profile ten times resulted in a low average correlation of 0.04 between IL-10 and its target genes and a low correlation of 0.05 between the IL-10 receptors and its targets.
We computed correlations in this way between the expression levels of each cytokine and its candidate targets for all 2,056 cytokine treatment profiles across The Cancer Genome Atlas (TCGA)18 and the Genotype-Tissue Expression (GTEx)19 cohorts. The distribution of correlations between the candidate target gene expression with the respective ligands and receptors was significantly higher than expected by chance (Fig. 1e). The correspondence between the expression of target genes and a cytokine ligand or receptor in independent human tissue samples suggests that our data collection is useful for modeling cytokine signaling events in human physiology.
Although most of the cytokine-response profiles derived from cell culture models are relevant in human in vivo settings, experimental conditions of some cytokines may not reflect physiological kinetics (Extended Data Fig. 1g and Supplementary Table 2). For analyses presented from this point forward, we only use differential expression signatures with significant positive correlations between expression levels of target genes and ligands or receptors in both TCGA and GTEx cohorts (false discovery rate (FDR) < 0.05; Methods). This criterion was met by 1,307 of 2,056 signatures.
For each cytokine, merging independent signatures can create a composite profile with superior performance than individual signatures as measured by the correlation metrics described above (Extended Data Fig. 1h). Each cytokine’s composite signature is composed of the median logFC across all experiments for each gene, reflecting target genes induced or repressed in most conditions. We compared the overall similarities of response by performing hierarchical clustering of the composite signatures of the 43 cytokines that had at least five high-quality independent expression profiles (Fig. 1f). A few subclusters contained cytokines with very high correlations. For example, the composite response signature of IL-27 was very similar to that of interferon gamma (IFN-γ), and to a lesser extent, to type I (IFN-I) and type III (IFNL) interferons. This observation is consistent with the downstream transcriptional similarity between IL-27 and IFN-γ, because they both act through STAT1 signaling20,21. Another cluster with high similarity contains tumor necrosis factor (TNF), IL-1A/IL-1B and CD40L (CD40L is both a soluble ligand and a cell surface molecule22), all of which activate NF-κB signaling23,24.
Although many cytokines have highly similar target responses, the same cytokine may also present context-specific differences in target response patterns. For example, the IFN-γ response signatures formed distinct clusters based on their cell origins. Macrophages and monocytes are clustered together and have different responses than other clusters, such as fibroblasts (Extended Data Fig. 1i).
Two regulatory cascades from primary to secondary cytokines
The hierarchical cascade of cytokine regulation is a paradigm in cellular signaling. For example, CXCL9, CXCL10 and CXCL11 (CXCR3 ligands) are immune-activating chemokines induced by IFN-γ25, which itself is regulated upstream by IL-12 (ref. 26). Within a signaling cascade, a cytokine can also inhibit downstream signals. For example, IL-4 can block IL-1 and TNF signaling in human monocytes27. These hierarchical activations and inhibitions are essential to ensure rapid clearance of different pathogen classes while at the same time preventing an overzealous immune response25,26,27. The activation and repression relationships among CXCL9/CXCL10/CXCL11, IFN-γ, IL-12, IL-4 and IL-1 according to the examples discussed above were all statistically significant in our dataset (Fig. 2a and Extended Data Fig. 2a).
a, Differential expression of target cytokines (x axis) in response to primary cytokines (y axis). Each dot represents a treatment profile. Blue solid points are profiles that pass quality controls, with numbers labeled on the top of box plots. The thick line represents the median value. The bottom and top of the boxes are the 25th and 75th percentiles (interquartile range). The whiskers encompass 1.5 times the interquartile range. The P value, testing whether group values are different from zero, was calculated using the two-sided Wilcoxon signed-rank test. b, Inter-cytokine regulation hierarchy. Between each pair of regulators (x axis) and targets (y axis), the logFC values are the medians among profiles that passed our quality controls. The plot only includes regulators and targets with at least one logFC value that was larger than two. The upper heat map includes target ligands, and the lower part includes target receptors with ligands in brackets. The NF-κB and interferon transcriptional groups are labeled with boxes. c, Differential logFC values of GZMA, GZMB and PRF1 upon TGF-β1 treatment, shown by violin plots, smoothed by a kernel density estimator. Each dot represents an independent profile. The P value, testing whether the T cell group values are different from zero, was calculated using the two-sided Wilcoxon signed-rank test (n = 11 independent treatment profiles for each gene). d, Perforin intracellular levels measured by flow cytometry upon TGF-β1 treatment in human primary CD8+ T cells. The x axis shows the signal intensity. The y axis shows the T cell fraction with modal normalization, scaling the maximum y axis to 100%. The vertical line indicates the percentage of T cells with signal intensity above the gate threshold (Extended Data Fig. 2d). e, Granzyme and perforin protein levels upon TGF-β treatment in human and mouse CD8+ T cells. The mean percentage of T cells with an intensity above the gate threshold is shown, with standard deviations as error bars (n = 3 cell culture replicates per condition). The two-sided Wilcoxon rank-sum test was used to compare groups.
We systematically examined the changes induced by primary cytokines on secondary signals and identified two distinct pro-inflammatory clusters (Fig. 2b). Each group activates a distinct set of secondary targets. The first group, including TNF, IL-1, IL-17A, IL-36 and CD40L, triggers IL-1, IL-6, CXCL1, CXCL2, CXCL5, CXCL6, CXCL8 and CCL20 (Fig. 2a,b). These primary cytokines have target genes enriched in the NF-κB signaling pathway (Extended Data Fig. 2b). The target chemokines of this group may attract or activate pro-inflammatory immune cells, such as neutrophils, fibroblasts and T cells28. We hereafter refer to this first group as the ‘NF-κB transcriptional group’.
The second group, including IFN-γ, IFN-I (IFN-α and IFN-β), IFNL and IL-27, trigger CXCL9, CXCL10, CXCL11 and TRAIL (Fig. 2b). These primary cytokines are known to be related to interferons because IL-27 has similar downstream transcriptional profiles with IFN-γ20 through STAT1 signaling21. Their secondary targets are chemokines for activated T cells (CXCL9, CXCL10 and CXCL11)25, or pro-apoptotic signals released by effector T cells (TRAIL)28. We hereafter refer to this second group as the ‘interferon transcriptional group’.
Besides regulating a ligand, cytokines can also modulate receptor activity as an alternative means of cascade regulation. For example, CytoSig found that activin A activates CXCR4, while GMCSF represses CXCR4 (Fig. 2a,b). This result is consistent with previous studies of CXCR4 regulation29,30. Regulation of receptors in this way appears less frequent than ligand regulation: we observed just 4 of 183 (2.2%) annotated receptors that had a logFC greater than two, whereas 33 of 253 (13%) annotated ligands had a logFC larger than two (Fig. 2b).
The CytoSig data reveal anti-inflammatory cytokines
In contrast to cytokines that induce secondary cytokines and chemokines, IL-4 and BMP6 repress other pro-inflammatory molecules, such as IL-1β, CXCL1, CXCL8 and CCL2 (Fig. 2b and Extended Data Fig. 2a). GSEA on target genes of IL-4 and BMP6 revealed a depletion of inflammatory response pathways (Extended Data Fig. 2b,c). IL-4 is a well-known anti-inflammatory cytokine that inhibits certain immune processes, although it may also cause allergic inflammation in a context-dependent manner31.
Besides IL-4 and BMP6, which directly suppress the transcription of downstream cytokines and chemokines, other anti-inflammatory molecules may counteract inflammation by alternative mechanisms. For example, a previous study in mouse models demonstrated that transforming growth factor (TGF)-β signaling directly targets cytotoxic T cell functions in mice32. Indeed, our collected data shows that TGF-β1 treatment in human T cells significantly downregulated granzyme A (GZMA), granzyme B (GZMB), and perforin (PRF1), which induce cell death in target cells attacked by T cells (Fig. 2c). Flow cytometry analysis in human and mouse primary T cells validated the inhibitory effect of TGF-β1 on GZMA, GZMB and PRF1 (Fig. 2d,e and Extended Data Fig. 2d; gating strategy). Therefore, our data can reveal broad categories of anti-inflammatory cytokines.
We next examined how cytokines cooperate with and antagonize each other with respect to target genes across the human genome. To test whether pairs of cytokines co-regulate target genes, we enumerated genes with significant logFC values from both cytokines under analysis. Then, we compared the gene counts against values when gene labels were shuffled to compute the FDR (Methods). We defined significant results by an FDR threshold of 0.05 (Fig. 3a). In 86% of statistically significant cases, cytokine pairs either enhanced or repressed target genes in concert (Fig. 3b). For example, TNF and IL-1β induced a similar set of genes and also repressed a similar set of genes when investigating average targets across all models in our data collection (Fig. 3a).
a, Co-regulation between IL-1β and other cytokines on target genes. Each dot represents a target gene. The x and y axes show the median logFC. Blue (co-enhance) represents targets with significant positive values on both axes (FDR < 0.05). Green (co-repress) represents targets with significant negative values on both axes. Orange (antagonize) represents targets with a significant positive score from one signal and a negative one from the other. Selected antagonized targets are labeled. b, Fractions of cytokine co-regulation type. For each cytokine pair and their targets, we counted the fractions of significant co-regulation in the three categories illustrated in a. c, Antagonistic relationships among cytokines on target genes. Each flat-end edge indicates that the first cytokine represses targets of the second one. The edge width is proportional to the number of antagonized targets. The node pie chart represents the in and out degrees. d, Antagonizing regulatory relationships between cytokines and anti-inflammatory signals. The heat map presents the number of target genes induced by a cytokine (row) but repressed by an anti-inflammatory signal (column). The similarities are shown with the hierarchical clustering trees using the Pearson correlation as the distance metric.
In 14% of the statistically significant cases, cytokines exhibited an antagonistic relationship, meaning that they had opposite signaling effects on downstream targets (Fig. 3b). For example, IL-4 and BMP6 downregulated many targets induced by IL-1β and TNF (Fig. 3a and Extended Data Fig. 3a). We also observed a similar relationship among four other cytokines: the IFN-γ target genes were antagonized by IL-10 and GCSF but enhanced by IL-27 (Extended Data Fig. 3b). Thus, our target co-regulation analysis identified four major anti-inflammatory regulators (IL-4, BMP6, IL-10 and GCSF), which antagonize chiefly pro-inflammatory molecules in two groups (Fig. 3c,d), referred to as the NF-κB and interferon transcriptional groups (Fig. 2b).
BMP6 antagonizes pro-inflammatory cytokine target genes
Previous work demonstrated that IL-4 could inhibit NF-κB transcription programs33, explaining the antagonistic relationship between IL-4 and cytokines in the NF-κB group. A previous study demonstrated that BMP6 could inhibit the CCL2 mRNA level induced by TNF34. However, to the best of our knowledge, no previous studies have reported BMP6 as an anti-inflammatory molecule that antagonizes many pro-inflammatory targets.
Our analysis indicated that BMP6 may antagonize the effect of IL-1β through downregulation of IL-1β-induced pro-inflammatory chemokines, with CXCL8 and CCL2 as the most significant targets (Fig. 3a). To validate our prediction, we first evaluated the intracellular protein levels of CXCL8 and CCL2 upon IL-1β and BMP6 treatments by flow cytometry in two human lung epithelial cell lines, A549 and NCI-H1299. Consistent with our data analysis, BMP6 treatment significantly inhibited the IL-1β induction of CXCL8 and CCL2 (Extended Data Fig. 4a–c). ELISA assays also indicated that levels of soluble CXCL8 and CCL2 were consistently lower in cells treated with BMP6 + IL-1β compared to IL-1β alone (Extended Data Fig. 4d; one-sided Wilcoxon signed-rank P value = 0.016).
CytoSig predicts signaling activities from expression data
Because the cytokine-response data in our collection reflect signaling relationships in human tissues (Fig. 1e), we created the CytoSig model to predict signaling activities using the transcriptome profile of an input sample. The output of CytoSig is different from standard cytokine assays such as ELISA, which measure cytokine levels instead of cytokine target activities. CytoSig utilized the ridge regression to search for features in an input transcriptome profile that can be explained by a cytokine’s influence on its target gene expression (Extended Data Fig. 5 and Methods).
As described in the introduction, redundancy and pleiotropy are major obstacles to modeling cytokine activity. To account for complications from signaling pleiotropy, our model only aims to predict each cytokine’s overall activity, instead of its effects on individual genes or pathways. We analyzed each cytokine’s composite signature, averaged across at least five independent experiments. Significant enrichment of the composite signature of a cytokine in the input sample’s transcriptome should indicate the presence of signaling events. To address signaling redundancy, we utilized a penalized linear model that avoids reporting a cytokine as active if other cytokines with similar composite signatures have influenced target gene expression to a greater extent. For any input profile, our model reports a significant score for a cytokine only if predicted activities were significantly higher than expected by chance (Methods).
Accuracy validation using cytokine-blocking clinical response
To test the model accuracy, we reasoned that the participant’s clinical response upon cytokine-blocking therapies should reflect authentic cytokine activities in human tissues. Therefore, we compared CytoSig predictions of cytokine activities with transcriptomic data before and after cytokine-blocking therapies in inflammatory diseases (Supplementary Table 3). For example, a microarray study measured the whole-blood transcriptome of individuals with arthritis at baseline and day 3 after anti-IL-1β canakinumab treatment and evaluated the therapy response at day 15 after therapy35. Upon IL-1β neutralizing therapy, the IL-1β activity reduction at day 3 predicted by CytoSig correlated significantly with the participant’s clinical response evaluated at day 15 (Fig. 4a). For another example, an IFN-α vaccine trial among patients with systemic lupus profiled both whole-blood transcriptomes and clinical response as the titer of IFN-α neutralizing antibodies in blood after immunization36. The IFN-I activity reduction predicted by CytoSig correlated significantly with the clinical response across patients (Fig. 4b).
a, IL-1β activities in blood predicted the anti-IL-1β therapy response in arthritis. Each dot represents a blood sample35. The x axis shows patients with similar therapy responses. The y axis shows the IL-1β activities predicted by CytoSig at an early time point, shown by violin plots smoothed by a kernel density estimator. Spearman correlation between clinical response in patient groups and the median IL-1β activity with a two-sided t-test P value is indicated. b, IFN-I activity in blood correlated with antibody titer upon IFN-α vaccine in patients with systemic lupus. Each dot represents a blood sample. The x axis presents the titer of IFN-α antibody after immunization36. The y axis presents the IFN-I differential activity predicted by CytoSig. Spearman correlation between x and y axes with a two-sided t-test P value is indicated (n = 36). c, Predicted activity change upon anti-cytokine therapies. Each dot represents an anti-cytokine therapy study in inflammatory diseases, with targets on the x axis. Gray labels represent mouse model studies for cytokines without clinical data. The y axis presents the average differential activity between posttreatment and pretreatment groups. The accuracy was computed as the fraction of cytokines with median activity reduction smaller than one, with P value from the two-sided Wilcoxon signed-rank test. d, TGF-β activity changes upon neutralizing antibody treatments. The first antibody inhibited all TGF-β isoforms (123; n = 7 mice) and the second antibody inhibited only TGF-β1 and TGF-β2 (12; n = 6 mice). The anti-TGF-β3 profile (123/12) was the differential profile between the pan-TGF-β- and the TGF-β1/2-specific groups. The TGF-β1 and TGF-β3 activities predicted by CytoSig are shown by bar plots, with two-sided P values from the permutation test with 10,000 randomizations. e, VEGF activities in pretreatment tumors predicted the anti-VEGF therapy response in sunitinib38 and bevacizumab clinical studies39. The Kaplan–Meier plot presents patient fractions (y axis) with survival length (x axis, progression-free or overall) among pretreatment tumors with high and low VEGFA activities predicted by CytoSig. The activity cutoff was selected through maximizing the difference between high and low groups. The P value was derived from the one-sided Wald test using continuous values without cutoffs.
Among all cytokine-blocking studies collected from GEO and AE databases, CytoSig predicted the activity reduction score to be at least negative one (one standard deviation below zero) for 85% of cytokines (Fig. 4c). The accuracy dropped to 0% when gene labels were permuted in the model. These results support the reliability of CytoSig on cytokine activity prediction in human tissues and demonstrate the clinical utility to guide therapy decisions.
Accuracy validation on TGF-β isoform-specific activities
CytoSig predicts different activities for cytokines from the same family sharing receptors, such as TGF-β1 and TGF-β3. The validation in the previous section established that CytoSig can perform with high accuracy on a broad set of cytokines. To validate the accuracy of CytoSig’s predictions of signaling activities among cytokine isoforms sharing the same receptors and similar downstream pathways, we performed in vivo experiments with the 4T1 breast cancer mouse model using neutralizing antibodies to TGF-β isoforms.
Specifically, we profiled the transcriptomes of mouse 4T1 tumors treated with neutralizing antibodies targeting all TGF-β isoforms and antibodies targeting only TGF-β1 and TGF-β2 (but not TGF-β3). The differential profile between pan-TGF-β and TGFβ1/2 antibodies can reflect the anti-TGF-β3 effects because the TGF-β3 isoform is the differential target between two antibodies. CytoSig predicted a significant reduction in TGF-β1 activity based on the differential transcriptomic profiles upon treatments for both anti-TGF-β antibodies, and a significant reduction of TGF-β3 activity only for the anti-TGF-β3 profile (Fig. 4d).
Accuracy validation in tumors and cancer therapy response
To further evaluate CytoSig model accuracy, we utilized the International Cancer Genome Consortium (ICGC) tumor cohort37, which has no overlap with the previous TCGA and GTEx data in model training. We assumed that tumors with ligand or receptor expression levels higher than one standard deviation above the average level in the entire dataset reflected positive activity for that cytokine. Under this assumption, we evaluated the accuracy of CytoSig on predicting samples with significant cytokine signaling activities. Based on the receiver operating characteristic (ROC) curve and area under the ROC curve (AUC), 35 of 43 cytokines had a performance significantly better than chance (Extended Data Fig. 6a,b). Therefore, the CytoSig model can predict target activities of most cytokines.
We also evaluated the CytoSig model in predicting the clinical outcome of anticancer therapies that inhibit cytokine signaling. Vascular endothelial growth factor (VEGF) blocking is a category of treatments inhibiting either VEGF ligands or VEGF receptors from promoting abnormal angiogenesis in tumors38,39. As the cancer driver, the pretreatment target pathway activity may predict targeted therapy efficacy and patient survival after treatment40. We found that high VEGF signaling activities predicted by CytoSig in pretreatment tumors, using data from two clinical studies38,39, were highly predictive of longer survival outcomes upon blocking the VEGF pathway through either ligand (bevacizumab) or receptors (sunitinib, inhibitor of multiple receptor tyrosine kinases, including VEGF receptors; Fig. 4e).
Immune checkpoint blockade is another treatment category whose responses depend on cytokine signaling by IFN-γ41. CD274, which encodes PDL1, is a target gene induced by IFN-γ signaling42; therefore, we evaluated the association between IFN-γ activity in pretreatment tumors and the anti-PDL1 therapy response, using data from an anti-PDL1 clinical trial in urothelial cancer43. IFN-γ activity predicted by CytoSig was highly predictive of overall survival outcome upon anti-PDL1 (Extended Data Fig. 6c). Moreover, for both anti-VEGF and anti-PDL1 clinical studies, the CytoSig predictions had better associations with the clinical outcome than other approaches, such as ligand or receptor expression and gene-set signatures (Extended Data Fig. 6d).
Accuracy validation in single-cell transcriptomic data
Encouraged by the reliable performance on bulk data, we further evaluated the capability of the CytoSig to predict signaling activities in single cells. The ideal evaluation standard for CytoSig predictions in single cells would be a method providing systematic measurements of both transcriptome and cytokine activities in each single cell. However, to our knowledge, no such method currently exists. To validate the accuracy in single-cell data, we used transcription factor (TF) activities as indicators of active cytokine signaling (Supplementary Table 4).
We computed TF activities for a single-cell transcriptomic profile using the RABIT framework, which leverages an extensive collection of chromatin immunoprecipitation and sequencing (ChIP–seq) profiles to predict TF activities through transcriptional patterns of TF target genes44. For example, using data from a COVID-19 single-cell study45, RABIT predicted that most CD8+ T cells have positive STAT1 TF activity, reflected as a higher expression level of STAT1 ChIP–seq target genes compared to other genes (Fig. 5a). A minor CD8+ T cell population showed negative STAT1 TF activity. Consistent with the dependence of interferons and IL-27 on STAT1 signaling (Supplementary Table 4), cells with positive TF activities had significantly higher signaling activities from the CytoSig model than cells with negative TF activities (Fig. 5a).
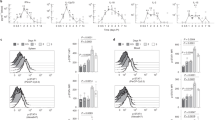
a, Single-cell signaling activities for STAT1-group cytokines among CD8+ T cells using data from a COVID-19 study45. For each single cell (dot), the x axis shows the STAT1 TF activities. The y axis presents the average signaling activities among interferons and IL-27. The color brightness indicates the local dot density. The P value was calculated using the two-sided Wilcoxon rank-sum test, comparing cytokine activities of single cells with positive (n = 1,323) and negative (n = 117) STAT1 activities. b, ROC curves for CD8+ T cell effector cytokines. The ROC curve presents false-positive rates against true-positive rates of TF activity prediction at different cytokine activity thresholds using the data in a. The diagonal line represents random expectation. c, Accuracy of single-cell cytokine activities prediction. The AUC for each cell type was computed for each TF and its cytokine families, using the COVID-19 single-cell dataset analyzed in a and b. Each dot represents a cell type (n = 12 cell types per box plot). The thick line represents the median value. The bottom and top of the boxes are the 25th and 75th percentiles (interquartile range). The whiskers encompass 1.5 times the interquartile range. d, Accuracy of single-cell cytokine activities prediction among many single-cell datasets. Each dot presents the median AUC among all cell types (y axis) in a single-cell dataset. The results are shown for all cytokine and TF combinations using box plots in c (n = 10 cytokine–TF combinations per box plot). NK, natural killer; Treg, regulatory T.
We utilized a ROC curve to measure the ability of CytoSig to predict TF activities based on the predicted cytokine activity. We found that the activity of CD8+ T cell effector cytokines, including interferons and TNF, all predicted downstream TF activities better than would be expected at random (Fig. 5b). Using the AUC values, we next evaluated the predictive performance of cytokine activities on their downstream TF activity for all cell types in the COVID-19 single-cell study. The AUC metrics were consistently higher than expected by chance for 10 of 11 pairs of cytokines and downstream TFs (Fig. 5c and Supplementary Table 4). We observed similar high performance in another cancer study (Extended Data Fig. 7a). We performed such evaluation on 18 single-cell datasets and found that AUC metrics were consistently higher than would be expected at random (Fig. 5d).
CytoSig identifies signaling markers of severe COVID-19
The global spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is an urgent health crisis. The symptoms of COVID-19 range from mild fever, cough and difficulty breathing to respiratory failure and death7. While more severe outcomes have been associated with an exaggerated immune response, referred to as the cytokine storm, the immune-response mechanisms underlying the dramatic differences in disease severity remain unclear.
We applied CytoSig to analyze single-cell RNA-seq data from bronchoalveolar lavage fluid45 and peripheral blood46 of patients with COVID-19 (Fig. 6a). These datasets were used earlier in this paper to establish reliable prediction accuracy in a single-cell RNA-seq data (Fig. 5c,d and Extended Data Fig. 7b). Many cytokine signaling activities are significantly associated with the severity differences in COVID-19 symptoms. For example, among macrophages from lavage fluid, IL-10 activity is significantly higher in patients with severe disease than in those with mild disease and in healthy controls (Fig. 6b,c and Supplementary Table 5). In contrast, among CD8+ T cells from blood, the IFN-I activity is highest in patients with mild illness compared to those with severe disease or healthy controls (Fig. 6d,e). This is consistent with previous studies reporting a lack of IFN-I response among both patients47 and cultured cells upon SARS-CoV-2 infection48.
a, Sample source sites. We analyzed single-cell RNA-seq datasets from the bronchoalveolar lavage and peripheral blood samples from patients with COVID-19. b, IL-10 activities in macrophages from lavage fluid. Each dot presents a single cell from the COVID-19 study with t-distributed stochastic neighbor embedding (t-SNE) coordinates computed per the original publication45. The color represents the predicted IL-10 activity. Circles highlight the cluster of macrophages. c, Enrichment of IL-10 activities in macrophages from lavage fluid from patients with severe illness. The violin plots present IL-10 activity distributions in different patient groups, smoothed by a kernel density estimator. The color legend is per b. The two-sided Wilcoxon rank-sum P value was computed to compare activities of severe (n = 6) and mild (n = 3) patient groups. d, IFN-I activities in CD8+ T cells from blood. The IFN-I activities in CD8+ T cells (circles) are shown as per b, with uniform manifold approximation and projection (UMAP) coordinates from the original publication46. e, Depletion of IFN-I activities in peripheral blood CD8+ T cells from patients with severe disease. The two-sided Wilcoxon rank-sum P value was computed to compare activities of severe (n = 14) and mild (n = 13) patient groups within each cell type. f, Differential signaling activities in macrophages from lavage fluid. The heat map presents cytokines whose predicted signaling activities were significantly different between individuals with severe disease, those with mild disease and healthy individuals. g, Summary of signaling activities in COVID-19. The heat map includes cell types with significant differences in signaling activities among individuals with severe disease, those with mild disease and healthy individuals. Each cell shows the median value across all individuals in each group. Only cytokines with at least three significant values in at least three cell types were included.
Analysis of differential activity among individuals with severe disease, those with mild disease and healthy individuals revealed a few cytokines as signaling markers of COVID-19 symptom severity. Among patients with severe COVID-19, we found elevated IL-1B activities in macrophages, the most populous cell type from bronchoalveolar lavage fluid45 (Fig. 6f). This result of IL-1B activation in macrophages is consistent with previous reports7. Meanwhile, IL-10 had high activity levels among patients with severe disease in lavage and blood myeloid cells, including macrophages, monocytes and neutrophils (Fig. 6g and Extended Data Fig. 8), a result also consistent with published studies49,50. IL-10 is known to inhibit antigen presentation by dendritic cells to cytotoxic T cells, thus impairing T cell-mediated antiviral immunity51. IL-10 can also suppress macrophage activation for fighting against intracellular pathogens52. Thus, in severe cases of COVID-19, the cytokine environment may compromise the antiviral immune response while triggering pathological inflammation. We observed that monocytes and macrophages have high expression levels of IL-10 and IL-1B, thus potentially serving as cytokine-producing cells (Extended Data Fig. 9).
CXCL8 as a candidate therapeutic target for severe COVID-19
Direct IL-1B or IL-10 blockade may compromise the antiviral response53 or fail to alleviate inflammation54. Therefore, inhibition of downstream targets could serve as alternative approaches. With this in mind, we analyzed the downstream targets induced by IL-1B or IL-10. We found that IL-1B target CXCL8 had higher expression levels in macrophages from lavage samples of patients with severe COVID-19 (Extended Data Fig. 10). Meanwhile, in blood neutrophils, IL-10 target CXCL1 had higher expression levels among individuals with severe illness than in those with mild illness and in healthy controls (Extended Data Fig. 10).
CXCL1 and CXCL8 all bind to the CXCR2 receptor and serve as primary chemokines in neutrophil recruitment. A high neutrophil-to-lymphocyte ratio in peripheral blood indicates severe disease and organ failure55. Aberrant formation of neutrophil extracellular traps may contribute to severe damage to the lung parenchyma in COVID-19 (ref. 56). A phase I clinical trial evaluated the CXCL8 blocking antibody in treating solid tumors and has not observed any dose-limiting toxicities57, which indicates the potential of therapy repurposing.
Discussion
We have introduced CytoSig, a data-driven platform to model cytokine activities. CytoSig complements existing cytokine release assays because it can predict cytokine target activities from bulk transcriptomic data available from many large-scale cohorts and single-cell RNA-seq data that provides resolution down to individual cells. The acquisition of both types of data is now routine, making CytoSig useful to a broad spectrum of research questions.
CytoSig offers particular advantages in analyzing single-cell data because it is not affected by the absence of cytokine-producing cells or zero read counts for ligand or receptor genes. This advantage is especially important because current single-cell technologies have difficulty capturing some cell types, such as neutrophils58. Many studies also sort cells using markers such as CD45, which may exclude cytokine-producing cell populations. Moreover, the dropout events, reflected as zero read counts, on transiently expressed cytokine genes further complicate analysis. The CytoSig model uses a reliable alternative strategy, analyzing receiver cells’ transcriptional patterns across many cytokine target genes.
A limitation of CytoSig is the ascertainment bias of public datasets, which leads to many experiments on a few cytokines, cell models or experimental conditions, and a lack of data on others. There are currently 67 human cytokines, 42 chemokines and 133 growth factors annotated in the literature28. However, our collection from public databases captures high-quality profiles for 43 of the 242 documented molecules (17.8%) due to the lack of data available for most signaling molecules. Also, most datasets were generated through a few models, such as monocytes or fibroblasts, without sufficient coverage on diverse cell lineages. Such a gap indicates a need for attention on a broad range of cytokines and cell models beyond a few deeply studied molecules and systems.
Despite these limitations, the CytoSig platform provides biologists and clinicians with a powerful resource to study signaling activities in laboratory or clinical samples. Furthermore, independent of CytoSig, the FDC is a general resource for data scientists to accelerate data curation projects. Using the FDC, our plans to continuously integrate new datasets will provide the community with an ever-growing repository for generating new biological insights.
Methods
The Framework for Data Curation from public databases
The FDC aims to automate the data curation process as much as possible with two components: (1) semiautomated metadata annotation; (2) automatic gene expression matrix extraction. Databases processed by the FDC include AE15, GEO14, ENA13 and SRA12. The FDC server is built using Python 3 and Dango 3 frameworks with MySQL 8 as the database backend. The natural language processing functions are created in the web browser frontend using the jQuery 3 JavaScript library.
The first FDC component for metadata annotation utilizes a three-stage approach. In the first stage, the users should query the GEO and AE databases with keywords related to their biological topic. The SRA and ENA metadata are available through the GEO and AE, respectively. The database query will generate a list of candidate datasets. After uploading the candidate list to the FDC, the users can define pattern-matching rules, implemented as regular expressions59, to narrow down query results. In the second stage, users should browse study summaries and determine which datasets are relevant to their study topic. To accelerate the process, users could define a set of highlighting rules, implemented as regular expressions59, so that curators only need to focus on the most relevant texts.
In the third stage, users will extract metadata fields for each experimental profile. The FDC aims to reduce human manual edits as much as possible, with automatic rules and text transformation functions defined by users. The FDC will automatically parse each sample’s study design information and summarize all potentially relevant fields in a candidate table. The users can define a set of automatic mapping rules to convert aliases, such as biological molecule and cell model names, to their standard names. The FDC also provides automated functions to extract and transform text information from candidate metadata columns. Based on these functions, curators will standardize metadata columns into controlled vocabularies.
The second FDC component can automatically extract MicroArray and RNA-seq public databases. For Affymetrix MicroArray data on the AE and GEO, we generated expression matrices from CEL files through the R Oligo package60. For other MicroArray platforms, we downloaded the processed data through the R GEOquery61 and Python Orange62 packages for GEO and AE, respectively. For RNA-seq data from the ENA and SRA databases, we downloaded the fragments per kilobase of transcript per million mapped reads (FPKM) data through the RNASeq-er Application Programming Interface63. In total, our framework extracted 27,181 independent human transcriptomics datasets deposited before 02 February 2020. Many datasets from public repositories are not gene expression studies, thus cannot be automatically analyzed by the second FDC component. However, the first FDC component can still assist the metadata annotation for non-transcriptomic data.
Besides the two primary components introduced above, the FDC also provides other assistant modules, such as curator management for a project and result proofreading panels if project managers want to review the annotations from curators. With the standardized metadata matrix from the first component and gene expression matrix from the second component, users can perform algorithmic data analysis for their biological topics.
Collection of human cytokine-response data based on the FDC
In the first step, we queried names and aliases of human cytokines, chemokines, growth factors28 and a few immunosuppressive signals in the tumor microenvironment64 through the query interface of GEO and AE. The SRA and ENA metadata are available through the GEO and AE, respectively. Our query returned 9,271 candidate series, 5,186 of which had processed data matrices from the FDC. The other 4,085 datasets did not have FDC-processed data for several reasons. Some studies using NanoString platforms only focus on hundreds of genes instead of genome wide. Some MicroArray or RNA-seq studies may have corrupted raw data, leading to FDC extraction failures. We also excluded all micro-RNA and noncoding RNA studies.
In the second step, we recruited Ph.D. scientists with immunology training for data annotations based on the FDC. Curators focused on data curation for several months. A second curator proofread all annotations of the first curator and corrected errors. Most studies among the 5,186 candidates only mentioned cytokines in their description but did not study the signaling response. The curators read descriptions of 5,186 experiments and identified 962 of them as cytokine-response studies, including 20,591 nonredundant samples (Supplementary Table 1). Then, curators read descriptions of 20,591 samples and labeled cytokine treatment, cell model, dose and duration, using the semiautomated functions on the FDC. We established a set of control vocabularies about signal names, cell models, concentrations and duration units.
Together with data matrices extracted and expert annotations, we generated differential gene expression profiles, defined as the logFC between treatment and control conditions. We only kept experiments with biological replicates and acquired 2,056 logFC vectors after merging biological replicates. Meanwhile, we merged IFN-α and IFN-β as IFN-I, representing type I interferons, due to the high Pearson correlation of 0.698 between their composite profiles. We also combined IL-36A and IL-36B as IL-36 due to the high correlation of 0.938 between their composite profiles.
Data quality control
To test the human physiological relevance of data collection, we defined a quality-control metric as the Pearson correlation between expression levels of cytokine target genes and the ligands or receptors in independent human tissue data. We used TCGA18 and GTEx19 datasets. Each TCGA and GTEx sample measures a bulk tissue’s average expression that contains both producer and receiver cells for cytokines.
To measure the overall expression of target genes, we performed a linear regression for each pair of cytokine-response and tissue expression profiles as ‘tissue expression = A × cytokine profile + B,’ and computed the cytokine profile’s target score as A/standard error(A) using the ordinary least-squares method65. The target score represents the enrichment of a response signature in the tissue expression profile. We then analyzed the Pearson correlation between the target score and ligand or receptor expression across tissue samples (Fig. 1e).
TCGA has 33 tumor cohorts, and GTEx has 27 tissue cohorts. For each cytokine profile, we utilized the one-sided Wilcoxon test to evaluate whether the correlations with the ligand or receptor were higher than zero across both TCGA and GTEx cohorts (FDR < 0.05 with Benjamini–Hochberg correction). We only included 1,307 profiles that passed the threshold in further analysis.
Target cooperation and antagonization analysis between cytokine pairs
We computed FDRs between each cytokine pair to test the statistical significance of co-regulating target genes. For each target gene C, there are three types of co-regulations from a cytokine pair:
-
1.
Cytokine A and B both induce target gene C.
-
2.
Cytokine A and B both suppress target gene C.
-
3.
Cytokine A (or B) induces target C, but the other cytokine B (or A) represses target C.
First, we defined two logFC thresholds of cytokine A and B for the FDR computation. For type 1 co-regulation (co-enhance), we computed the \(FDR\,(thres_A,\,thres_B)\) as \(Random\,count\,(logFC_A \ge thres_A,\,logFC_B \ge thres_B)\)\(/gene\,count\,(logFC_A \ge thres_A,\,logFC_B \ge thres_B)\). The gene count derives directly from the data. The random count is equal to \(N \times probability\,(logFC_A \ge thres_A) \times probability\,(logFC_B \ge thres_B)\). N represents the total number of genes. We computed both probabilities from the logFC rank of each gene. In summary, the FDR computations are as follows:
Similarly, for type 2 co-regulation (co-repress), we computed the FDR as:
For type 3 co-regulation (antagonize), we computed the FDR as:
After computing the FDR at each threshold combination (thres_A and thres_B), we adjusted FDRs into monotonically decreasing values with respect to increasing threshold values, following the q-value procedure66. Finally, for the triplet of each cytokine pair and target gene, its statistical significance is the FDR (logFC_A and logFC_B) under each co-regulation category.
Penalized linear model to predict cytokine target activities
The CytoSig linear model is programmed through a combination of Python 3 and GNU Compiler Collection 4C++. We only included 43 cytokines with at least five high-quality experiments (‘Data quality control’). We utilized a linear model to identify each signaling molecule’s signature patterns in an input sample’s expression profile. Composite profiles of cytokine response were the explanatory variables, and an input sample’s transcriptomic profile was the response variable. The regression coefficients represent cytokine target activities. The linear regression with all cytokine composite profiles as explanatory variables will reduce a cytokine’s coefficient if other cytokines with similar response profiles have more extensive impacts on the sample’s transcriptomic pattern67.
The expression values, from either RNA-seq or MicroArray, should be transformed by log2(x + 1). We also recommend quantile normalization across conditions. Some software packages, such as RMA or DESeq, will automatically include all normalizations. We recommend input differential profiles between the two conditions. If data are from a sample collection without pairs, the value of each gene across all samples should be mean centralized.
Many cytokine profiles were highly similar (Fig. 1f); such signature collinearity will create large result variance in a regular linear regression65. Therefore, we used the penalized ridge regression, which trades off the result bias to reduce the variance. The vector y is the input sample’s expression profile. The matrix X contains composite profiles of 43 cytokines. The parameter λ is the penalty. The ridge regression aims to minimize the objective function \((y - X\beta )^T(y - X\beta ) + \lambda \times \beta ^T\beta\). The coefficient β represents signaling activities.
To optimize parameters, we evaluated two types of model performance:
-
1.
Prediction performance. This evaluates how the fitted model of cytokine activities predicts a sample’s gene expression profile. We use the fivefold cross-validation R2 ratio as the prediction performance metrics.
-
2.
Inference performance. This evaluates whether coefficients on cytokine covariates of the fitted model represent the actual cytokine activities in a sample. We used the correlation between model coefficients and the ligand or receptor expression across samples as the inference performance metrics for each cytokine.
Typically, the training of ridge regression models only evaluates the prediction performance through cross-validation to determine the optimal penalization factor and coefficients65. However, we also evaluated the inference performance because the goal of the CytoSig model is to infer cytokine signaling activities. The collinearity among cytokine-response profiles may not affect the prediction performance but will induce significant variance on model coefficients65, thus undermining the inference performance. The penalty factor in the ridge model will reduce the model variance at the cost of lowering prediction performance. Thus, we aimed to find a penalty factor as a trade-off between two performance aspects.
On average, the cross-validation R2 metric reaches its maximal point at a low penalty factor and deteriorates while the penalty factor is increasing (Extended Data Fig. 5a,b). In contrast, the inference performance, measured as correlation values, monotonically increases with increasing penalties (Extended Data Fig. 5c,d). Therefore, we selected a value of 10,000, which is the minimal lambda to achieve 80% best-inference performance and 70% best-prediction performance. Such a penalty will control both result variance and bias in the ridge regression.
We also evaluated XG Boost, a popular machine learning algorithm68 (Extended Data Fig. 5). The XG boost with tree learners outperformed ridge regression in prediction but does not provide any coefficients on cytokine covariates for the inference purpose due to the tree structure of learners. The prediction performance of XG boost with linear learners quickly deteriorates to zero with increasing penalties although it has a high inference performance. Ridge regression is the only method with reasonable performance in both prediction and inference metrics.
We utilized a permutation test to estimate ridge coefficients’ standard errors after shuffling gene identities 1,000 times. The z-scores (coefficient − random_average_coefficient)/standard_deviation on each cytokine represents its target activity.
T cell activation and TGF-β1 treatment assay
Human primary T cells were sourced from Hong Kong Red Cross Transfusion Service. Peripheral blood mononuclear cells were isolated from healthy donors using the Ficoll Paque Plus (GE healthcare, 17-1440-03) via density gradient centrifugation. CD8+ T cells were purified from fresh peripheral blood mononuclear cells by magnetic negative selection using the human CD8+ T cell isolation kit (Miltenyi Biotec, 130-096-495). Isolated cells were stimulated with the human T cell TransAct (Miltenyi Biotec, 130-111-160) in the presence or absence of human recombinant TGF-β1 (R&D systems, 240-B-002) at 5 ng ml−1 for 72 h. Cells were cultured in MACS GMP medium, which is TexMACS GMP medium (Miltenyi Biotec, 170-076-309) supplemented with 10% inactivated fetal bovine serum (FBS; Gibco, 10082147), 50 µM 2-mercaptoethanol (Gibco, 21985023), 10 mM N-acetyl-l-cysteine and 1% penicillin–streptomycin (P/S; Gibco, 15140122) at 1 × 106 cells per ml.
Mouse CD8+ T cells were isolated from splenocytes of one 8-week-old male C57BL/6J mouse using the CD8a T cell isolation kit (Miltenyi Biotec, 130-104-075) by magnetic negative selection. Isolated CD8+ T cells were stimulated with plate-bound anti-mouse CD3 (BioLegend,100202, clone 17A2) at 5 µg ml−1 (1:100 dilution) and soluble anti-mouse CD28 (BioLegend, 102102, clone 37.51) at 2 µg ml−1 (1:250 dilution) in the presence or absence of human recombinant TGF-β1 (R&D systems, 240-B-002) at 5 ng ml−1 for 72 h. Cells were cultured in complete RPMI 1640 medium, which is RPMI 1640 Medium (Gibco, 11875119) supplemented with 10% inactivated FBS (Gibco, 10082147), 20 mM HEPES (Gibo, 15630080), 1 mM sodium pyruvate (Gibco, 11360070), 50 µM 2-mercaptoethanol (Gibco, 21985023), 2 mM l-glutamine (Gibco, 25030024) and 1% P/S (Gibco, 15140122) at 1 × 106 cells per ml.
Human inadequate whole blood was collected following informed consent and protocols were approved by the ethics committee at the University of Hong Kong and the Hong Kong Red Cross Blood Transfusion Service. Animal experiments were approved by the committee of the Use of Live Animals in Teaching and Research at the University of Hong Kong and performed strictly according to the animal protocol 5310-20. C57BL/6J mice were purchased from the Laboratory Animal Unit of the University of Hong Kong.
BMP6 and IL-1β in vitro treatment combinations
NCI-H1299 (CRL-5803) and A549 (CCL-185) cells were purchased from American Type Culture Collection (ATCC). NCI-H1299 cells are cultured in high-glucose DMEM medium (Gibco) supplemented with 10% FBS (Gibco BRL) and 100 IU per ml P/S. Human A549 cells were cultured in F12-K medium (ATCC, 30-2004) supplemented with 10% FBS and 100 IU per ml P/S.
NCI-H1299 and A549 cells were seeded in a six-well plate at the density of 2 × 105 cells per well. On the next day, cells were treated with human recombinant IL-1β (R&D systems, 201-LB-005, 10 ng ml−1) alone or in combination with human recombinant BMP6 (R&D systems, 507-BP-020, 10 ng ml−1) for 12 h. In an alternative sequential treatment schedule, cells were pretreated with IL-1β first for 12 h, then BMP6 or media control for another 12 h. Reconstitution buffers of the IL-1β (PBS containing 0.1% BSA) and BMP6 (4 mM HCl containing 0.1% BSA) were used as negative controls.
Flow cytometry
For the evaluation of intracellular markers on A549 and H1299 cells, the following antibodies were used at the indicated dilutions:
PE anti-human PRF1 (BioLegend, 353303, clone B-D48, 1:50 dilution),
PE anti-mouse PRF1 (BioLegend, 154305, clone S16009A, 1:50 dilution),
PE anti-human GZMA (BioLegend, 507206, clone CB9, 1:50 dilution),
PE anti-mouse GZMA (BioLegend, 149703, clone 3G8.5, 1:100 dilution),
FITC anti-human/mouse GZMB (BioLegend, 515403, clone GB11, 1:50 dilution),
APC anti-human/mouse MCP-1 (CCL2, BioLegend, 505909, clone 2H5, 1:200 dilution),
FITC anti-human CXCL8 (BioLegend, 511406, clone E8N1, 1:50 dilution).
Cells were fixed before permeabilization according to the manufacturer’s instructions of wash buffer (BioLegend, 421002), and followed by intracellular staining with the antibodies. Flow cytometry was performed on an ACEA NovoCyte Quanteon and raw data were analyzed using FlowJo (Version 10.7).
To determine the gating threshold to detect marker-positive cells, we used the forward scatter height (FSC-H) and side scatter height (SSC-H) for dead cell and debris removal. FSC-H/width and SSC-H/width were used to select single cells. We included unstained cells to define the threshold that separates positive populations from negative control cells (Extended Data Figs. 2d and 4c).
CXCL8 and CCL2 detection by enzyme-linked immunosorbent assay
A549 cells were seeded in a six-well plate at a density of 2 × 105 cells per well. On the next day, cells were treated with human recombinant IL-1β (R&D systems, 201-LB-005, 10 ng ml−1), human recombinant BMP6 (R&D systems, 507-BP-020, 10 ng ml−1) and combinations of IL-1β and BMP6 for 24 h. Reconstitution buffers of IL-1β (PBS containing 0.1% BSA) and BMP6 (4 mM HCl containing 0.1% BSA) were used as negative controls.
The amount of released CCL2 and CXCL8 from tumor cells in the supernatants was measured by ELISA assay using human CCL2 DuoSet ELISA kit (R&D systems, DY279) and human CXCL8 DuoSet ELISA kit (R&D systems, DY208). Optical density value was determined using a microplate reader (TECAN, Infinite 200) at a wavelength of 450 nm with the correction wavelength set at 570 nm.
Upon treatment combinations of IL-1β and BMP6 after 24 h, supernatants from different conditions were 200× diluted and measured. The experiment was repeated independently in three batches. In each batch, a standard curve was created to measure the relationship between fluorescence values and seven 2× concentration dilutions from 2,000 pg ml−1 and 1,000 pg ml−1 for CXCL8 and CCL2, respectively. The In(concentration + 1) and fluorescence values followed a linear relationship. We fitted a linear regression model to convert the fluorescence measurements to concentrations.
Anti-TGF-β1 animal studies
XOMA068 (pan-TGF-β1, TGF-β2, TGF-β3), XOMA089 (TGF-β1, TGF-β2 selective) and anti-KLH (control) antibodies, supplied by XOMA, were all fully human IgG2(κ) antibodies generated by phage display and affinity maturation in our previous study69. Briefly, fully human antibody phage display libraries were used to discover a number of antibodies that bind and neutralize various combinations of TGF-β1, TGF-β2 or TGF-β3. The primary panning did not yield any uniformly potent pan-isoform neutralizing antibodies; therefore, an antibody that displayed potent TGF-β1 and TGF-β2 inhibition but more modest affinity versus TGF-β3, was affinity matured by shuffling with a light chain sub-library and further screening. This process yielded the high-affinity pan-isoform neutralizing clone. Antibodies were diluted in 10 mM histidine and 142 mM l-arginine (pH 6.0) buffer ‘vehicle’ for in vivo studies.
Animal studies were conducted under protocol LC-070, approved by the Animal Care and Use Committee of the NCI. The animals were on a 12 h:12 h light:dark cycle. The ambient temperature was 72 ± 2 °F, and the humidity was kept between 30–70%. Around 40,000 4T1 mouse mammary tumor cells were surgically implanted into four mammary fat pads of 8-week-old female BALB/c mice. From day one after surgery, mice were treated with TGF-β antibodies at 5 mg per kg body weight intraperitoneally three times per week for 2 weeks. Tumors were surgically resected on day 13 when they reached 0.8–1 cm in diameter and were snap frozen for molecular analysis. The NCI Ethics Committee requires that animals must be euthanized at the time of observation if the tumor size is approaching 20 mm, in any dimension. None of the tumors in our experiment exceeded this limitation.
RNA was isolated from tumor samples using the RNeasy method (Qiagen) according to the manufacturer’s instructions following tissue lysis with a Precellys 24 Homogenizer (Bertin Instruments). Tumor RNA that passed quality control (RNA Integrity Number > 7) was sequenced on HighSeq 2500 using Illumina TruSeq v4 chemistry, generating 50–100 million pass-filtered reads per sample. There were six mice in the XOMA089 group, and seven mice in the XOMA068 group and anti-KLH group. No data points were excluded from the analysis.
Identification of signaling signatures in COVID-19 severe symptoms
For each single-cell dataset, we computed the cytokine activities for individual cells using the CytoSig model and got the mean value for each cell type in each patient. Then, for each cell type, we compared activities between different patient groups using the two-sided Wilcoxon rank-sum test and converted the P values to FDRs by the Benjamini–Hochberg correction. FDR < 0.05 is the threshold for the result significance.
For the COVID-19 study on bronchoalveolar lavage45 and peripheral blood46 samples, we performed comparisons between individuals with severe and mild disease, and between individuals with disease (severe and mild) and healthy individuals. Our analysis only reported results identified in both comparisons with an FDR < 0.05. We made an exception for the analysis of neutrophils; among neutrophils from peripheral blood, a few cytokines’ signaling differences between patients with severe and mild disease achieved a statistical significance of FDR = 0.051. We believe these results are still significant, and thus have reported them in our analysis.
We used the original coordinates of two-dimensional embedding from each publication (Fig. 6b,d). The bronchoalveolar lavage study45 utilized t-SNE that projects the single-cell RNA-seq profiles in two dimensions with distances between dots representing the profile similarities. The peripheral blood study46 utilized UMAP, a dimensionality reduction approach.
Statistics and reproducibility
All comparisons between two groups used the two-sided Wilcoxon rank-sum test, a non-parametric test without any assumptions on the data distribution. Similarly, all comparisons between group values and zero used the non-parametric Wilcoxon signed-rank test. No data were excluded from any analyses.
No statistical method was used to predetermine sample size. Instead, we selected a fixed sample size in the following experiments. In the in vitro validation of TGF-β1’s inhibitory role (Fig. 2e) and BMP6’s anti-inflammatory role (Extended Data Fig. 4), we used a sample size of three, the minimum number to achieve statistical significance of P value ≤ 0.05 in the two-sided Wilcoxon rank-sum test. All cell culture replicates lead to reproducible successful results (Fig. 2e and Extended Data Fig. 4b,d). In the TGF-β blocking in vivo experiment (Fig. 4d), we used a minimal mouse number of six, suggested by a previous study to detect differential expression events through RNA-seq70. Our recent study demonstrated that four tumors (smaller than our sample size of six) for each condition would be sufficient to detect differentially expressed genes between conditions71.
Mouse identities were randomized before in vivo experiments. Randomizations were not performed for in vitro cell cultures because all conditions were derived from a homogeneous cell line population. Blinding was not performed in our experiments because the robust phenotype of our results is based on strictly objective measurements by equipment instead of any human estimations. The outcome assessments included flow cytometry (Fig. 2e and Extended Data Fig. 4a–c) ELISA assay plate reader (Extended Data Fig. 4d) and RNA-seq (Fig. 4d). None of these measurements involved human subjective perception.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The processed data of cytokine treatment response are available at https://cytosig.ccr.cancer.gov/, visible after user registration and login. The RNA-seq data of TGF-β neutralizing antibody treatment is available at NCBI GEO under accession GSE174686.
The TCGA data are available at https://gdc.cancer.gov/ and were downloaded on 28 January 2020.
The ICGC data are available at https://dcc.icgc.org/ and were downloaded on 09 April 2020.
The GTEx data are available at https://gtexportal.org/home/datasets/ and were downloaded on 26 October 2019.
The gene expression datasets of human inflammatory disease (Fig. 4a–c) are available under the GEO accession codes listed in Supplementary Table 3. Other individual datasets analyzed are available under the database accession codes listed in Supplementary Table 6. Source data are provided with this paper.
Code availability
The interactive analysis modules of CytoSig are available at https://cytosig.ccr.cancer.gov/. The source code of CytoSig is available at https://github.com/data2intelligence/CytoSig/. Both applications are available under the NCI CytoSig software use agreement. The FDC server is available at https://curate.ccr.cancer.gov/ under the NCI FDC software use agreement.
Besides the software released above, we also provide source codes for essential analysis steps. The source code for automatic processing of treatment response data from the FDC server is available at https://github.com/data2intelligence/FDC_treatment_profile/. The source code for quality control of cytokine-response data when building the CytoSig model is available at https://github.com/data2intelligence/CytoSig_data_filter/.
References
Lin, J.-X. & Leonard, W. J. Fine-tuning cytokine signals. Annu. Rev. Immunol. 37, 295–324 (2019).
Zhang, Y., Guan, X.-Y. & Jiang, P. Cytokine and chemokine signals of T cell exclusion in tumors. Front. Immunol. 11, 594609 (2020).
Ozaki, K. & Leonard, W. J. Cytokine and cytokine receptor pleiotropy and redundancy. J. Biol. Chem. 277, 29355–29358 (2002).
Stenken, J. A. & Poschenrieder, A. J. Bioanalytical chemistry of cytokines—a review. Anal. Chim. Acta 853, 95–115 (2015).
Rusinova, I. et al. Interferome v2.0: an updated database of annotated interferon-regulated genes. Nucleic Acids Res. 41, D1040–D1046 (2013).
Subramanian, A. et al. Gene-set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity 52, 910–941 (2020).
Long, Q.-X. et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 26, 1200–1204 (2020).
Rotz, S. J. et al. Severe cytokine release syndrome in a patient receiving PD-1-directed therapy. Pediatr. Blood Cancer 64, e26642 (2017).
Yildizhan, E. & Kaynar, L. Cytokine release syndrome. J. Oncol. Sci. 4, 134–141 (2018).
Shimabukuro-Vornhagen, A. et al. Cytokine release syndrome. J. Immunother. Cancer 6, 56 (2018).
Leinonen, R., Sugawara, H. & Shumway, M., International Nucleotide Sequence Database Collaboration. The Sequence Read Archive. Nucleic Acids Res. 39, D19–D21 (2011).
Amid, C. et al. The European Nucleotide Archive in 2019. Nucleic Acids Res. 48, D70–D76 (2020).
Barrett, T. et al. NCBI GEO: archive for functional genomics datasets–update. Nucleic Acids Res. 41, D991–D995 (2013).
Parkinson, H. et al. ArrayExpress—a public database of microarray experiments and gene expression profiles. Nucleic Acids Res. 35, D747–D750 (2007).
Heine, A. et al. Generation and functional characterization of MDSC-like cells. Oncoimmunology 6, e1295203 (2017).
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Weinstein, J. N. et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Bender, H. et al. Interleukin-27 displays interferon-gamma-like functions in human hepatoma cells and hepatocytes. Hepatology 50, 585–591 (2009).
Kalliolias, G. D. & Ivashkiv, L. B. IL-27 activates human monocytes via STAT1 and suppresses IL-10 production but the inflammatory functions of IL-27 are abrogated by TLRs and p38. J. Immunol. 180, 6325–6333 (2008).
Elgueta, R. et al. Molecular mechanism and function of CD40/CD40L engagement in the immune system. Immunol. Rev. 229, 152–172 (2009).
Hostager, B. S. & Bishop, G. A. CD40-mediated activation of the NF-κB2 pathway. Front. Immunol. 4, 376 (2013).
Lawrence, T. The nuclear factor NF-κB pathway in inflammation. Cold Spring Harb. Perspect. Biol. 1, a001651 (2009).
Tokunaga, R. et al. CXCL9, CXCL10, CXCL11/CXCR3 axis for immune activation—a target for novel cancer therapy. Cancer Treat. Rev. 63, 40–47 (2018).
Mal, X. & Trinchieri, G. Regulation of interleukin-12 production in antigen-presenting cells. Adv. Immunol. https://doi.org/10.1016/s0065-2776(01)79002-5 (2001).
Hart, P. H. et al. Potential antiinflammatory effects of interleukin 4: suppression of human monocyte tumor necrosis factor alpha, interleukin 1 and prostaglandin E2. Proc. Natl Acad. Sci. USA 86, 3803–3807 (1989).
Murphy, K. & Weaver, C. Janeway’s Immunobiology (Garland Science, 2016).
Katoh & Katoh Integrative genomic analyses of CXCR4: transcriptional regulation of CXCR4 based on TGF-β, nodal, activin signaling and POU5F1, FOXA2, FOXC2, FOXH1, SOX17 and GFI1 transcription factors. Int. J. Oncol. 36, 415–420 (2009).
Nagase, H. et al. Cytokine-mediated regulation of CXCR4 expression in human neutrophils. J. Leukoc. Biol. 71, 711–717 (2002).
Gour, N. & Wills-Karp, M. IL-4 and IL-13 signaling in allergic airway disease. Cytokine 75, 68–78 (2015).
Thomas, D. A. & Massagué, J. TGF-β directly targets cytotoxic T cell functions during tumor evasion of immune surveillance. Cancer Cell 8, 369–380 (2005).
Ohmori, Y. & Hamilton, T. A. Interleukin-4/STAT6 represses STAT1 and NF-kappa B-dependent transcription through distinct mechanisms. J. Biol. Chem. 275, 38095–38103 (2000).
Varas, A. et al. Blockade of bone morphogenetic protein signaling potentiates the pro-inflammatory phenotype induced by interleukin-17 and tumor necrosis factor-α combination in rheumatoid synoviocytes. Arthritis Res. Ther. 17, 192 (2015).
Brachat, A. H. et al. Early changes in gene expression and inflammatory proteins in systemic juvenile idiopathic arthritis patients on canakinumab therapy. Arthritis Res. Ther. 19, 13 (2017).
Ducreux, J. et al. Interferon α kinoid induces neutralizing anti-interferon-α antibodies that decrease the expression of interferon-induced and B cell activation associated transcripts: analysis of extended follow-up data from the interferon-α kinoid phase I/II study. Rheumatology 55, 1901–1905 (2016).
International Cancer Genome Consortium. International network of cancer genome projects. Nature 464, 993–998 (2010).
Beuselinck, B. et al. Molecular subtypes of clear cell renal cell carcinoma are associated with sunitinib response in the metastatic setting. Clin. Cancer Res. 21, 1329–1339 (2015).
Erdem-Eraslan, L. et al. Identification of patients with recurrent glioblastoma who may benefit from combined bevacizumab and CCNU therapy: a report from the BELOB Trial. Cancer Res. 76, 525–534 (2016).
Jiang, P., Sellers, W. R. & Liu, X. S. Big data approaches for modeling response and resistance to cancer drugs. Annu. Rev. Biomed. Data Sci. 1, 1–27 (2018).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Garcia-Diaz, A. et al. Interferon receptor signaling pathways regulating PD-L1 and PD-L2 expression. Cell Rep. 19, 1189–1201 (2019).
Mariathasan, S. et al. TGF-β attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Jiang, P., Freedman, M. L., Liu, J. S. & Liu, X. S. Inference of transcriptional regulation in cancers. Proc. Natl Acad. Sci. USA 112, 7731–7736 (2015).
Liao, M. et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 26, 842–844 (2020).
Schulte-Schrepping, J. et al. Severe COVID-19 is marked by a dysregulated myeloid. Cell 182, 1419–1440 (2020).
Hadjadj, J. et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369, 718–724 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 (2020).
Zhao, Y. et al. Longitudinal COVID-19 profiling associates IL-1RA and IL-10 with disease severity and RANTES with mild disease. JCI Insight 5, e139834 (2020).
Han, H. et al. Profiling serum cytokines in COVID-19 patients reveals IL-6 and IL-10 are disease severity predictors. Emerg. Microbes Infect. 9, 1123–1130 (2020).
Mittal, S. K. & Roche, P. A. Suppression of antigen presentation by IL-10. Curr. Opin. Immunol. 34, 22–27 (2015).
Katakura, T., Miyazaki, M., Kobayashi, M., Herndon, D. N. & Suzuki, F. CCL17 and IL-10 as effectors that enable alternatively activated macrophages to inhibit the generation of classically activated macrophages. J. Immunol. 172, 1407–1413 (2004).
Orzalli, M. H. et al. An antiviral branch of the IL-1 signaling pathway restricts immune-evasive virus replication. Mol. Cell 71, 825–840 (2018).
Sun, J., Madan, R., Karp, C. L. & Braciale, T. J. Effector T cells control lung inflammation during acute influenza virus infection by producing IL-10. Nat. Med. 15, 277–284 (2009).
Kuri-Cervantes, L. et al. Comprehensive mapping of immune perturbations associated with severe COVID-19. Sci. Immunol. 5, eabd7114 (2020).
Barnes, B. J. et al. Targeting potential drivers of COVID-19: neutrophil extracellular traps. J. Exp. Med. 217, e20200652 (2020).
Bilusic, M. et al. Phase I trial of HuMax-IL8 (BMS-986253), an anti-IL-8 monoclonal antibody, in patients with metastatic or unresectable solid tumors. J. Immunother. Cancer 7, 240 (2019).
Chen, J. et al. PBMC fixation and processing for chromium single-cell RNA sequencing. J. Transl. Med. 16, 198 (2018).
Friedl, J. Mastering Regular Expressions (O’Reilly Media, 2006).
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847 (2007).
Demšar, J. et al. Orange: data mining toolbox in Python. J. Mach. Learn. Res. 14, 2349–2353 (2013).
Petryszak, R. et al. The RNASeq-er API—a gateway to systematically updated analysis of public RNA-seq data. Bioinformatics 33, 2218–2220 (2017).
Martinez, M. & Moon, E. K. CAR T cells for solid tumors: new strategies for finding, infiltrating, and surviving in the tumor microenvironment. Front. Immunol. 10, 128 (2019).
James, G., Witten, D., Hastie, T. & Tibshirani, R. An Introduction to Statistical Learning: with Applications in R (Springer Science & Business Media, 2013).
Storey, J. D. & Tibshirani, R. Statistical significance for genome-wide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Freedman, D. Statistical Models: Theory and Practice https://doi.org/10.1017/cbo9781139165495 (Cambridge University Press, 2005).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. https://doi.org/10.1145/2939672.2939785 (2016).
Bedinger, D. et al. Development and characterization of human monoclonal antibodies that neutralize multiple TGF-β isoforms. MAbs 8, 389–404 (2016).
Schurch, N. J. et al. How many biological replicates are needed in an RNA-seq experiment and which differential expression tool should you use? RNA 22, 839–851 (2016).
Yang, Y. et al. The outcome of TGF-β antagonism in metastatic breast cancer models in vivo reflects a complex balance between tumor-suppressive and proprogression activities of TGF-β. Clin. Cancer Res. 26, 643–656 (2020).
Zheng, C. et al. Landscape of infiltrating T cells in liver cancer revealed by single-cell sequencing. Cell 169, 1342–1356 (2017).
Acknowledgements
P.J. is supported by the intramural budget allocation from the NCI. L.L. is supported by grant K12HL138037 from the National Heart, Lung, and Blood Institute through the Yale Scholars in Implementation Science Career Development Program. K.W.W. is supported by the Ludwig Center at Harvard Medical School and National Institute of Health (NIH) grants R01 CA238039, R01 CA251599, P01 CA163222 and P01 CA236749. We thank J. Levine and the Center for Biomedical Informatics & Information Technology at NCI for setting up the web hosting infrastructures of CytoSig and FDC servers. This work utilized the computational resources of the NIH HPC Biowulf cluster (https://hpc.nih.gov/).
Author information
Authors and Affiliations
Contributions
P.J. and K.W.W. designed the study and wrote the manuscript. P.J. and Y.Z. curated all datasets. P.J. and B.R. performed computational analysis. P.J. created the CytoSig and FDC servers. R.P. deployed all servers under Amazon Web Services. Y.Z. performed cytokine treatments, flow cytometry and ELISA experiments in cell models. Y.Y. and L.W. performed the anti-TGF-β experiments in mice and prepared the RNA for sequencing. A.M. provided the TGF-β isoform-selective antibodies. G.A.-B., L.L. and E.R. participated in discussions.
Corresponding authors
Ethics declarations
Competing interests
K.W.W. is cofounder of Immunitas Therapeutics and also serves as the advisory board member of Immunitas Therapeutics, TCR2 Therapeutics, T-Scan Therapeutics, S.Q.Z. Biotech and Nextech Invest. K.W.W. received sponsored research funding from Novartis. The other authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks Nima Aghaeepour, Wendy Fantl and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Madhura Mukhopadhyay was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Curation of Human Cytokine Response Data.
a, The histogram of experimental cell-type models among collected treatment profiles. For each cell-type model, we counted the number of differential expression profiles upon cytokine treatment in that model as the Profile Count. b, The count of treatment response profiles for top-20 most frequently used cell models. c, The histogram of treatment durations among collected treatment profiles. For each duration, we counted the number of differential expression profiles upon cytokine treatment with that duration as the Profile Count. d, The count of treatment response profiles for top-20 most frequently used durations. e, The histogram of experimental doses among collected treatment profiles. For each dose, we counted the number of differential expression profiles upon cytokine treatment with that dose as the Profile Count. f, The count of treatment response profiles for top-20 most frequently used doses. g, The association between cytokine response profiles’ quality and treatment duration. We evaluated the quality for each treatment response profile as the correlation of expression levels between target gene scores and its ligand or receptor. Each dot represents one cytokine response profile with its treatment duration on the X-axis and median correlation across all TCGA or GTEx cohorts on the Y-axis. Activin A has high quality data with long treatment durations, while EGF has high quality data with transient treatment duration. h, The composite profile quality depends on the number of independent experiments merged. Four cytokines have more than 100 response profiles passing quality filters. We evaluated the quality of composite profiles after down-sampling the number of independent experiments merged. The quality metric is the correlation of expression between target genes and its ligand (blue) or receptor (yellow). The dots and error bars represent the median and standard deviation from 100 randomizations. Black triangle dots represent the down-sampling point that achieves 90% of the highest correlation. i, Similarities among IFNG response signatures from diverse cell models. An average response signature was computed for each cell model. Then, average response signatures were hierarchically clustered based on Pearson correlations.
Extended Data Fig. 2 Target Genes in Response to Cytokine Treatments.
a, Differential expression of target cytokines (x-axis) regulated by IL4 or BMP6 (y-axis). The thick line represents the median value. The bottom and top of the boxes are the 25th and 75th percentiles (interquartile range). The whiskers encompass 1.5 times the interquartile range. The difference between group values and zero among treatment profiles that passed quality controls was tested using the two-sided Wilcoxon signed-rank test, with p-values and sample counts labeled. b, Gene set enrichment among cytokine response targets. The normalized enrichment score from the gene set enrichment analysis (GSEA)6 represents the overall enrichment of pathway members among target genes from each cytokine’s composite signature. c, Normalized enrichment scores from panel b among pairs of hallmark categories and cytokines. d, Gating strategy of flow analysis. The forward scatter height (FSC-H) and side scatter height (SSC-H) are used for dead cell and cell debris removal. FSC-H width and SSC-H width are used to gate the single cells. We include unstained cells to define the gate threshold that separates positive populations and negative control cells.
Extended Data Fig. 3 Co-regulation between Cytokines on Target Genes.
a, Target co-regulation between TNFA and other anti-inflammatory cytokines. b, Target co-regulation between IFNG and other cytokines.
Extended Data Fig. 4 BMP6 represses CXCL8 and CCL2 induced by IL1B.
a, Representative plots of CXCL8 and CCL2 protein levels upon BMP6 and IL1B cotreatments in A549 cells. We utilized flow cytometry to measure CXCL8 and CCL2 intracellular protein levels after 12-hour treatments with IL1B, IL1B + BMP6, and media control. The X-axis shows the signal intensity measured by flow cytometry. The Y-axis shows the A549 fraction distribution with modal normalization, scaling the maximum Y-axis value to 100%. The percentage of cells with signal intensity above the gate threshold (vertical line, panel c) is indicated. b, Summary plots of CXCL8 and CCL2 protein levels upon BMP6 and IL1B treatments. In three cell-culture replicates, A549 and H1299 cells were treated either simultaneously or sequentially with combinations of BMP6 and IL1B. The mean fraction of cells with an intensity above the gate thresholds (defined in panel c) is plotted with standard deviations as error bars (n = 3 cell-culture replicates per condition). The two-sided Wilcoxon rank-sum test p-values were computed to compare groups. c, Gating strategy of flow analysis. The forward scatter height (FSC-H) and side scatter height (SSC-H) are used for dead cell and cell debris removal. FSC-H width and SSC-H width are used to gate the single cells. We include unstained cells to define the gate threshold that separates positive populations and negative control cells. d, CXCL8 and CCL2 soluble protein levels by ELISA in A549 cells. A549 cells were treated with BMP6 and IL1B in combinations for 24 hours. The mean soluble cytokine levels were measured by ELISA with standard error of the mean as error bars (n = 3 independent experiments).
Extended Data Fig. 5 CytoSig model selection.
a, Prediction performance measured as the cross-validation (CV) R2 in TCGA cohorts. For each algorithm, median 5-fold CV R2 metrics across all TCGA datasets were shown at different penalty values with standard error of the mean (SEM) as error bands. The horizontal line indicates 70% of the optimal CV R2 of ridge regression. The vertical line marks the lambda value of 10,000, the penalty used in the CytoSig model. b, Prediction performance measured as the cross-validation (CV) R2 metrics in GTEx cohorts, shown as panel a. c, Inference performance measured as the correlation between model coefficients and cytokine expression in TCGA cohorts. For each algorithm, we computed the median correlation values between model coefficients and cytokine ligand or receptor expression at different penalty values, with SEM as error bands. The vertical line marks the Lambda value of 10000, which is the penalty reaching 80% of the optimal correlation. XG Boost with tree learner cannot be evaluated for inference performance because its tree structure cannot provide coefficients as cytokine response. d, Inference performance measured as the correlation between model coefficients and cytokine expression in GTEx cohorts, shown as panel c.
Extended Data Fig. 6 CytoSig Predicts Cytokine Activities in Tumors and Cancer Therapy Response.
a, Receiver Operating Characteristic (ROC) curve of IFNG activity prediction. For each dataset from the ICGC, the ROC curve presents false-positive rates against true-positive rates at different IFNG activity thresholds. b, Area under the ROC Curve (AUC) as the prediction accuracy. We computed the ROC curves for all cytokines following the procedure in a. Each bar represents the median AUC among all ICGC cohorts, with standard errors of the mean as error bars (n = 9 independent datasets). The AUC baseline is 0.5, representing a random prediction. We applied the one-sided Wilcoxon signed-rank test to evaluate whether AUC values are higher than 0.5 for each signal, and converted p-values to false discovery rates (FDR) through the Benjamini-Hochberg correction. c, IFNG activity predicts overall survival upon Atezolizumab treatment in urothelial carcinoma43. The Kaplan-Meier plot presents patient fractions (Y-axis) with different overall survival (X-axis) among pre-treatment tumors with high and low IFNG activities predicted by CytoSig. The activity cutoff is selected through maximizing the difference between high and low groups. The p-value was from the two-sided Wald test using continuous values without cutoffs. d, Signaling activity computed by CytoSig better predicts clinical outcome than other metrics. We compared three approaches to compute cytokine activities, including expression levels of ligand, receptors, and CytoSig predictions. For IFNG activity, we also utilized a geneset signature developed by Merck to predict checkpoint blockade response41, as well as the PDL1 expression. The association between activity and survival outcome was computed as the Wald test z-score in Cox-PH regression.
Extended Data Fig. 7 CytoSig Reliably Predicts Cytokine Activity in Single Cells.
a, Result for a liver cancer cohort72 (n = 11 cell types per boxplot). The thick line represents the median value. The bottom and top of the boxes are the 25th and 75th percentiles (interquartile range). The whiskers encompass 1.5 times the interquartile range. b, Result for a COVID-19 peripheral blood cohort46 (n = 15 cell types per boxplot). The boxplot is defined as panel a.
Extended Data Fig. 8 Differential signaling activities from COVID-19 samples.
The heatmap presents cytokines whose predicted signaling activities are significantly different among severe, mild, healthy individuals in a COVID-19 peripheral blood cohort46 (false discovery rate < 0.05).
Extended Data Fig. 9 Gene expression of cytokines with differential signaling activities from COVID-19 samples.
IL1B and IL10 gene expression in diverse cell types. The violin plots present the expression of IL1B and IL10 in patient groups, with distributions smoothed by a kernel density estimator.
Extended Data Fig. 10 Potential Therapeutic Targets to Overcome COVID-19 Induced Tissue Damage.
a, Target genes of cytokines with elevated activities among severe patients. Arrow-headed edges indicate up-regulation, and flat-headed edges indicate down-regulation. For each cell type, red square nodes represent cytokines with high activity scores among severe patients, and blue diamonds represent anti-inflammatory signals with low activities. The network only includes targets whose expression values are significantly higher compared between severe and mild patients, and between disease and healthy controls. b, Example of gene expression in different patient groups. The violin plots present gene expression distributions in patient groups, smoothed by a kernel density estimator. Examples from lavage macrophages45 are in the left, and peripheral blood neutrophils46 are in the right. Y axis indicated values from individual cells.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Jiang, P., Zhang, Y., Ru, B. et al. Systematic investigation of cytokine signaling activity at the tissue and single-cell levels. Nat Methods 18, 1181–1191 (2021). https://doi.org/10.1038/s41592-021-01274-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01274-5
This article is cited by
-
Aberrant R-loop–mediated immune evasion, cellular communication, and metabolic reprogramming affect cancer progression: a single-cell analysis
Molecular Cancer (2024)
-
Dictionary of immune responses to cytokines at single-cell resolution
Nature (2024)
-
A single-cell atlas of lung homeostasis reveals dynamic changes during development and aging
Communications Biology (2024)
-
Integrating single-cell multi-omics and prior biological knowledge for a functional characterization of the immune system
Nature Immunology (2024)
-
Comprehensive characterization of IFNγ signaling in acute myeloid leukemia reveals prognostic and therapeutic strategies
Nature Communications (2024)