Abstract
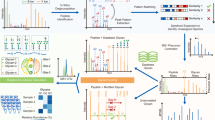
Recent advances in methods for enrichment and mass spectrometric analysis of intact glycopeptides have produced large-scale glycoproteomics datasets, but interpreting these data remains challenging. We present MSFragger-Glyco, a glycoproteomics mode of the MSFragger search engine, for fast and sensitive identification of N- and O-linked glycopeptides and open glycan searches. Reanalysis of recent N-glycoproteomics data resulted in annotation of 80% more glycopeptide spectrum matches (glycoPSMs) than previously reported. In published O-glycoproteomics data, our method more than doubled the number of glycoPSMs annotated when searching the same glycans as the original search, and yielded 4- to 6-fold increases when expanding searches to include additional glycan compositions and other modifications. Expanded searches also revealed many sulfated and complex glycans that remained hidden to the original search. With greatly improved spectral annotation, coupled with the speed of index-based scoring, MSFragger-Glyco makes it possible to comprehensively interrogate glycoproteomics data and illuminate the many roles of glycosylation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
N- and O-linked glycoproteomics raw data were downloaded from the PRIDE Archive53 and Proteome Xchange54 with accession numbers PXD011533 (Riley et al.10 N-glycan data), PXD009476 (Yang et al.12 O-glycan data), and PXD005411, PXD005412, PXD005413, PXD005553 and PXD005555 (Liu et al.8 N-glycan data). Processed search results (raw data, MSFragger output files and processed peak tables) that support the findings of this study are available in PRIDE (accession number PXD021196).
Code availability
The MSFragger-Glyco program was developed in the cross-platform Java language, and incorporated in the MSFragger search engine starting with v.3.0, which can be accessed at https://msfragger.nesvilab.org/.
References
Varki, A. Biological roles of glycans. Glycobiology 27, 3–49 (2017).
Thaysen-Andersen, M., Packer, N. H. & Schulz, B. L. Maturing glycoproteomics technologies provide unique structural insights into the N-glycoproteome and its regulation in health and disease. Mol. Cell. Proteomics 15, 1773–1790 (2016).
Chang, D. & Zala, J. Why glycosylation matters in building a better flu vaccine. Mol. Cell. Proteomics 18, 2348–2358.
Marsico, G., Russo, L., Quondamatteo, F. & Pandit, A. Glycosylation and integrin regulation in cancer. Trends Cancer 4, 537–552 (2018).
Schedin-Weiss, S., Winblad, B. & Tjernberg, L. O. The role of protein glycosylation in Alzheimer disease. FEBS J. 281, 46–62 (2014).
Wohlgemuth, J., Karas, M., Eichhorn, T., Hendriks, R. & Andrecht, S. Quantitative site-specific analysis of protein glycosylation by LC–MS using different glycopeptide-enrichment strategies. Anal. Biochem. 395, 178–188 (2009).
Rudd, P. M. & Dwek, R. A. Glycosylation: heterogeneity and the 3D structure of proteins. Crit. Rev. Biochem. Mol. Biol. 32, 1–100 (1997).
Liu, M. Q. et al. PGlyco 2.0 enables precision N-glycoproteomics with comprehensive quality control and one-step mass spectrometry for intact glycopeptide identification. Nat. Commun. 8, 438 (2017).
Suttapitugsakul, S., Sun, F. & Wu, R. Recent advances in glycoproteomicanalysis by mass spectrometry. Anal. Chem. 92, 267–291 (2020).
Riley, N. M., Hebert, A. S., Westphall, M. S. & Coon, J. J. Capturing site-specific heterogeneity with large-scale N-glycoproteome analysis. Nat. Communications 10, 1311–1311 (2019).
Reiding, K. R., Bondt, A., Franc, V. & Heck, A. J. R. The benefits of hybrid fragmentation methods for glycoproteomics. Trends Analyt. Chem. 108, 260–268 (2018).
Yang, W., Ao, M., Hu, Y., Li, Q. K. & Zhang, H. Mapping the O‐glycoproteome using site‐specific extraction of O‐linked glycopeptides (EXoO). Mol. Syst. Biol. 14, e8486 (2018).
King, S. L. et al. Characterizing the O-glycosylation landscape of human plasma, platelets, and endothelial cells. Blood Adv. 1, 429–442 (2017).
Bollineni, R. C., Koehler, C. J., Gislefoss, R. E., Anonsen, J. H. & Thiede, B. Large-scale intact glycopeptide identification by Mascot database search. Sci. Rep 8, https://doi.org/10.1038/s41598-018-20331-2 (2018).
Bern, M., Kil, Y. J. & Becker, C. Byonic: advanced peptide and protein identification software. Curr. Protoc. Bioinformatics 40, 13.20.1–13.20.14 (2012).
Eng, J. K., McCormack, A. L. & Yates, J. R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spec. 5, 976–989 (1994).
Zhu, Z., Hua, D., Clark, D. F., Go, E. P. & Desaire, H. GlycoPep detector: a tool for assigning mass spectrometry data of N-linked glycopeptides on the basis of their electron transfer dissociation spectra. Anal. Chem. 85, 5023–5032 (2013).
Yu, C. Y. et al. Automated glycan sequencing from tandem mass spectra of N-linked glycopeptides. Anal. Chem. 88, 5725–5732 (2016).
He, L., Xin, L., Shan, B., Lajoie, G. A. & Ma, B. GlycoMaster DB: software to assist the automated identification of N-linked glycopeptides by tandem mass spectrometry. J. Proteome Res. 13, 3881–3895 (2014).
Mayampurath, A. et al. Computational framework for identification of intact glycopeptides in complex samples. Anal. Chem. 86, 453–463 (2014).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D. & Nesvizhskii, A. I. MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry-based proteomics. Nat. Methods 14, 513–520 (2017).
Solntsev, S. K., Shortreed, M. R., Frey, B. L. & Smith, L. M. Enhanced global post-translational modification discovery with metamorpheus. J. Proteome Res. 17, 1844–1851 (2018).
Creasy, D. M. & Cottrell, J. S. Error tolerant searching of uninterpreted tandem mass spectrometry data. Proteomics 2, 1426–1434 (2002).
Ma, C. W. M. & Lam, H. Hunting for unexpected post-translational modifications by spectral library searching with tier-wise scoring. J. Proteome Res. 13, 2262–2271 (2014).
Chick, J. M. et al. A mass-tolerant database search identifies a large proportion of unassigned spectra in shotgun proteomics as modified peptides. Nat. Biotech. 33, 743–749 (2015).
Ahrné, E., Nikitin, F., Lisacek, F. & Müller, M. QuickMod: a tool for open modification spectrum library searches. J. Proteome Res. 10, 2913–2921 (2011).
Chi, H. et al. Comprehensive identification of peptides in tandem mass spectra using an efficient open search engine. Nat. Biotech. 36, 1059–1066 (2018).
Na, S., Bandeira, N. & Paek, E. Fast multi-blind modification search through tandem mass spectrometry. Mol. Cell. Proteomics 11, https://doi.org/10.1074/mcp.M111.010199 (2012).
Swearingen, K. E. et al. A tandem mass spectrometry sequence database search method for identification of O-fucosylated proteins by mass spectrometry. J. Proteome Res. 18, 652–663 (2019).
Trinidad, J. C., Schoepfer, R., Burlingame, A. L. & Medzihradszky, K. F. N- and O-Glycosylation in the murine synaptosome. Mol. Cell. Proteomics 12, 3474–3488 (2013).
Chalkley, R. J., Baker, P. R., Medzihardszky, K. F., Lynn, A. J. & Burlingame, A. L. In-depth analysis of tandem mass spectrometry data from disparate instrument types. Mol. Cell. Proteomics 7, 2386–2398 (2008).
Chalkley, R. J. & Baker, P. R. Use of a glycosylation site database to improve glycopeptide identification from complex mixtures. Anal. Bioanal. Chem. 409, 571–577 (2017).
Medzihradszky, K. F., Kaasik, K. & Chalkley, R. J. Tissue-specific glycosylation at the glycopeptide level. Mol. Cell. Proteomics 14, 2103–2110 (2015).
Yu, F. et al. Identification of modified peptides using localization-aware open search. Nat. Communications 11, 4065 (2020).
Seipert, R. R. et al. Factors that influence fragmentation behavior of N-linked glycopeptide ions. Anal. Chem. 80, 3684–3692 (2008).
Wuhrer, M., Deelder, A. M. & Van Der Burgt, Y. E. M. Mass spectrometric glycan rearrangements. Mass Spec. Rev. 30, 664–680 (2011).
Ledvina, A. R. et al. Infrared photoactivation reduces peptide folding and hydrogenatom migration following ETD tandem mass spectrometry. Angew. Chem. Int. Ed. 48, 8526–8528 (2009).
Vékey, K. et al. Fragmentation characteristics of glycopeptides. Int. J. Mass Spec. 345–347, 71–79 (2013).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Leprevost, F. D. et al. Philosopher: a versatile toolkit for shotgun proteomics data analysis. Nat. Methods 17, 869–870 (2020).
Hang, H. C. & Bertozzi, C. R. The chemistry and biology of mucin-type O-linked glycosylation. Bioorg. Med. Chem. 13, 5021–5034 (2005).
Jensen, P. H., Kolarich, D. & Packer, N. H. Mucin-type O-glycosylation—putting the pieces together. FEBS J. 277, 81–94 (2010).
Yang, Z. et al. The GalNAc-type O-glycoproteome of CHO cells characterized by the simplecell strategy. Mol. Cell Proteomics 13, 3224–3235 (2014).
Potel, C. M., Lemeer, S. & Heck, A. J. R. Phosphopeptide fragmentation and site localization by mass spectrometry: an update. Anal. Chem. 91, 126–141 (2019).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Hu, H., Khatri, K., Klein, J., Leymarie, N. & Zaia, J. A review of methods for interpretation of glycopeptide tandem mass spectral data. Glycoconjugate J. 33, 285–296 (2016).
Hu, H., Khatri, K. & Zaia, J. Algorithms and design strategies towards automated glycoproteomics analysis. Mass Spec. Rev. 36, 475–498 (2017).
Li, K., Vaudel, M., Zhang, B., Ren, Y. & Wen, B. PDV: an integrative proteomics data viewer. Bioinformatics 35, 1249–1251 (2019).
Röst, H. L., Schmitt, U., Aebersold, R. & Malmström, L. pyOpenMS: a Python-based interface to the OpenMS mass-spectrometry algorithm library. Proteomics 14, 74–77 (2014).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Deutsch, E. W. et al. The ProteomeXchange consortium in 2017: supporting the cultural change in proteomics public data deposition. Nucleic Acids Res. 45, D1100–D1106 (2017).
Acknowledgements
This work was funded in part by NIH grant nos. R01-GM-094231 and U24-CA210967.
Author information
Authors and Affiliations
Contributions
D.A.P., F.Y. and G.C.T. developed the algorithm. D.A.P. analyzed the data. A.I.N. conceived and supervised the project. D.A.P. and A.I.N. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Arunima, Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–3 and Tables 1–9.
Rights and permissions
About this article
Cite this article
Polasky, D.A., Yu, F., Teo, G.C. et al. Fast and comprehensive N- and O-glycoproteomics analysis with MSFragger-Glyco. Nat Methods 17, 1125–1132 (2020). https://doi.org/10.1038/s41592-020-0967-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0967-9
This article is cited by
-
Prediction of glycopeptide fragment mass spectra by deep learning
Nature Communications (2024)
-
Integrative proteomics and n-glycoproteomics reveal the synergistic anti-tumor effects of aspirin- and gemcitabine-based chemotherapy on pancreatic cancer cells
Cellular Oncology (2024)
-
A Novel Integrated Pipeline for Site-Specific Quantification of N-glycosylation
Phenomics (2024)
-
Particle elasticity influences polymeric artificial antigen presenting cell effectiveness in vivo via CD8+ T cell activation, macrophage uptake, and the protein corona
Nano Research (2024)
-
Oxonium ion scanning mass spectrometry for large-scale plasma glycoproteomics
Nature Biomedical Engineering (2023)