Abstract
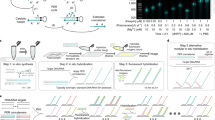
We present split-FISH, a multiplexed fluorescence in situ hybridization method that leverages a split-probe design to achieve enhanced specificity. Split-FISH reduces off-target background fluorescence, decreases false positives and enables accurate RNA profiling in uncleared tissues. We demonstrate the efficacy of split-FISH on various mouse tissues by quantifying the distribution and abundance of 317 genes in single cells and reveal diverse localization patterns for spatial regulation of the transcriptome in complex tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
RNA coordinates data are available at https://github.com/khchenLab/split-fish. All data obtained during this study are available from the corresponding author upon request. Source data are provided with this paper.
Code availability
The software, including the analysis pipeline, spots viewer, probe design and scripts to recreate the figures, can be downloaded from https://github.com/khchenLab/split-fish.
Change history
26 July 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41592-020-0931-8
References
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat. Methods 11, 360–361 (2014).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Codeluppi, S. et al. Spatial organization of the somatosensory cortex revealed by osmFISH. Nat. Methods 15, 932–935 (2018).
Mateo, L. J. et al. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 568, 49–54 (2019).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Xia, C., Fan, J., Emanuel, G., Hao, J. & Zhuang, X. Spatial transcriptome profiling by MERFISH reveals subcellular RNA compartmentalization and cell cycle-dependent gene expression. Proc. Natl Acad. Sci. USA 116, 201912459 (2019).
Wu, C. et al. RollFISH achieves robust quantification of single-molecule RNA biomarkers in paraffin-embedded tumor tissue samples. Commun. Biol. 1, 1–8 (2018).
Rouhanifard, S. H. et al. ClampFISH detects individual nucleic acid molecules using click chemistry–based amplification. Nat. Biotechnol. 37, 84–94 (2019).
Xia, C., Babcock, H. P., Moffitt, J. R. & Zhuang, X. Multiplexed detection of RNA using MERFISH and branched DNA amplification. Sci. Rep. 9, 1–13 (2019).
Kishi, J. Y. et al. SABER amplifies FISH: enhanced multiplexed imaging of RNA and DNA in cells and tissues. Nat. Methods 16, 533–544 (2019).
Choi, H. M. T. et al. Programmable in situ amplification for multiplexed imaging of mRNA expression. Nat. Biotechnol. 28, 1208–1212 (2010).
Moffitt, J. R. et al. High-performance multiplexed fluorescence in situ hybridization in culture and tissue with matrix imprinting and clearing. Proc. Natl Acad. Sci. USA 113, 201617699 (2016).
Wu, B., Chen, J. & Singer, R. H. Background free imaging of single mRNAs in live cells using split fluorescent proteins. Sci. Rep. 4, 3615 (2014).
Wang, F. et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagnostics 14, 22–29 (2012).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, 1–10 (2018).
Cabili, M. N. et al. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 16, 20 (2015).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 1–5 (2017).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 113, 201612826 (2016).
Lyubimova, A. et al. Single-molecule mRNA detection and counting in mammalian tissue. Nat. Protoc. 8, 1743–1758 (2013).
Farack, L. et al. Transcriptional heterogeneity of beta cells in the intact pancreas. Dev. Cell 48, 115–125 (2019).
Moffitt, J. R. & Zhuang, X. RNA imaging with multiplexed error-robust fluorescence in situ hybridization. Methods Enzymol. 572, 1–49 (2016).
Davis, C. A. et al. The Encyclopedia of DNA elements (ENCODE): data portal update. Nucleic Acids Res. 46, D794–D801 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Acknowledgements
We thank S. Prabhakar, M. Huang, J. Toh, C. Goh, V. Singhal, J. Liu, M. Lee and K. Mehta for insightful discussions and technical assistance. We thank the Advanced Molecular Pathology Laboratory at IMCB, A*STAR, Singapore for mouse necropsy and cryo-sectioning services. We thank the next-generation sequencing platform at GIS, A*STAR, Singapore for sequencing services. We thank Applied Materials for in-kind contributions. This research was supported by a National Medical Research Council of Singapore grant OFYIRG15nov017 (K.H.C.), Agency for Science, Technology, and Research grant I1801E0029 (K.H.C.) and an AXA Post-Doctoral Fellowship (K.H.C.). N.C. is grateful for support from A*STAR Graduate Academy. Z.W.Z. is grateful for a National Medical Research Council of Singapore grant OFYIRG18nov-0019 (Z.W.Z.) and National University of Singapore Presidential Young Professorship start-up fund (Z.W.Z.).
Author information
Authors and Affiliations
Contributions
K.H.C. conceived the study. K.H.C., J.J.L.G. and N.C. designed the experiments. K.H.C. and N.C. built the instruments. N.C. and K.H.C. developed the computational software and performed the image analysis. J.J.L.G. and K.H.C. performed the experiments. W.Y.S., N.H. and C.P.P.C. assisted with the experiments and replicated the results. K.H.C., J.J.L.G. and N.C. analyzed the data and interpreted the results. W.Y.S. and Y.-C.C. assisted with data analysis. Z.W.Z. assisted with building the instruments. K.H.C., N.C. and J.J.L.G. wrote the manuscript with input from Z.W.Z., W.Y.S. and N.H.
Corresponding author
Ethics declarations
Competing interests
The authors have submitted a provisional patent application that is based on the method described in this manuscript.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of the bridge sequence length.
(a) Split probes were designed to target a polymorphic repeat region of the MUC5AC transcripts in A549 cell lines. Sample RNA FISH images of split bridge sequence length (x) ranging from 7 to 12 nucleotides (nt) in (b) unpaired and (c) paired split probes (orange and light blue sequences). Shorter (7–9 nucleotides) bridge lengths were able to suppress the binding of unpaired probes. However, using bridge lengths that were too short (7 + 7 nucleotides) resulted in poor binding even in paired probes. 9 + 9 nucleotides appeared to be the most optimal length. This experiment was repeated once with similar results.
Extended Data Fig. 2 Optimization of the split probe construct.
(a–f) Six different constructs - circular, cruciform, double ‘C’, and double ‘Z’, conventional, and unpaired were tested. The target and probe sequences for MUC5AC are shown. (g–k) Sample RNA FISH images of the tested constructs with DAPI nucleus (blue) staining. We found that the circular construct (g) resulted in the best RNA signals, which achieved similar brightness to the conventional scheme (k). (l) In contrast, unpaired probe showed no signal (negative control). This set of experiments was repeated twice with similar results. (m) Box plots of the brightness of single RNA molecules (n = 1,000 randomly selected RNAs from 5 FOVs) for each of the probe constructs. Center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range.
Extended Data Fig. 3 Two channels co-localization control for the split probe construct.
(a) 75 unique probes (Cy3) against non-repeat regions on MUC5AC transcripts were simultaneously hybridized with split probe constructs (Cy5) - circular, cruciform, double ‘C’, double ‘Z’, and conventional. (b–e) Sample RNA FISH images from Cy3 and Cy5 channels for the circular and double ‘Z’ constructs, with DAPI staining (blue) for cell nucleus. Double ‘Z’ Cy5 is displayed at 4x enhanced contrast compared to ‘circular’. This experiment was repeated twice with similar results. (f) Box plots of the fraction of Cy5 spots that co-localized with Cy3 spots (n = 10 FOVs) for each of the probe constructs. Center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range.
Extended Data Fig. 4 Split probe construct eliminates false positive signals associated with known off targets.
(a) RNA FISH for XLOC_010514, for which one of the probes was known to have off-target binding in the nucleus17. Using conventional read-out, we also observed the false positive signals in the nucleus (blue). (b) Removal of the ‘rogue’ probe (red) eliminated the false positive signals in the nucleus. (c) RNA FISH using split readouts. No false positive signals were observed in the nucleus despite the presence of the ‘rogue’ sequence. (d) Readout probes are unable to bind to the unpaired ‘rogue’ sequence. This experiment was repeated once with similar results.
Extended Data Fig. 5 Optimization of split-FISH workflow.
Split-FISH image (a) with, and (b) without PCR amplification primers removed from the probes via restriction digestion. This was observed in 3 independent libraries. (c) Same as b, but at 10x contrast. (d) Normalized RNA brightness after hybridization of dye labelled bridge probe (blue) versus conventional readout probe (red) at 1, 5, 10, 30, and 60 min. This experiment was repeated once with similar results. An additional round of dye-labelled readout probe hybridization (~10 min) is needed for split-FISH (Fig. 2a).
Extended Data Fig. 6 Optimized split-FISH allows repeated cycles of hybridization and wash.
(a) Alternating hybridization and wash of the FLNA transcripts in the same A549 cells for 20 cycles. This experiment was repeated once with similar results. (b) Box plots of number of spots detected per cell (n = 38 cells) over the 20 hybridization and wash cycles. Center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range. (c) Box plots of RNA brightness (n = 1,000 randomly selected RNAs from 4 FOVs) over the 20 hybridizations. Center line, median; box limits, upper and lower quartiles; whiskers, 1.5x interquartile range.
Extended Data Fig. 7 Tissue auto-fluorescence was negligible compared to real RNA signals.
(a) Sample split-FISH image (same dataset as Fig. 1) overlaid with DAPI stain (blue). (b) Post-wash images, showing no detectable RNA spots. (c) Same image as b, but at 10x contrast, to highlight tissue auto-fluorescence and un-washed single fluorescent dye molecules. Similar results were observed in different tissues.
Extended Data Fig. 8 Correlation between total counts and bulk RNA-sequencing for conventional multiplexed FISH.
(a) AML12 (b) Liver (c) Brain (n = 1 section per tissue). Log Pearson correlation (r) values in red calculated using all the 133 genes.
Extended Data Fig. 9 Nuclear enrichment of gene expression in liver tissue.
Zoomed in high-resolution image of mouse liver tissue (same dataset as Fig. 3d) to show selected genes highly enriched in the nucleus: Son (red) and Abcc2 (yellow). DAPI image is overlaid in blue. Grayscale background image is the raw image from a single bit. Across the dataset (n = 1, all 49 FOVs, ~7000 cells), we estimated that Son and Abcc2 were enriched 5.6X and 6.4X respectively in the nucleus compared to outside the nucleus (see methods for details on segmentation of DAPI image).
Extended Data Fig. 10 Overview of the image analysis pipeline.
Step by step scheme: parsing of raw images, preprocessing (color channel correction, registration and filtering), normalization across bits, decoding of RNA identities, and stitching of FOVs into a complete mosaic.
Supplementary information
Supplementary Information
Supplementary Figures 1–3.
Supplementary Table 1
317-gene split-probe library template sequences. Template sequences include the forward and reverse primer sequences necessary for library amplification.
Supplementary Table 2
Codebook for each gene in the 317-gene split-probe library. The binary codeword assigned to each gene in the 317-gene split-probe library.
Supplementary Table 3
Bridge sequence for each bit in the 317-gene split-probe library. Each bridge sequence consists of three blocks (separated by spaces): an encoding probe binding the block in the center, flanked by two readout binding blocks. In the encoding-probe binding block, the barcode sequences are in lowercase. Bridge sequences used in AML12, kidney, frontal cortex and liver experiments are shown. B1 to B13 were read out by Alexa Fluor 750, and B14 to B26 were read out by Cy5. For ovary experiments, B1, B3, B8 to B13, B15 and B17 to B20 were read out by Cy5 and B2, B4 to B7, B14, B16 and B21 to B26 were read out by Alexa Fluor 750.
Supplementary Table 4
133-gene conventional library template sequences. Template sequences include the forward and reverse primer sequences necessary for library amplification. The primers used for PCR are 5′-TGGTTCAATCGTATGCCCGT-3′ and 5′-TAATACGACTCACTATAGGGGTCACTTAGCCAACGCCGAT-3′.
Supplementary Table 5
Readout probe sequences for each of the 16 bits used in the 133-gene conventional library.
Supplementary Table 6
Codebook for each gene in the 133-gene conventional library. The binary codeword assigned to each gene in the 133-gene conventional library.
Supplementary Table 7
Probe sequences for the conventional, split-probe pairs, and readout probe used in the XLOC_010514 experiment (Extended Data Fig. 4). The known off-target sequence is shown in red.
Supplementary Table 8
Probe sequences used in the MUC5AC experiment (Extended Data Figs. 1, 2, 3 and 5). Sheet 8a: Sequences of the unpaired, paired (circular) and readout probes used in Supplementary Fig. 1. Sheet 8b: Sequences of the MUC5AC split-probe constructs and readout probe used for Supplementary Fig. 2. Sheet 8c: Sequences of the MUC5AC colocalization and readout probes used for Supplementary Fig. 3. Sheet 8d: Sequences of the MUC5AC split-probe, conventional probe, bridge probe and readout probes used for the kinetic experiment in Supplementary Fig. 5. Lowercase letters denote the target gene (MUC5AC) binding sequence. Uppercase letters denote the three-nucleotide linker and readout binding sequence.
Supplementary Table 9
Probe sequences used in the FLNA experiment (Extended Data Fig. 6). The template sequence includes the forward and reverse primer sequences for amplifying the template sequence. The primers used for PCR amplification are 5′-TACCATCTCGTGTTCGTACC-3′ and 5′-TAATACGACTCACTATAGTTCGTTCCGCTACTCACCAC-3′.
Supplementary Table 10
Reference FKPM values for AML12, mouse kidney, liver, frontal cortex and ovary for the 317 genes in the split library.
Source data
Source Data Fig. 1
Raw image files for AML12 and brain tissue shown in Fig. 1.
Source Data Fig. 2
Raw gene counts and cell counts for AML12, mouse brain, liver, kidney and ovary datasets shown in Fig. 2.
Source Data Fig. 3
RNA coordinates for mouse brain, liver, kidney and ovary shown in Fig. 3.
Rights and permissions
About this article
Cite this article
Goh, J.J.L., Chou, N., Seow, W.Y. et al. Highly specific multiplexed RNA imaging in tissues with split-FISH. Nat Methods 17, 689–693 (2020). https://doi.org/10.1038/s41592-020-0858-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0858-0
This article is cited by
-
Highly sensitive spatial transcriptomics using FISHnCHIPs of multiple co-expressed genes
Nature Communications (2024)
-
SRTsim: spatial pattern preserving simulations for spatially resolved transcriptomics
Genome Biology (2023)
-
HT-smFISH: a cost-effective and flexible workflow for high-throughput single-molecule RNA imaging
Nature Protocols (2023)
-
The expanding vistas of spatial transcriptomics
Nature Biotechnology (2023)
-
Interpretable and context-free deconvolution of multi-scale whole transcriptomic data with UniCell deconvolve
Nature Communications (2023)