Abstract
Serotonin (5-HT) is an important signaling monoamine and neurotransmitter. We report structure-guided engineering of a green fluorescent, genetically encoded serotonin sensor (G-GESS) from a 5-HT-binding lipocalin in the soft tick Argas monolakensis. G-GESS shows fast response kinetics and high affinity, specificity, brightness and photostability. We used G-GESS to image 5-HT dynamics in cultured cells, brain slices and behaving mice.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The gene sequence for G-GESS has been deposited with GenBank under the accession number MW027675. The plasmids for pDisplay-G-GESS-M (plasmid 154273), pAAV-hSyn-G-GESS-M (plasmid 154274), pLenti-hSyn-G-GESS-M (plasmid 154275), pDisplay-G-dGESS-M (plasmid 154276), pAAV-hSyn-G-dGESS-M (plasmid 154277) and pLenti-hSyn-G-GESS-M (plasmid 154278) and their sequence information have been deposited with Addgene. Source data are provided with this paper.
Change history
23 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41592-021-01148-w
References
Berger, M., Gray, J. A. & Roth, B. L. The expanded biology of serotonin. Annu. Rev. Med. 60, 355–366 (2009).
Sabatini, B. L. & Tian, L. Imaging neurotransmitter and neuromodulator dynamics in vivo with genetically encoded indicators. Neuron 108, 17–32 (2020).
Chefer, V. I., Thompson, A. C., Zapata, A. & Shippenberg, T. S. Overview of brain microdialysis. Curr. Protoc. Neurosci. Chapter 7, Unit 7.1 (2009).
Puthongkham, P. & Venton, B. J. Recent advances in fast-scan cyclic voltammetry. Analyst 145, 1087–1102 (2020).
Jeong, S. et al. High-throughput evolution of near-infrared serotonin nanosensors. Sci. Adv. 5, eaay3771 (2019).
Hettie, K. S. & Glass, T. E. Turn-on near-infrared fluorescent sensor for selectively imaging serotonin. ACS Chem. Neurosci. 7, 21–25 (2016).
Borden, P. M. et al. A fast genetically encoded fluorescent sensor for faithful in vivo acetylcholine detection in mice, fish, worms and flies. Preprint at bioRxiv https://doi.org/10.1101/2020.04.18.048397 (2020).
Tubio, M. R. et al. Expression of a G protein-coupled receptor (GPCR) leads to attenuation of signaling by other GPCRs: experimental evidence for a spontaneous GPCR constitutive inactive form. J. Biol. Chem. 285, 14990–14998 (2010).
Mans, B. J., Ribeiro, J. M. & Andersen, J. F. Structure, function, and evolution of biogenic amine-binding proteins in soft ticks. J. Biol. Chem. 283, 18721–18733 (2008).
Mans, B. J., Calvo, E., Ribeiro, J. M. & Andersen, J. F. The crystal structure of D7r4, a salivary biogenic amine-binding protein from the malaria mosquito Anopheles gambiae. J. Biol. Chem. 282, 36626–36633 (2007).
Hofherr, A., Fakler, B. & Klöcker, N. Selective Golgi export of Kir2.1 controls the stoichiometry of functional Kir2.x channel heteromers. J. Cell Sci. 118, 1935–1943 (2005).
Cataldo Bascuñan, L. R., Lyons, C., Bennet, H., Artner, I. & Fex, M. Serotonergic regulation of insulin secretion. Acta Physiol. (Oxf.) 225, e13101 (2019).
Rohm, B. et al. Nonivamide, a capsaicin analog, increases dopamine and serotonin release in SH-SY5Y cells via a TRPV1-independent pathway. Mol. Nutr. Food Res. 57, 2008–2018 (2013).
Fletcher, P. J., Korth, K. M., Robinson, S. R. & Baker, G. B. Multiple 5-HT receptors are involved in the effects of acute MDMA treatment: studies on locomotor activity and responding for conditioned reinforcement. Psychopharmacol. (Berl.) 162, 282–291 (2002).
Bauer, E. P. Serotonin in fear conditioning processes. Behav. Brain Res. 277, 68–77 (2015).
Fujino, K. et al. Increased serotonin release in mice frontal cortex and hippocampus induced by acute physiological stressors. Neurosci. Lett. 320, 91–95 (2002).
Bindels, D. S. et al. mScarlet: a bright monomeric red fluorescent protein for cellular imaging. Nat. Methods 14, 53–56 (2017).
Paxinos, G., Burt, J., Atrens, D. M. & Jackson, D. M. 5-hydroxytryptamine depletion with para-chlorophenylalanine: effects on eating, drinking, irritability, muricide and copulation. Pharmacol. Biochem. Behav. 6, 439–447 (1977).
Unger, E. K. et al. Directed evolution of a selective and sensitive serotonin sensor via machine learning. Cell 183, 1986–2002.e1926 (2020).
Wan, J. et al. A genetically encoded GRAB sensor for measuring serotonin dynamics. Preprint at bioRxiv https://doi.org/10.1101/2020.02.24.962282 (2020).
Flower, D. R., North, A. C. & Sansom, C. E. The lipocalin protein family: structural and sequence overview. Biochim. Biophys. Acta 1482, 9–24 (2000).
Skerra, A. Alternative binding proteins: anticalins - harnessing the structural plasticity of the lipocalin ligand pocket to engineer novel binding activities. FEBS J. 275, 2677–2683 (2008).
Marvin, J. S. et al. An optimized fluorescent probe for visualizing glutamate neurotransmission. Nat. Methods 10, 162–170 (2013).
Hochbaum, D. R. et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat. Methods 11, 825–833 (2014).
Lin, J. Y., Knutsen, P. M., Muller, A., Kleinfeld, D. & Tsien, R. Y. ReaChR: a red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nat. Neurosci. 16, 1499–1508 (2013).
Zhao, Y. et al. An expanded palette of genetically encoded Ca(2)(+) indicators. Science 333, 1888–1891 (2011).
Marvin, J. S., Schreiter, E. R., Echevarría, I. M. & Looger, L. L. A genetically encoded, high-signal-to-noise maltose sensor. Proteins 79, 3025–3036 (2011).
Chen, S., Chen, Z. J., Ren, W. & Ai, H. W. Reaction-based genetically encoded fluorescent hydrogen sulfide sensors. J. Am. Chem. Soc. 134, 9589–9592 (2012).
Fan, Y., Chen, Z. & Ai, H. W. Monitoring redox dynamics in living cells with a redox-sensitive red fluorescent protein. Anal. Chem. 87, 2802–2810 (2015).
Kardash, E., Bandemer, J. & Raz, E. Imaging protein activity in live embryos using fluorescence resonance energy transfer biosensors. Nat. Protoc. 6, 1835–1846 (2011).
Chen, Z. & Ai, H. W. Single fluorescent protein-based indicators for zinc ion (Zn2+). Anal. Chem. 88, 9029–9036 (2016).
Molina, R. S. et al. Understanding the fluorescence change in red genetically encoded calcium ion indicators. Biophys. J. 116, 1873–1886 (2019).
Drobizhev, M., Molina, R. S. & Hughes, T. E. Characterizing the two-photon absorption properties of fluorescent molecules in the 680-1300 nm spectral range. Bio-Protoc. 10, e3498 (2020).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Chen, M. et al. Genetically encoded, photostable indicators to image dynamic Zn(2+) secretion of pancreatic islets. Anal. Chem. 91, 12212–12219 (2019).
Burroughs, S. L. et al. Plate reader-based assays for measuring cell viability, neuroprotection and calcium in primary neuronal cultures. J. Neurosci. Methods 203, 141–145 (2012).
Rego, M. et al. Improved yield of AAV2 and rAAV2-retro serotypes following sugar supplementation during the viral production phase. bioRxiv https://doi.org/10.1101/488585 (2018).
Ding, B. & Kilpatrick, D. L. Lentiviral vector production, titration, and transduction of primary neurons. Methods Mol. Biol. 1018, 119–131 (2013).
Tetteh, H., Lee, J., Lee, J., Kim, J. G. & Yang, S. Investigating long-term synaptic plasticity in interlamellar hippocampus CA1 by electrophysiological field recording. J. Vis. Exp. https://doi.org/10.3791/59879 (2019).
Otis, J. M. et al. Prefrontal cortex output circuits guide reward seeking through divergent cue encoding. Nature 543, 103–107 (2017).
Holtmaat, A. et al. Long-term, high-resolution imaging in the mouse neocortex through a chronic cranial window. Nat. Protoc. 4, 1128–1144 (2009).
Acknowledgements
We thank S. Chen (Weill Cornell) for sharing MIN6 cells, Z. Chen for early exploration of this project, R. Molina for technical help with analyzing two-photon spectra and Y. Wang, Z. Li and other members in the Ai group for helpful discussion and assistance with experiments. Research reported in this publication was supported by funding to H.-w.A. (University of Virginia Start-up Package and NIH grant nos R01 DK122253, R01 GM118675 and R01 GM129291) and to M.D. (NIH grant nos U01 NS094246 and U24 NS109107).
Author information
Authors and Affiliations
Contributions
H.-w.A. conceived and supervised the project. X.L. and S. Zhao performed experiments involving brain slices and live mice. M.D. determined photophysical parameters and recorded two-photon excitation spectra. S. Zhang performed all other experiments, including protein engineering, characterization in vitro and in cultured cells, and preparation of viral vectors. All authors analyzed the data and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Methods thanks Bryan Roth, Jing Ren and Oliver Griesbeck for their contribution to the peer review of this work. Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
>This article has been retracted. Please see the retraction notice for more detail: https://dx.doi.org/10.1038/s41592-021-01148-w
Extended data
Extended Data Fig. 1 Structural analysis to identify potential cpGFP insertion sites.
a, Cartoon presentation of AM10. Loops showing ligand-dependent changes in dihedral angle (defined by the Cα atoms spanning every four sequential residues) are highlighted in red. b, Cα dihedral differences calculated from dihedrals between the ligand-bound (PDB 3BU1) and apo (PDB 3BS2) states of AM10 plotted against residue numbers. c, Dihedral angles for residue 113 of AM10 in the ligand-bound and apo states. d, Cartoon presentation of D7r4. Loop 2 showing a 5-HT-dependent change in dihedral angle is highlighted in red. e, Cα dihedral differences calculated from dihedrals between the ligand-bound (PDB 2QEH) and apo (PDB 2QEV) states of D7r4 plotted against residue numbers. f, Dihedral angles for residue 29 of D7r4 in the ligand-bound and apo states.
Extended Data Fig. 2 Process to engineer G-GESS.
a, Flowchart to illustrate our integrated, multi-step process to derive G-GESS. ΔF/F0 for each sensor generation is also presented. From G-GESS0.3 to G-GESS0.4, several N-terminal residues uninvolved in ligand binding were deleted, resulting in increased fluorescence change. However, it was found later in G-GESS0.5 that these residues were important for achieving good cell surface localization, so they were added back to G-GESS0.6. From G-GESS0.6 to G-GESS0.7, superfolder mutations (previously reported for sfGFP) were introduced into the cpGFP fragment of the sensor, resulting in improved brightness and fluorescence change. b, Sequence alignment of G-GESS with two earlier variants.
Extended Data Fig. 3 Two-photon fluorescence characterization of G-GESS.
a, Two-photon fluorescence excitation spectra in the presence (green) and absence (black) of 5-HT. b, Ratio of two-photon excitation in the presence over absence of 5-HT plotted against wavelength. These experiments were repeated twice with similar results.
Extended Data Fig. 4 Characterization of G-GESS-M for membrane localization in HEK 293 T cells.
a, Representative fluorescence images of HEK 293 T cells co-transfected with pDisplay-G-GESS-M and pCMV-R-GECO1. Red fluorescent Ca2+ sensor R-GECO1 was co-expressed as a whole-cell label. Scale bar, 20 µm. b, Fluorescence intensity measured over the white line shown in the ‘overlay’ image of panel a. These experiments were repeated three times with similar results using independent cultures.
Extended Data Fig. 5 Construction of G-dGESS, an unresponsive variant of G-GESS.
a, Cartoon presentation of AM182 (PDB 3BRN), highlighting the 5-HT ligand (magenta) and two residues (F58 and D106; colored in cyan and numbered according to PDB 3BRN) important for ligand binding. Mutations corresponding to F58A and D106L were introduced into G-GESS to create an unresponsive variant named ‘G-dGESS’. b, Representative fluorescence images of G-dGESS-M at the surface of HEK 293 T cells, in the presence or absence of 5-HT. This experiment was repeated three times with similar results using independent cell cultures. Scale bar, 20 µm.
Extended Data Fig. 6 Imaging of 5-HT release from β-cells and neuroblastoma cells.
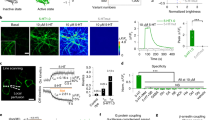
a, Representative images of G-GESS-M at the surface of mouse pancreatic MIN6 β-cells. Scale bar, 20 µm. b, Fluorescence traces of G-GESS-M in response to high glucose (25 mM). n = 7 cells for G-GESS-M and 6 cells for G-dGESS-M from 3 independent repeats. c, Representative images of G-GESS-M at the surface of SH-SY5Y neuroblastoma cells. Scale bar, 20 µm. d, Fluorescence traces of G-GESS-M in response to nonivamide (0.1 µM). n = 13 cells from 3 repeats for each group. Data are presented as mean ± s.d. of indicated replicates.
Extended Data Fig. 7 Imaging of 5-HT release in primary mouse neuron cultures.
a, Representative images of G-GESS-M at the surface of primary mouse neurons in response to 5-HT (10 µM). Also shown is a pseudocolor ratiometric image of the neuron in the presence over absence of 5-HT. Scale bar, 20 µm. This experiment was repeated three times with similar results using independent cultures. b, Fluorescence traces of G-GESS-M at the surface of primary mouse neurons, in response to high potassium induced membrane polarization. n = 19 cells for G-GESS-M and 12 cells for G-dGESS-M from 4 repeats. Data are presented as mean ± s.d. overlaid on the top of the traces of individual cells.
Extended Data Fig. 8 Evaluation of G-GESS-M cytotoxicity in primary mouse neurons.
a, MTT assay of primary neurons transfected with pDisplay-G-GESS-M or pDisplay-sfGFP-M in the absence or presence of the oxidative stress inducer, tert-butyl hydroperoxide (tBHP). P values determined by unpaired, two-tailed Student’s t-test. n = 6 cultures each group. b, Ca2+ mobilization assay of primary neurons transfected with pDisplay-G-GESS-M or pDisplay-sfGFP-M in response to depolarization with 100 mM KCl at the indicated time point. Intracellular Ca2+ was monitored with an orange fluorescent Ca2+ indicator (Enzo GFP-Certified Fluoforte Calcium Assay Kit). P values determined by unpaired, two-tailed Student’s t-test. n = 4 cultures each group. Individual data points and mean ± s.d. are presented. n.s., not significant.
Extended Data Fig. 9 Imaging of 5-HT release in acute mouse brain slices.
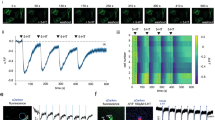
a, Schematic diagram illustrating the preparation of acute mouse brain slices with viral expression of G-GESS-M. b, Representative fluorescence images of G-GESS-M-expressing mouse brain slices. Scale bar, 200 µm. This experiment was repeated five times with three slice preparations and similar results were obtained. c, Representative fluorescence images of G-GESS-M-expressing hippocampal neurons in mouse brain slices in response to electrical stimuli (20 pulses at 64 Hz). Also shown is a pseudocolor ratiometric image (peak F/F0) of a representative neuron. Scale bar, 20 µm. This experiment was repeated six times with three slice preparations and similar results were obtained. d, Fluorescence traces of G-GESS-M in response to 20 pulses of electric stimuli at 50 V and the indicated frequencies. e, Fluorescence traces of G-GESS-M in response to 20 pulses of electric stimuli at 30 V and the indicated frequencies. f, Maximal fluorescence ratio change (peak F/F0) plotted against stimulation frequency at 50 V or 30 V. P values determined by 2-way ANOVA with Sidak’s multiple comparisons test. n = 6 cells on 3 slices from 2 mice for each group stimulated at 50 V, and 3 cells on 2 slices from 2 mice for each group stimulated at 30 V. Data are presented as mean ± s.d in all panels.
Extended Data Fig. 10 Bicistronic co-expression of G-GESS-M with a red fluorescent protein mScarlet-I to detect pharmacological modulation of 5-HT by p-chlorophenylalanine methyl ester (PCPA), a tryptophan hydroxylase inhibitor.
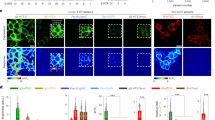
a, Schematic diagram illustrating the co-expression construct including G-GESS-M, a P2A peptide self-cleaving site, and mScarlet-I. b, Representative fluorescence images of the co-expression construct expressed in HEK 293 T cells in the absence and presence of 5-HT acquired with a Leica DMi8 SPE confocal microscope. Scale bar, 20 µm. c, Quantitative analysis of the green-to-red intensity ratios of HEK 293 T cells, in the absence or presence of 5-HT for G-dGESS-M-P2A-mScarlet-I and G-GESS-M-P2A-mScarlet-I. P values determined by unpaired, two-tailed Student’s t-test. n = 12 random cells from three independent cultures. d, Schematic diagram illustrating the mouse treatment timeline, including intracranial injection of AAV on Day 0 and PCPA injection from Day 7 to Day 11. Acute brain slices were prepared and imaged on Day 12. e, Representative fluorescence images of the co-expression construct expressed in the hippocampus of mice with or without PCPA treatment. Images were acquired with a Scientifica SliceScope Pro 1000. Scale bar, 20 µm. f, Quantitative analysis of the green-to-red intensity ratios of mouse brain slices, in the presence or absence of PCPA. P values determined by one-way ANOVA with Tukey’s multiple comparisons test. n = 7 cells on one 5-HT-treated slice expressing G-GESS-M-P2A-mScarlet-I, and 35 cells on 5 slices from two mice for all other groups. Individual data points and mean ± s.d. are presented. n.s., not significant.
Supplementary information
Supplementary Information
Supplementary Table 1 and Figs. 1 and 2.
Source data
Source Data Fig. 1
Figure and statistical source data.
Source Data Fig. 2
Figure and statistical source data.
Source Data Extended Data Fig. 1
Figure and statistical source data.
Source Data Extended Data Fig. 3
Figure and statistical source data.
Source Data Extended Data Fig. 4
Figure and Statistical source data.
Source Data Extended Data Fig. 6
Figure and statistical source data.
Source Data Extended Data Fig. 7
Figure and statistical source data.
Source Data Extended Data Fig. 8
Figure and statistical source data.
Source Data Extended Data Fig. 9
Figure and statistical source data.
Source Data Extended Data Fig. 10
Figure and statistical source data.
About this article
Cite this article
Zhang, S., Li, X., Zhao, S. et al. RETRACTED ARTICLE: A fast, high-affinity fluorescent serotonin biosensor engineered from a tick lipocalin. Nat Methods 18, 258–261 (2021). https://doi.org/10.1038/s41592-021-01078-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-021-01078-7
This article is cited by
-
Hierarchical TiO2–Ag/hydrogel coating used for outstanding antifouling detection of serotonin
Macromolecular Research (2024)
-
Making a tick protein talk as a serotonin sensor
Nature Methods (2021)