Abstract
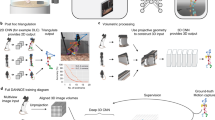
Markerless three-dimensional (3D) pose estimation has become an indispensable tool for kinematic studies of laboratory animals. Most current methods recover 3D poses by multi-view triangulation of deep network-based two-dimensional (2D) pose estimates. However, triangulation requires multiple synchronized cameras and elaborate calibration protocols that hinder its widespread adoption in laboratory studies. Here we describe LiftPose3D, a deep network-based method that overcomes these barriers by reconstructing 3D poses from a single 2D camera view. We illustrate LiftPose3D’s versatility by applying it to multiple experimental systems using flies, mice, rats and macaques, and in circumstances where 3D triangulation is impractical or impossible. Our framework achieves accurate lifting for stereotypical and nonstereotypical behaviors from different camera angles. Thus, LiftPose3D permits high-quality 3D pose estimation in the absence of complex camera arrays and tedious calibration procedures and despite occluded body parts in freely behaving animals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The datasets used in this paper and their sources are listed in Supplementary Table 2. Data used for Figs. 1–3 and Extended Data Figs. 1 and 2 can be downloaded from https://doi.org/10.7910/DVN/KHFAEI. Source data are provided with this paper.
Code availability
LiftPose3D code can be installed as a pip package at https://pypi.org/project/liftpose/. The source code and custom software used to acquire images with the LiftPose3D station is available at https://doi.org/10.5281/zenodo.5031774. The code is licensed under GNU General Public License v.3.0.
References
Pereira, T. D. et al. Fast animal pose estimation using deep neural networks. Nat. Methods 16, 117–125 (2019).
Mathis, A. et al. DeepLabCut: markerless pose estimation of user-defined body parts with deep learning. Nat. Neurosci. 21, 1281–1289 (2018).
Günel, S. et al. DeepFly3D, a deep learning-based approach for 3D limb and appendage tracking in tethered, adult Drosophila. eLife 8, 3686 (2019).
C. Bala, P. et al. Automated markerless pose estimation in freely moving macaques with OpenMonkeyStudio. Nat. Commun. 11, 4560 (2020).
Newell, A., Yang, K. & Deng, J. Stacked hourglass networks for human pose estimation. In Proc. European Conference on Computer Vision (ECCV) (2016).
Graving, J. M. et al. Deepposekit, a software toolkit for fast and robust animal pose estimation using deep learning. eLife 8, e47994 (2019).
Fang, H.-S., Xie, S., Tai, Y.-W. & Lu, C. RMPE: Regional multi-person pose estimation. In Proc. IEEE International Conference on Computer Vision (ICCV) (2017).
Wei, S.-E., Ramakrishna, V., Kanade, T. & Sheikh, Y. Convolutional pose machines. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2016).
Cao, Z., Simon, T., Wei, S.-E. & Sheikh, Y. Realtime multi-person 2D pose estimation using part affinity fields. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2017).
Hartley, R. & Zisserman, A. Multiple View Geometry in Computer Vision 2nd edn (Cambridge University Press, Cambridge, 2003).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Seelig, J. D. et al. Two-photon calcium imaging from head-fixed Drosophila during optomotor walking behavior. Nat. Methods 7, 535–540 (2010).
Gaudry, Q., Hong, E. J., Kain, J., de Bivort, B. L. & Wilson, R. I. Asymmetric neurotransmitter release enables rapid odour lateralization in Drosophila. Nature 493, 424–428 (2013).
Machado, A. S., Darmohray, D. M., Fayad, J., Marques, H. G. & Carey, M. R. A quantitative framework for whole-body coordination reveals specific deficits in freely walking ataxic mice. eLife 4, e07892 (2015).
Isakov, A. et al. Recovery of locomotion after injury in Drosophila melanogaster depends on proprioception. J. Exp. Biol. 219, 1760–1771 (2016).
Uhlmann, V., Ramdya, P., Delgado-Gonzalo, R., Benton, R. & Unser, M. Flylimbtracker: an active contour based approach for leg segment tracking in unmarked, freely behaving Drosophila. PLoS ONE 12, e0173433 (2017).
DeAngelis, B. D., Zavatone-Veth, J. A. & Clark, D. A. The manifold structure of limb coordination in walking Drosophila. eLife 8, 137 (2019).
Lee, H.-J. & Chen, Z. Determination of 3D human body postures from a single view. Comp. Vis. Graphics Image Proc. 30, 148–168 (1985).
Taylor, C. J. Reconstruction of articulated objects from point correspondences in a single uncalibrated image. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2000).
Chen, C. & Ramanan, D. 3D human pose estimation = 2D pose estimation + matching. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2017).
Gupta, A., Martinez, J., Little, J. J. & Woodham, R. J. 3D pose from motion for cross-view action recognition via non-linear circulant temporal encoding. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2014).
Sun, J. J. et al. View-invariant probabilistic embedding for human pose. In Proc. European Conference on Computer Vision (ECCV) (2020).
Nibali, A., He, Z., Morgan, S. & Prendergast, L. 3D human pose estimation with 2D marginal heatmaps. In Proc. IEEE Winter Conference on Applications of Computer Vision (WACV) (2019).
Zhao, L., Peng, X., Tian, Y., Kapadia, M. & Metaxas, D. N. Semantic graph convolutional networks for 3D human pose regression. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2019).
Iskakov, K., Burkov, E., Lempitsky, V. & Malkov, Y. Learnable triangulation of human pose. In Proc. International Conference on Computer Vision (ICCV) (2019).
Kanazawa, A., Zhang, J. Y., Felsen, P. & Malik, J. Learning 3D human dynamics from video. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2019).
Mehta, D. et al. XNect: real-time multi-person 3D motion capture with a single RGB camera. In Proc. ACM Transactions on Graphics (2020).
Rematas, K., Nguyen, C. H., Ritschel, T., Fritz, M. & Tuytelaars, T. Novel views of objects from a single image. IEEE Trans. Patt. Anal. Machine Intell. 39, 1576–1590 (2017).
Rhodin, H., Constantin, V., Katircioglu, I., Salzmann, M. & Fua, P. Neural scene decomposition for multi-person motion capture. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2019).
Martinez, J., Hossain, R., Romero, J. & Little, J. J. A simple yet effective baseline for 3D human pose estimation. In Proc. IEEE International Conference on Computer Vision (ICCV) (2017).
Pavllo, D., Feichtenhofer, C., Grangier, D. & Auli, M. 3D human pose estimation in video with temporal convolutions and semi-supervised training. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2019).
Liu, J., Guang, Y. & Rojas, J. GAST-Net: graph attention spatio-temporal convolutional networks for 3D human pose estimation in video. Preprint at https://arxiv.org/abs/2003.14179 (2020).
Cai, Y. et al. Exploiting spatial-temporal relationships for 3D pose estimation via graph convolutional networks. In Proc. IEEE International Conference on Computer Vision (ICCV) (2019).
Yiannakides, A., Aristidou, A. & Chrysanthou, Y. Real-time 3D human pose and motion reconstruction from monocular RGB videos. Comput. Animat. Virt. Worlds (2019).
Card, G. & Dickinson, M. H. Visually mediated motor planning in the escape response of Drosophila. Curr. Biol. 18, 1300–1307 (2008).
Wosnitza, A., Bockemühl, T., Dübbert, M., Scholz, H. & Büschges, A. Inter-leg coordination in the control of walking speed in Drosophila. J. Exp. Biol. 216, 480–491 (2013).
Marshall, J. D. et al. Continuous whole-body 3D kinematic recordings across the rodent behavioral repertoire. Neuron 109, 420–437.e8 (2021).
De Bono, M. & Bargmann, C. I. Natural variation in a neuropeptide Y receptor homolog modifies social behavior and food response in C. elegans. Cell 94, 679–689 (1998).
Budick, S. A. & O’Malley, D. M. Locomotor repertoire of the larval zebrafish: swimming, turning and prey capture. J. Exp. Biol. 203, 2565–2579 (2000).
Louis, M., Huber, T., Benton, R., Sakmar, T. P. & Vosshall, L. B. Bilateral olfactory sensory input enhances chemotaxis behavior. Nat. Neurosci. 11, 187–199 (2008).
Strauss, R. & Heisenberg, M. Coordination of legs during straight walking and turning in Drosophila melanogaster. J. Comp. Physiol. A. 167, 403–412 (1990).
Clarke, K. & Still, J. Gait analysis in the mouse. Physiol. Behav. 66, 723–729 (1999).
Wiltschko, A. B. et al. Mapping sub-second structure in mouse behavior. Neuron 88, 1121–1135 (2015).
Hong, W. et al. Automated measurement of mouse social behaviors using depth sensing, video tracking, and machine learning. Proc. Natl. Acad. Sci. USA 112, E5351–E5360 (2015).
Mendes, C. S., Bartos, I., Akay, T., Márka, S. & Mann, R. S. Quantification of gait parameters in freely walking wild type and sensory deprived Drosophila melanogaster. eLife 2, 231 (2013).
Feng, K. et al. Distributed control of motor circuits for backward walking in Drosophila. Nat. Commun. 11, 6166 (2020).
Alp Güler, R., Neverova, N. & Kokkinos, I. Densepose: dense human pose estimation in the wild. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2018).
Güler, R. A. & Kokkinos, I. Holopose: holistic 3D human reconstruction in-the-wild. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2019).
Loper, M., Mahmood, N., Romero, J., Pons-Moll, G. & Black, M. J. SMPL: a skinned multi-person linear model. ACM Trans. Graphics (Proc. SIGGRAPH Asia) 34, 248:1–248:16 (2015).
Zhang, J. Y., Felsen, P., Kanazawa, A. & Malik, J. Predicting 3D human dynamics from video. In Proc. IEEE International Conference on Computer Vision (ICCV) (2019).
Zuffi, S., Kanazawa, A., Berger-Wolf, T. & Black, M. J. Three-D safari: learning to estimate zebra pose, shape, and texture from images ‘in the wild’. In Proc. IEEE International Conference on Computer Vision (ICCV) (2019).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Nair, V. & Hinton, G. E. Rectified linear units improve restricted Boltzmann machines. In Proc. International Conference on Machine Learning (ICML) (2010).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2016).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. In Proc. International Conference on Learning Representations (ICLR) (2015).
Ioffe, S. & Szegedy, C. Batch normalization: accelerating deep network training by reducing internal covariate shift. In Proc. International Conference on Machine Learning (ICML) (2015).
Wandt, B., Rudolph, M., Zell, P., Rhodin, H. & Rosenhahn, B. CanonPose: self-supervised monocular 3D human pose estimation in the wild. In Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2021).
Cao, J. et al. Cross-domain adaptation for animal pose estimation. In Proc. IEEE International Conference on Computer Vision (ICCV) (2019).
Sanakoyeu, A., Khalidov, V., McCarthy, M. S., Vedaldi, A. & Neverova, N. Transferring dense pose to proximal animal classes. In IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2020).
Sridhar, V. H., Roche, D. G. & Gingins, S. Tracktor: image-based automated tracking of animal movement and behaviour. Meth. Ecol. Evol. 10, 815–820 (2019).
Acknowledgements
P.R. acknowledges support from an SNSF Project grant (no. 175667) and an SNSF Eccellenza grant (no. 181239). A.G. acknowledges support from an HFSP Cross-disciplinary Postdoctoral Fellowship (grant no. LT000669/2020-C). S.G. acknowledges support from an EPFL SV iPhD grant. D.M. holds a Marie Curie EuroTech postdoctoral fellowship and acknowledges that this project has received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement no. 754462. V.L.-R. acknowledges support from the Mexican National Council for Science and Technology, CONACYT, under the grant number 709993. We thank the Ölveczky laboratory (Center for Brain Science, Harvard University, Boston, MA, USA) for providing us with the CAPTURE dataset. We thank the Carey laboratory (Champalimaud Centre for the Unknown, Lisbon, Portugal) for the LocoMouse dataset.
Author information
Authors and Affiliations
Contributions
A.G. was responsible for conceptualization, methodology, software, hardware (Drosophila prism-mirror system), formal analysis, data curation, writing the original draft and reviewing and editing. S.G. was responsible for conceptualization, methodology, software, formal analysis, data curation, writing the original draft and reviewing and editing the paper. V.L.-R. was responsible for software and hardware (LiftPose3D station, low-resolution Drosophila ventral view system), data curation, and reviewing and editing the paper. M.P.A. was responsible for methodology, software (LiftPose3D), preliminary analysis of DeepFly3D dataset, data curation, and reviewing and editing the paper. D.M. was responsible for investigation (low-resolution Drosophila experiments), and reviewing and editing the paper. H.R. was responsible for conceptualization, and reviewing and editing the paper. P.F. was responsible for reviewing and editing the paper and funding acquisition. P.R. was responsible for conceptualization, hardware (Drosophila prism-mirror system), resources, writing the original draft, and reviewing and editing the paper, supervision, project administration and funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks Katarzyna Bozek, Giorgio Glilestro and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Joint angles resulting from lifting compared with 3D triangulated ground truth and 2D projections.
Joint angles α, β, γ, and ω for the front, mid, and hind left legs during forward walking. Shown are angles computed from 3D triangulation using DeepFly3D (blue), LiftPose3D predictions (red), and ventral 2D projections \(\alpha ^{\prime} ,\beta ^{\prime} ,\gamma ^{\prime}\), and \(\omega ^{\prime}\) (green). The mean (solid lines) and standard deviation of joint error distributions (transparency) are shown. Joint angles were computed by Monte Carlo sampling and errors were computed by taking the fluctuation in bone lengths.
Extended Data Fig. 2 Training and test loss convergence of the LiftPose3D network applied to a variety of datasets.
A-E Absolute test errors of LiftPose3D for all joints as a function of optimization epoch. A Two-camera data of Drosophila on a spherical treadmill (each color denotes a different pair of diametrically opposed cameras). B OpenMonkeyStudio dataset (each color denotes a different training run). C Single-camera data of Drosophila behaving freely in the right-angle prism-mirror system. D LocoMouse dataset. E CAPTURE dataset.
Extended Data Fig. 3 Drosophila LiftPose3D station.
A CAD drawing of the LiftPose3D station indicating major components (color-coded). B Photo of the LiftPose3D station. C Electronic circuit for building the illumination module on a pre-fabricated prototyping board (see Supplementary Table 1), electronic components and additional wiring are indicated (color-coded). D Printed circuit board provided as an alternative to the pre-fabricated board for building the illumination module.
Supplementary information
Supplementary Information
Supplementary Note 1 and Table 1.
Supplementary Video 1
3D pose lifting for backward walking in tethered Drosophila obtained from two side cameras. Videos obtained from cameras 2 (top left) and 5 (bottom left). DeepFly3D-derived 2D poses are superimposed. Orange circle indicates that the optogenetic stimulation LED light is on, activating MDNs to elicit backward walking. Right, 3D poses obtained by triangulating six camera views using DeepFly3D (solid lines), or lifting two camera views using LiftPose3D (dashed lines).
Supplementary Video 2
3D pose lifting for antennal grooming in tethered Drosophila obtained from two side cameras. Videos obtained from cameras 2 (top left) and 5 (bottom left). DeepFly3D-derived 2D poses are superimposed. Orange circle indicates that the optogenetic stimulation LED light is on, activating aDNs to elicit antennal grooming. Right, 3D poses obtained by triangulating six camera views using DeepFly3D (solid lines), or lifting two camera views using LiftPose3D (dashed lines).
Supplementary Video 3
3D pose lifting for irregular spontaneous limb movements in tethered Drosophila obtained from two side cameras. Videos obtained from cameras 2 (top left) and 5 (bottom left). DeepFly3D-derived 2D poses are superimposed. Right, 3D poses obtained by triangulating six camera views using DeepFly3D (solid lines), or lifting two camera views using LiftPose3D (dashed lines).
Supplementary Video 4
3D pose lifting of previously published OpenMonkeyStudio dataset4 of a freely moving macaque. Left, Single image drawn randomly from one of 62 cameras. Middle, ground truth 3D poses based on triangulation of 2D poses from up to 62 cameras (solid lines), or lifting from a single camera view using LiftPose3D (dashed lines). Right, error distribution across the 62 cameras for a given pose. Camera locations (circles) are color-coded by error. Gray circles denote cameras for which an image was not available. Green circle denotes the camera from which the image was used.
Supplementary Video 5
3D pose lifting of freely behaving Drosophila when triangulation is only partially possible. Single camera images of the ventral (top left) and side (bottom left) views. DeepLabCut-derived 2D poses are superimposed. Right, 3D poses obtained by triangulating partially available multi-view 2D poses (solid lines), or by lifting the ventral 2D pose using LiftPose3D (dashed lines).
Supplementary Video 6
3D pose lifting of previously published freely behaving mouse data14 when triangulation is only partially possible. Side (top left) and ventral (bottom left) views of a freely walking mouse. Superimposed are keypoints on the paws, mouth and proximal tail tracked using the LocoMouse software (blue circles). Using only the ventral view 2D pose, a trained LiftPose3D network can accurately track keypoints in the side view (orange circles).
Supplementary Video 7
3D pose lifting for freely behaving rats in a naturalistic arena. Left, ground truth 3D poses triangulated from six cameras (solid lines) superimposed with LiftPose3D’s predictions using 2D poses from one camera (dashed lines). Right, images from one camera with 2D poses acquired using CAPTURE are superimposed.
Supplementary Video 8
3D pose lifting for low-resolution videos of freely behaving flies when triangulation is impossible. Top, three freely behaving Drosophila in a rounded square arena and recorded ventrally using a single low-resolution camera. Of these, fly 0 is tracked, cropped and rotated leftward. Superimposed are 2D poses for 24 visible joints. (bottom) 3D poses lifted from ventral view 2D poses (x–y plane) permit analysis of leg kinematics in the otherwise unobserved x–z plane.
Supplementary Video 9
3D pose lifting of previously published ventral view videos16 of freely behaving flies when triangulation is impossible. Top, video from a freely behaving fly within a pill-shaped arena and recorded ventrally using a single high-resolution camera. Bottom left, following tracking, a region-of-interest containing the fly was cropped and rotated to maintain a leftward orientation. Superimposed are 2D poses estimated for 24 visible joints. Bottom middle, 3D poses obtained by lifting ventral view 2D poses. Bottom right, 3D poses lifted from ventral view 2D poses (top) permit analysis of leg kinematics in the otherwise unobserved x–z plane (bottom).
Supplementary Video 10
3D pose lifting of data from the Drosophila LiftPose3D station. Left, video of a freely behaving fly in the LiftPose3D station arena. Middle, cropped video around the centroid of the tracked fly, superimposed with 2D pose predictions. Right, lifted 3D poses obtained using ventral 2D poses.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Gosztolai, A., Günel, S., Lobato-Ríos, V. et al. LiftPose3D, a deep learning-based approach for transforming two-dimensional to three-dimensional poses in laboratory animals. Nat Methods 18, 975–981 (2021). https://doi.org/10.1038/s41592-021-01226-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01226-z
This article is cited by
-
WormSwin: Instance segmentation of C. elegans using vision transformer
Scientific Reports (2023)
-
3D mouse pose from single-view video and a new dataset
Scientific Reports (2023)
-
Automated phenotyping of postoperative delirium-like behaviour in mice reveals the therapeutic efficacy of dexmedetomidine
Communications Biology (2023)
-
Three-dimensional surface motion capture of multiple freely moving pigs using MAMMAL
Nature Communications (2023)
-
Three-dimensional unsupervised probabilistic pose reconstruction (3D-UPPER) for freely moving animals
Scientific Reports (2023)