Abstract
The discovery and design of biologically important RNA molecules is outpacing three-dimensional structural characterization. Here, we demonstrate that cryo-electron microscopy can routinely resolve maps of RNA-only systems and that these maps enable subnanometer-resolution coordinate estimation when complemented with multidimensional chemical mapping and Rosetta DRRAFTER computational modeling. This hybrid ‘Ribosolve’ pipeline detects and falsifies homologies and conformational rearrangements in 11 previously unknown 119- to 338-nucleotide protein-free RNA structures: full-length Tetrahymena ribozyme, hc16 ligase with and without substrate, full-length Vibrio cholerae and Fusobacterium nucleatum glycine riboswitch aptamers with and without glycine, Mycobacterium SAM-IV riboswitch with and without S-adenosylmethionine, and the computer-designed ATP-TTR-3 aptamer with and without AMP. Simulation benchmarks, blind challenges, compensatory mutagenesis, cross-RNA homologies and internal controls demonstrate that Ribosolve can accurately resolve the global architectures of RNA molecules but does not resolve atomic details. These tests offer guidelines for making inferences in future RNA structural studies with similarly accelerated throughput.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Cryo-EM maps are available in the EMDB with accession codes EMD-21831, EMD-21832, EMD-21833, EMD-21834, EMD-21835, EMD-21836, EMD-21838, EMD-21839, EMD-21840, EMD-21841 and EMD-21842. Models (best-case) are available in the PDB with accession codes 6WLJ, 6WLK, 6WLL, 6WLM, 6WLN, 6WLO, 6WLQ, 6WLR, 6WLS, 6WLT and 6WLU. Fully automated models are available in the Supplementary Data. M2-seq and mutate-map-rescue data are available in the RMDB with accession codes RB1UTR_DMS_0000, 243RNA_DMS_0000, ATPAPO_DMS_0000, ATPAMP_DMS_0000, U1SNRNA_DMS_0000, SCARNA6_DMS_0000, VCKTAPO_DMS_0000, VCKTGLY_DMS_0000, DPRGLN_DMS_0000, ETERNA3_DMS_0000, SPINACH_DMS_0000, FNKTAPO_DMS_0000, FNKTGLY_DMS_0000, HC16APO_DMS_0000, HC16PRO_DMS_0000, SAM4APO_DMS_0000, SAM4SAM_DMS_0000, L21RNA_DMS_0000, HC16M2R_1M7_0001, HC16M2R_1M7_0002 and HC16M2R_1M7_0003. The raw particle image data that support the findings of this study are available from the corresponding author upon request.
Code availability
The auto-DRRAFTER software is freely available to academic users as part of the Rosetta software package. Documentation is available at https://www.rosettacommons.org/docs/latest/application_documentation/rna/auto-drrafter and a demo is available at https://www.rosettacommons.org/demos/latest/public/auto-drrafter/README. A limited version of the software is also freely available through an online ROSIE server at https://rosie.rosettacommons.org/auto-drrafter.
Change history
22 July 2020
The links to the EMDB and PDB accession codes in the Data Availability section have been corrected in the HTML and PDF versions of the article.
References
Atkins, J. F., Gesteland, R. F. & Cech, T. RNA Worlds: From Life’s Origins to Diversity in Gene Regulation (Cold Spring Harbor Laboratory Press, 2011).
Cech, T. R. & Steitz, J. A. The noncoding RNA revolution—trashing old rules to forge new ones. Cell 157, 77–94 (2014).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Hangauer, M. J., Vaughn, I. W. & McManus, M. T. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet. 9, e1003569 (2013).
Zhang, J. W. & Ferre-D’Amare, A. R. New molecular engineering approaches for crystallographic studies of large RNAs. Curr. Opin. Struc. Biol. 26, 9–15 (2014).
Zhang, H. & Keane, S. C. Advances that facilitate the study of large RNA structure and dynamics by nuclear magnetic resonance spectroscopy. Wiley Interdiscip. Rev. RNA 10, e1541 (2019).
Bergfors, T. Screening and optimization methods for nonautomated crystallization laboratories. Methods Mol. Biol. 363, 131–151 (2007).
McMullan, G., Faruqi, A. R., Clare, D. & Henderson, R. Comparison of optimal performance at 300 keV of three direct electron detectors for use in low dose electron microscopy. Ultramicroscopy 147, 156–163 (2014).
Ognjenovic, J., Grisshammer, R. & Subramaniam, S. Frontiers in cryo electron microscopy of complex macromolecular assemblies. Annu. Rev. Biomed. Eng. 21, 395–415 (2019).
Zhang, K. M. et al. Structure of the 30 kDa HIV-1 RNA dimerization signal by a hybrid Cryo-EM, NMR, and molecular dynamics approach. Structure 26, 490–498.e3 (2018).
Kappel, K. et al. De novo computational RNA modeling into cryo-EM maps of large ribonucleoprotein complexes. Nat. Methods 15, 947 (2018).
Kruger, K. et al. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31, 147–157 (1982).
Jaeger, L., Wright, M. C. & Joyce, G. F. A complex ligase ribozyme evolved in vitro from a group I ribozyme domain. Proc. Natl Acad. Sci. USA 96, 14712–14717 (1999).
Horning, D. P. & Joyce, G. F. Amplification of RNA by an RNA polymerase ribozyme. Proc. Natl Acad. Sci. USA 113, 9786–9791 (2016).
Mandal, M. et al. A glycine-dependent riboswitch that uses cooperative binding to control gene expression. Science 306, 275–279 (2004).
Weinberg, Z. et al. The aptamer core of SAM-IV riboswitches mimics the ligand-binding site of SAM-I riboswitches. RNA 14, 822–828 (2008).
Ames, T. D. & Breaker, R. R. Bacterial aptamers that selectively bind glutamine. RNA Biol. 8, 82–89 (2011).
Yesselman, J. D. et al. Computational design of three-dimensional RNA structure and function. Nat. Nanotechnol. 14, 866–873 (2019).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Lee, J. et al. RNA design rules from a massive open laboratory. Proc. Natl Acad. Sci. USA 111, 2122–2127 (2014).
Huang, L. L., Serganov, A. & Patel, D. J. Structural insights into ligand recognition by a sensing domain of the cooperative glycine riboswitch. Mol. Cell 40, 774–786 (2010).
Guo, F., Gooding, A. R. & Cech, T. R. Structure of the Tetrahymena ribozyme: base triple sandwich and metal ion at the active site. Mol. Cell 16, 351–362 (2004).
Butler, E. B., Xiong, Y., Wang, J. & Strobel, S. A. Structural basis of cooperative ligand binding by the glycine riboswitch. Chem. Biol. 18, 293–298 (2011).
Golden, B. L., Gooding, A. R., Podell, E. R. & Cech, T. R. A preorganized active site in the crystal structure of the Tetrahymena ribozyme. Science 282, 259–264 (1998).
Marz, M. et al. Animal snoRNAs and scaRNAs with exceptional structures. RNA Biol. 8, 938–946 (2011).
Krummel, D. A. P., Oubridge, C., Leung, A., Li, J. D. & Nagai, K. Crystal structure of a ten-subunit human spliceosomal U1 snRNP at 5.5 angstrom resolution. Biophys. J. 100, 198–198 (2011).
Kutchko, K. M. et al. Multiple conformations are a conserved and regulatory feature of the RB1 5′ UTR. RNA 21, 1274–1285 (2015).
Russel, D. et al. Putting the pieces together: integrative modeling platform software for structure determination of macromolecular assemblies. PloS Biol. 10, e1001244 (2012).
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015).
Cheng, C. Y., Kladwang, W., Yesselman, J. D. & Das, R. RNA structure inference through chemical mapping after accidental or intentional mutations. Proc. Natl Acad. Sci. USA 114, 9876–9881 (2017).
Li, S. et al. Structural basis of amino acid surveillance by higher-order tRNA-mRNA interactions. Nat. Struct. Mol. Biol. 26, 1094–1105 (2019).
Zhang, K. et al. Cryo-EM structure of a 40-kDa SAM-IV riboswitch RNA at 3.7 Å resolution. Nat. Commun. 10, 5511 (2019).
Schmitt, E. et al. Structure of the ternary initiation complex aIF2–GDPNP–methionylated initiator tRNA. Nat. Struct. Mol. Biol. 19, 450–454 (2012).
Lane, S. W. et al. Construction and crystal structure of recombinant STNV capsids. J. Mol. Biol. 413, 41–50 (2011).
Kladwang, W., Chou, F. C. & Das, R. Automated RNA structure prediction uncovers a kink-turn linker in double glycine riboswitches. J. Am. Chem. Soc. 134, 1404–1407 (2012).
Cheng, C. Y. et al. Consistent global structures of complex RNA states through multidimensional chemical mapping. eLife 4, e07600 (2015).
Trausch, J. J. et al. Structural basis for diversity in the SAM clan of riboswitches. Proc. Natl Acad. Sci. USA 111, 6624–6629 (2014).
Montange, R. K. & Batey, R. T. Structure of the S-adenosylmethionine riboswitch regulatory mRNA element. Nature 441, 1172–1175 (2006).
Lehnert, V., Jaeger, L., Michel, F. & Westhof, E. New loop–loop tertiary interactions in self-splicing introns of subgroup IC and ID: a complete 3D model of the Tetrahymena thermophila ribozyme. Chem. Biol. 3, 993–1009 (1996).
Tian, S. Q., Cordero, P., Kladwang, W. & Das, R. High-throughput mutate-map-rescue evaluates SHAPE-directed RNA structure and uncovers excited states. RNA 20, 1815–1826 (2014).
Tian, S., Kladwang, W. & Das, R. Allosteric mechanism of the V. vulnificus adenine riboswitch resolved by four-dimensional chemical mapping. eLife 7, e29602 (2018).
Zhao, D. Q. & Jardetzky, O. An assessment of the precision and accuracy of protein structures determined by Nmr—dependence on distance errors. J. Mol. Biol. 239, 601–607 (1994).
Tian, S., Yesselman, J. D., Cordero, P. & Das, R. Primerize: automated primer assembly for transcribing non-coding RNA domains. Nucleic Acids Res. 43, W522–W526 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290 (2017).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Das, R., Karanicolas, J. & Baker, D. Atomic accuracy in predicting and designing noncanonical RNA structure. Nat. Methods 7, 291–294 (2010).
DiMaio, F. & Chiu, W. Tools for model building and optimization into near-atomic resolution electron cryo-microscopy density maps. Methods Enzymol. 579, 255–276 (2016).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Leontis, N. B. & Zirbel, C. L. Nonredundant 3D Structure Datasets for RNA Knowledge Extraction and Benchmarking. In RNA 3D Structure Analysis and Prediction (eds Leontis, N. & Westhof, E.) 281–298 (Springer, 2012).
Huang, L., Ishibe-Murakami, S., Patel, D. J. & Serganov, A. Long-range pseudoknot interactions dictate the regulatory response in the tetrahydrofolate riboswitch. Proc. Natl Acad. Sci. USA 108, 14801–14806 (2011).
Gao, A. & Serganov, A. Structural insights into recognition of c-di-AMP by the ydaO riboswitch. Nat. Chem. Biol. 10, 787–792 (2014).
Kempf, G., Wild, K. & Sinning, I. Structure of the complete bacterial SRP Alu domain. Nucleic Acids Res. 42, 12284–12294 (2014).
Serganov, A., Huang, L. L. & Patel, D. J. Coenzyme recognition and gene regulation by a flavin mononucleotide riboswitch. Nature 458, 233–U210 (2009).
Lu, C. R. et al. SAM recognition and conformational switching mechanism in the Bacillus subtilis yitJ S Box/SAM-I riboswitch. J. Mol. Biol. 404, 803–818 (2010).
Cate, J. H. et al. Crystal structure of a group I ribozyme domain: principles of RNA packing. Science 273, 1678–1685 (1996).
Serganov, A., Huang, L. L. & Patel, D. J. Structural insights into amino acid binding and gene control by a lysine riboswitch. Nature 455, 1263–U1276 (2008).
Meyer, M. et al. Speciation of a group I intron into a lariat capping ribozyme. Proc. Natl Acad. Sci. USA 111, 7659–7664 (2014).
Wang, R. Y. R. et al. Automated structure refinement of macromolecular assemblies from cryo-EM maps using Rosetta. eLife 5, e17219 (2016).
Acknowledgements
We thank members of the Das laboratory for useful discussions; members of the Rosetta community for discussions and code sharing; M. Summers for permission to perform the HIV-1 DIS blind modeling challenge; V. Kosaraju and J. Nicol for helping to develop the Eterna3D interface; and Eterna participant J.R. for designing the eterna3D-JR_1 construct through the Eterna3D interface. Calculations were performed on the Stanford Sherlock cluster. Molecular graphics and analyses were performed with UCSF Chimera, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from NIH grant no. P41-GM103311. This work was supported by a Gabilan Stanford Graduate Fellowship (K.K.), the National Science Foundation (GRFP to K.K. and R.R.) and the National Institutes of Health (grant nos. P41GM103832, R01GM079429, P01AI120943, U54GM103297 and S10 OD021600 to W.C.; grant nos. R35 GM112579 and R21 AI145647 to R.D.).
Author information
Authors and Affiliations
Contributions
K.K., R.D. and W.C. conceptualized and designed the research. K.K. prepared the RNA samples for cryo-EM. K.Z., Z.S., S.L. and G.P. collected and analyzed the cryo-EM data. K.K. and V.V.T. collected and analyzed the M2-seq data. W.K. collected the mutate-map-rescue data. W.K. and K.K. analyzed the mutate-map-rescue data. K.K. developed, implemented and tested the computational approach with input from R.R., A.M.W. and R.D. K.K., A.M.W. and R.D. performed the blind DIS modeling. K.K. performed modeling for all other RNA systems. I.N.Z. prepared the 24-3 ribozyme RNA for cryo-EM and performed functional validation. J.D.Y. developed Eterna3D. A.M.W. developed the auto-DRRAFTER webserver. K.K. and R.D. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Arunima Singh and Allison Doerr were the primary editors on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Native gel screens for all RNAs in this study.
Gel images for (1) and (2) RB1 5′ UTR, (3) and (4) scaRNA6, (5) and (6) U1 snRNA, (7) and (8) SAM-IV riboswitch (apo), (9) and (10) 24-3, (11) V. cholerae glycine riboswitch (apo), (12) hc16, (13) Tetrahymena ribozyme, (14) F. nucleatum glycine riboswitch (apo), (15) and (16) Eterna3D-JR_1, (17) and (18) spinach-TTR-3, (19) F. nucleatum glycine riboswitch (apo), (20) ATP-TTR-3 (apo), (21) Eterna3D-JR_1, (22) SAM-IV riboswitch (apo), (23) downstream peptide riboswitch, (24) hc16, and (25) hc16 product. Samples 1-13, 24, 25 were run on an 8% polyacrylamide gel. Samples 14-23 were run on a 12% polyacrylamide gel. All samples were run in 10 mM MgCl2, 67 mM HEPES, 33 mM Tris, pH 7.2. All gels that were run are shown here (experiments were not repeated beyond results shown here).
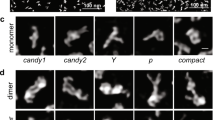
Extended Data Fig. 2 Representative micrographs for all RNAs in this study.
Micrographs shown in (a), (b), (c), (g), (h), (i), (j), (k), (l), (o), (q), and (r) were taken with the Talos Arctica. All others were taken with the Titan Krios. A Volta phase plate was used for (a), (b), (c), (g), (h), (i), (j), (k), (l), (o), and (q). The total numbers of micrographs collected are listed in Supplementary Table 1.
Extended Data Fig. 3 Example cryo-EM data processing workflow.
Shown here for the V. cholerae glycine riboswitch without glycine.
Extended Data Fig. 4 Cryo-EM 2D class averages and 3D reconstructions for all RNA systems in this study.
One dataset was collected for all RNAs except for the SAM-IV riboswitch without SAM and V. cholerae glycine riboswitch without glycine, for which smaller preliminary datasets were initially collected (see Methods). Results from these preliminary datasets were similar, though map resolution was lower. The numbers of particles used for the 3D reconstructions are listed in Supplementary Table 1.
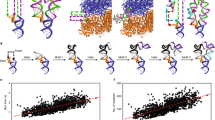
Extended Data Fig. 5 Local cryo-EM map resolution for RNA-only structures.
Calculated with ResMap48 for (a) Tetrahymena ribozyme, (b) hc16 product, (c) hc16, (d) V. cholerae glycine riboswitch with glycine, (e) V. cholerae glycine riboswitch without glycine, (f) F. nucleatum glycine riboswitch with glycine, (g) F. nucleatum glycine riboswitch without glycine, (h) ATP-TTR-3 with AMP, (i) ATP-TTR-3 without AMP, (j) SAM-IV riboswitch with SAM, and (k) SAM-IV riboswitch without SAM.
Extended Data Fig. 6 auto-DRRAFTER overview.
The F. nucleatum glycine riboswitch is shown here as an example. a, Secondary structure elements that connect to just one other helix or junction (‘end nodes’) are circled. b, The cryo-EM density map is low-pass filtered to 20 Å and points are placed (spheres) throughout the map to identify possible placements for ‘end nodes’ in the map (red spheres). The circled end node was randomly selected for initial helix placement. A probe helix (black) was then fit into the density map. The location of the probe helix was optimized, while the distances between the C1’ atom of nucleotide 6 (nucleotides 1-6 labeled) of the probe helix (black sphere) and the end node and the neighboring map node were monitored (see Supplementary Note 4). c, 3D models are built for each of the elements circled in (a) and fit into the density map in the location of the circled point in (b). These elements are kept fixed while the rest of the RNA is built into the density map. d, The top ten best scoring models after round 1. The overall convergence of these models is above the 10 Å threshold (convergence = 20.2 Å), so another round of modeling is performed. e, For the second round of modeling, regions that have converged are extracted from the top scoring models and kept fixed while the rest of the RNA is built into the density map. f, The top ten scoring models after round 2. The convergence is below the 10 Å threshold (convergence = 6.2 Å), so there is only one initial model for the third round of modeling, composed of converged regions from the top ten scoring models from round 2 g,. These regions are allowed to move from their initial positions during this modeling round. h, The best scoring models from round 3. i, Again, converged regions are extracted from top scoring models to form the initial model for the final round of modeling. These regions are kept fixed during the fragment assembly stage of auto-DRRAFTER modeling, but allowed to move during final refinement. j, The top ten scoring models built independently into each half map. Helical regions are depicted with bright colors in (c)-(j) and match colors in the secondary structure diagram (A). Non-helical regions are colored gray.
Extended Data Fig. 7 Experimental M2-seq z-score plots.
(a) Tetrahymena ribozyme (n = 65041 sequences), (b) hc16 product (n = 459852 sequences), (c) hc16 (n = 451568 sequences), (d) human scaRNA6 (n = 866158 seqeunces), (e) V. cholerae glycine riboswitch with glycine (n = 928767 sequences), (f) V. cholerae glycine riboswitch without glycine (n = 974343 sequences), (g) human RB1 5′ UTR (n = 889700 sequences), (h) 24-3 (n = 515486 sequences), (i) human U1 snRNA (n = 1185187), (j) F. nucleatum glycine riboswitch with glycine (n = 803254 sequences), (k) F. nucleatum glycine riboswitch without glycine (n = 670301 sequences), (l) eterna3D-JR_1 (n = 994048 sequences), (m) spinach-TTR-3 (n = 1090666 sequences), (n) ATP-TTR-3 with AMP (n = 914892 sequences), (o) ATP-TTR-3 without AMP (n = 712801 sequences), (p) SAM IV riboswitch with SAM (n = 131464 sequences), (q) SAM IV riboswitch without SAM (n = 991972 sequences), and (r) downstream peptide riboswitch (n = 1012081 sequences).
Extended Data Fig. 8 Secondary structures automatically derived from M2-seq data and revisions for best-case auto-DRRAFTER modeling based on sequence covariation and previously solved crystal structures.
(a) Tetrahymena ribozyme, (b) hc16 product, (c) hc16, (d) V. cholerae glycine riboswitch with glycine, (e) V. cholerae glycine riboswitch without glycine, (f) F. nucleatum glycine riboswitch with glycine, (g) F. nucleatum glycine riboswitch without glycine, (h) ATP-TTR-3 with AMP (I) ATP-TTR-3 without AMP, (j) SAM-IV riboswitch with SAM, and (k) SAM-IV riboswitch without SAM. (a-k) Blue lines indicate base pairs that are present in the best-case, but not automated secondary structures. Red lines indicate base pairs that are present in the automated, but not best-case secondary structures. (l) The previously proposed hc16 secondary structure4. (m) M2-seq and mutate-map-rescue experiments suggest that hc16 contains alt-P4 rather than P4. (n) Additional modeling and experiments suggest further modifications to the hc16 secondary structure: alt-P4 is extended, a pseudoknot is formed between the hairpin loops of P5c and P1, P5 is not formed, and P10 is formed. Helical regions are depicted with bright colors that match those shown in Fig. 3. Non-helical regions are colored black.
Extended Data Fig. 9 Secondary structures for best-case Ribosolve models.
(a) Tetrahymena ribozyme, (b) hc16 product, (c) hc16, (d) V. cholerae glycine riboswitch with glycine, (e) V. cholerae glycine riboswitch without glycine, (f) F. nucleatum glycine riboswitch with glycine, (g) F. nucleatum glycine riboswitch without glycine, (h) eterna3D-JR_1 (see Fig. 6), (i) spinach-TTR-3 (discussed in ‘Limitations of the Ribosolve pipeline’ in the main text), (j) ATP-TTR-3 with AMP (k) ATP-TTR-3 without AMP, (l) SAM-IV riboswitch with SAM, and (m) SAM-IV riboswitch without SAM. Helical regions are depicted with bright colors that match those shown in Fig. 3. Non-helical regions are colored black.
Extended Data Fig. 10 Benchmarking auto-DRRAFTER accuracy.
(a–h) The top ten scoring auto-DRRAFTER models (models #2-10 are transparent) built into 10 Å simulated density maps (left) and the corresponding crystal structures (right) for (a) THF riboswitch, (b) c-di-AMP riboswitch, (c) bacterial SRP Alu domain, (d) FMN riboswitch, (e) SAM-I riboswitch, (f) Tetrahymena ribozyme P4-P6 domain, (g) lysine riboswitch, and (h) lariat capping ribozyme. (i) Ribosolve models for THF riboswitch built into the previously solved 2.9 Å crystallographic density map (left) and the crystal structure (right). (j) Ribosolve models for the B. subtilis T-box-tRNA complex built into a 4.9 Å cryo-EM map (left) and previously modeled coordinates (right) based on high-resolution crystal structures30. Helical regions are depicted with bright colors. Non-helical regions are colored gray.
Supplementary information
Supplementary Information
Supplementary Results, Notes 1–7, Figs. 1–8, Tables 1–4, 6 and 7, and captions for Supplementary Table 5 and Supplementary Video 1.
Supplementary Table 5
DNA and RNA sequences used in this study.
Supplementary Video 1
RNA structures determined by the Ribosolve pipeline.
Supplementary Data
Auto-DRRAFTER models built using the fully automated procedure.
Rights and permissions
About this article
Cite this article
Kappel, K., Zhang, K., Su, Z. et al. Accelerated cryo-EM-guided determination of three-dimensional RNA-only structures. Nat Methods 17, 699–707 (2020). https://doi.org/10.1038/s41592-020-0878-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0878-9
This article is cited by
-
A deep learning-based method for modeling of RNA structures from cryo-EM maps
Nature Biotechnology (2024)
-
All-atom RNA structure determination from cryo-EM maps
Nature Biotechnology (2024)
-
Nano-DMS-MaP allows isoform-specific RNA structure determination
Nature Methods (2023)
-
Structure, folding and flexibility of co-transcriptional RNA origami
Nature Nanotechnology (2023)
-
CryoREAD provides fully automated DNA–RNA structure modeling for cryo-EM maps
Nature Methods (2023)