Abstract
CRISPR–Cas9 screens have emerged as a transformative approach to systematically probe gene functions. The quality and success of these screens depends on the frequencies of loss-of-function alleles, particularly in negative-selection screens widely applied for probing essential genes. Using optimized screening workflows, we performed essentialome screens in cancer cell lines and embryonic stem cells and achieved dropout efficiencies that could not be explained by common frameshift frequencies. We find that these superior effect sizes are mainly determined by the impact of in-frame mutations on protein function, which can be predicted based on amino acid composition and conservation. We integrate protein features into a ‘Bioscore’ and fuse it with improved predictors of single-guide RNA activity and indel formation to establish a score that captures all relevant processes in CRISPR–Cas9 mutagenesis. This Vienna Bioactivity CRISPR score (www.vbc-score.org) outperforms previous prediction tools and enables the selection of sgRNAs that effectively produce loss-of-function alleles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
All code used for this study including a python script to predict VBC score and Bioscore from feature data is openly available at https://github.com/GMichlits/VBC-Score. An online tool for sgRNA predictions is available at www.vbc-score.org.
References
Koike-Yusa, H., Li, Y., Tan, E.-P., Velasco-Herrera, M. D. C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2013).
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Shi, J. et al. Discovery of cancer drug targets by CRISPR–Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015).
Tzelepis, K. et al. A CRISPR dropout screen identifies genetic vulnerabilities and therapeutic targets in acute myeloid leukemia. Cell Rep. 17, 1193–1205 (2016).
Munoz, D. M. et al. CRISPR screens provide a comprehensive assessment of cancer vulnerabilities but generate false-positive hits for highly amplified genomic regions. Cancer Disco. 6, 900–913 (2016).
Aguirre, A. J. et al. Genomic copy number dictates a gene-independent cell response to CRISPR/Cas9 targeting. Cancer Disco. 6, 914–929 (2016).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Wang, T. et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic ras. Cell 168, 890–903.e15 (2017).
Sanson, K. R. et al. Optimized libraries for CRISPR–Cas9 genetic screens with multiple modalities. Nat. Commun. 9, 5416 (2018).
Steinhart, Z. et al. Genome-wide CRISPR screens reveal a Wnt-FZD5 signaling circuit as a druggable vulnerability of RNF43-mutant pancreatic tumors. Nat. Med. 23, 60–68 (2017).
Martin, T. D. et al. A role for mitochondrial translation in promotion of viability in K-Ras mutant cells. Cell Rep. 20, 427–438 (2017).
Behan, F. M. et al. Prioritization of cancer therapeutic targets using CRISPR–Cas9 screens. Nature 568, 511–516 (2019).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. Genes Genome. Genet. 7, 2719–2727 (2017).
Hart, T., Brown, K. R., Sircoulomb, F., Rottapel, R. & Moffat, J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol. Syst. Biol. 10, 733–733 (2014).
van Overbeek, M. et al. DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol. Cell 63, 633–646 (2016).
Allen, F. et al. Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat. Biotechnol. 498, 349 (2018).
Shen, M. W. et al. Predictable and precise template-free CRISPR editing of pathogenic variants. Nature 563, 646–651 (2018).
Chakrabarti, A. M. et al. Target-specific precision of CRISPR-mediated genome editing. Mol. Cell 73, 699–713 (2018).
Smits, A. H. et al. Biological plasticity rescues target activity in CRISPR knock outs. Nat. Methods 346, 1258096–1258097 (2019).
Anderson, J. L. et al. mRNA processing in mutant zebrafish lines generated by chemical and CRISPR-mediated mutagenesis produces unexpected transcripts that escape nonsense-mediated decay. PLoS Genet. 13, e1007105 (2017).
Mou, H. et al. CRISPR/Cas9-mediated genome editing induces exon skipping by alternative splicing or exon deletion. Genome Biol. 18, 108–108 (2017).
Tuladhar, R. et al. CRISPR–Cas9-based mutagenesis frequently provokes on-target mRNA misregulation. Nat. Commun. 10, 4010–4056 (2019).
Schoonenberg, V. A. C. et al. CRISPRO: identification of functional protein coding sequences based on genome editing dense mutagenesis. Genome Biol. 19, 169 (2018).
Tarumoto, Y. et al. LKB1, Salt-inducible kinases, and MEF2C are linked dependencies in acute myeloid leukemia. Mol. Cell 69, 1017–1027.e6 (2018).
Michlits, G. et al. CRISPR-UMI: single-cell lineage tracing of pooled CRISPR–Cas9 screens. Nat. Methods 14, 1191–1197 (2017).
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014).
Andersson, B. S. et al. KBM-7, a human myeloid leukemia cell line with double Philadelphia chromosomes lacking normal c-ABL and BCR transcripts. Leukemia 9, 2100–2108 (1995).
Kotecki, M., Reddy, P. S. & Cochran, B. H. Isolation and characterization of a near-haploid human cell line. Exp. Cell Res. 252, 273–280 (1999).
Doench, J. G. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Elling, U. et al. Forward and reverse genetics through derivation of haploid mouse embryonic stem cells. Cell. Stem Cell 9, 563–574 (2011).
Elling, U. et al. A reversible haploid mouse embryonic stem cell biobank resource for functional genomics. Nature 550, 114–118 (2017).
Allen, F. et al. Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat. Biotechnol. 37, 64–72 (2019).
He, W. et al. De novo identification of essential protein domains from CRISPR–Cas9 tiling-sgRNA knockout screens. Nat. Commun. 10, 4510–4541 (2019).
Blanchette, M. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Ma, J. et al. CRISPR-DO for genome-wide CRISPR design and optimization. Bioinformatics 32, 3336–3338 (2016).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Sim, N.-L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–W457 (2012).
Chari, R., Yeo, N. C., Chavez, A. & Church, G. M. sgRNA Scorer 2.0: a species-independent model to predict CRISPR/Cas9 activity. ACS Synth. Biol. 6, 902–904 (2017).
Labuhn, M. et al. Refined sgRNA efficacy prediction improves large- and small-scale CRISPR-Cas9 applications. Nucleic Acids Res. 46, 1375–1385 (2018).
Chuai, G. et al. DeepCRISPR: optimized CRISPR guide RNA design by deep learning. Genome Biol. 19, 18–80 (2018).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat. Genet. 25, 25–29 (2000).
Michlits, G., Burkard, T. R., Novatchkova, M. & Elling, U. CRISPR-UMI step by step: a protocol for robust CRISPR screening. Protoc. Exch. https://doi.org/10.1038/protex.2017.111 (2017).
Capra, J. A. & Singh, M. Predicting functionally important residues from sequence conservation. Bioinformatics 23, 1875–1882 (2007).
Rauscher, B., Heigwer, F., Breinig, M., Winter, J. & Boutros, M. GenomeCRISPR—a database for high-throughput CRISPR/Cas9 screens. Nucleic Acids Res. 45, D679–D686 (2017).
Finn, R. D. et al. The pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Rosenbloom, K. R. et al. The UCSC genome browser database: 2015 update. Nucleic Acids Res. 43, D670–D681 (2015).
Acknowledgements
We thank all members of our laboratories for constructive discussions and M. Weißenböck, T. Hoffmann and O. Kaya for experimental support. We are grateful to A. Stark and J. Brennecke for advice throughout this study, and to A. Andersen (Life Science Editors) for manuscript editing. We thank M. Schuler, M. Lenter, D. Mennerich and B. Ingelheim for support with library generation, A. Sommer and his team at the Vienna BioCenter Core Facilities (VBCF) (www.vbcf.ac.at) for NGS services, IMBA/IMP services for continuous support and the VBCF Information Management team and the IMBA/IMP IT team for help with building the website. We thank D. Hoepfner for continuous helpful discussions and I. Kaminski (kaminskigrafik.ch) for logo design. This work was funded by the European Research Council (ERC-StG-336860 to J.Z.) and the Austrian Science Fund (SFB-F4710 to J.Z.). Research at IMP/IMBA is generously supported by Boehringer Ingelheim, the Austrian Research Promotion Agency (Headquarter grant FFG-852936) and the Austrian Academy of Sciences.
Author information
Authors and Affiliations
Contributions
G.M. performed screens, indel mapping and all other experiments in mESCs with support from M.Hu and M.G., and performed bioinformatic analyses with support from U.E., A.S., T.R.B., G.V. and T.N. J.Z., J.J. and T.N. designed sgRNA libraries. J.J. devised protocols for sgRNA library cloning and screening with support from M.Hi and constructed genome-wide sgRNA libraries with support from M.F. J.J., M.Hi, M.dA. and A.T. derived Cas9 clones and performed screens in human cell lines. G.M. and U.E. conceived and developed the Vienna Bioactivity CRISPR score with support from J.Z. and T.N. U.E., J.Z. and G.M. cowrote the manuscript with input from all coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Lei Tang was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Viability screens performance comparisons.
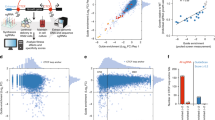
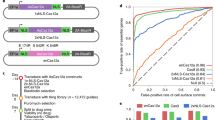
a, Comparative analysis of high-performance Vienna sgRNA library-based screen versus published11 screen in diploid RKO cells. b-e, Depletion of sgRNAs targeting core essential genes (red) and other genes (grey) on the x-axis, sgRNA representation in the plasmid library is shown on the y-axis in a screen in b, haploid mESCs using the Vienna Ms library, c, diploid mESCs using the Vienna Ms library, d, haploid mESCs using the UMI Ms library, e, haploid mESCs using the UMI Ms library. Diagonal lines of dropout on the left edge of the plot show sgRNAs depleting to 0 reads in the sample. f, Quality parameter for viability screens, dAUC (delta area under the curve) essential gene recall. Bars represent different libraries, data points individual screens with that library.
Extended Data Fig. 2 Clonal Cas9 expressing cell lines and fitness screens in diploid and haploid mouse embryonic stem cells.
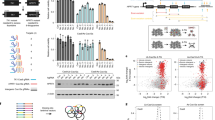
a, KO efficiency and translated log2 fold change of GFP loss in 96 single-cell-derived Cas9-expressing clones (diamonds) and in the bulk population (bar). Bar show mean with s.d. n = 96 individually picked single cell derived clonal populations with presumably independent Cas9 integration sites. b, Same experiment as in a using a different Cas9 vector (Cas9-P2A-BlastiR) and a different mouse embryonic cell line. Left panel: GFP KO Efficiency and corresponding log 2-fold change when infecting with sgRNA against ROSA (grey) or sgRNA against GFP in a standard tracr (tracrV1) backbone (green) or enhanced tracr (tracrV2) in (pink) after 2 weeks, ‘clones’ shows data of 94 single cell derived clones, ‘pool’ is an experiment without picking subclones carried out in triplicate, shown are 6 data points corresponding to technical duplicates of 3 pools of cells independently infected with Cas9-P2A-Blasti. Right panel: Experiment as in left panel. GFP KO efficiency after 1 week, lines connect identical clones. c, Depletion kinetics of experiment shown in a over 3 weeks. d, Effect of increasing selection antibiotic concentration from standard 10ug/ml to 50ug/ml in polyclonal Cas9 cell lines. Depletion kinetics over 3 weeks. shown are 6 data points corresponding to technical duplicates of 3 pools of cells independently infected with Cas9-P2A-BlastiR. b, c and d all bars show mean. e, Model of differential depletion of genes in diploid versus haploid cells. For each dot, representing a sgRNA, frameshift rate is 60–99% (purple and green), 100% (grey). Dominant mutations or haploinsufficient genes are shown in purple, recessive mutations in green. f, Depletion of genes in genome-wide viability screen in isogenic diploid and haploid mouse embryonic stem cells carried out with the Vienna sgRNA library. Most differentially depleting genes are shown in purple and green. g, Same experiment as in f, using an independent library containing mostly nuclear genes (UMI-library) Genes among the top 100 most diploid or top 100 most haploid depleting genes in the Vienna Ms library are highlighted if the same Gene was also present in the UMI-library. h, Shown are the Genes from the top 300 most diploid or top 300 most haploid depleting genes in the Vienna Ms library that were also present in the UMI library. The orthogonal distance of those Genes in the UMI screen to the linear regression trendline (dotted line in c) is plotted. For weak to medium depleting genes with LFC > −5 there is a separation of most diploid and most haploid genes also in the UMI-library (p-value 3.7e-6) reproducing the separation observed in the Vienna Ms library. This is not true for strongly depleting genes with LFC < −5 (p-value 0.4) p-values are heteroscedastic two-sided t-test. centre is mean error bars show s.d.
Extended Data Fig. 3 Analysis of indel fitness for individual sgRNAs.
Showing individual sgRNAs in addition to pooled results shown in Fig. 3e and 3f. Quantification of individual mutations of individual sgRNAs in a, control, b, weak, c, medium and d, strong sgRNAs y-axis fraction relative to wt sequence. Positive bars show quantification of individual mutations at day2, negative bars show quantification at day18. The x-axis indicates the 15 most frequently found mutations for each sgRNA. Error bars are standard deviation of 3 separately infected experiments. Annotations: d – deletion, i – insertion, d:6:−2 indicates a deletion of 6bp starting 2 bp upstream of the expected cut site. bars show median error bars s.d.
Extended Data Fig. 4 Conservation score optimization.
Predictive power (R-score) of individual and combined features using linear regression to predict strongly depleting sgRNA within essential genes. Individual Parameter optimization for a, various classes of Pfam annotations, b, window size of amino acids around CRISPR cut site to predict conservation of domains. Bars represent the median R-Score on the test set error bars standard deviation n = 9. c, Optimization of species sets available for human alignment used for generation of amino acid conservation score. d, Optimization of window size for phylogenetic conservation on DNA level by phylo scores and e, phastCons scores. Error bars show standard deviation of one data set 9 times independently split into training and testing sets. f, Extension to Fig 3b. showing multiple examples of essential genes targeted with a tiled sgRNA library illustrate correlation between depletion rate and amino acid conservation. Every circle represents a sgRNA. The x-axis represents the 5’ to 3’ position along the CDS coding DNA sequence and correlates with the gene body depicted in black, Pfam annotated regions are marked with a wide black box. AA conservation 1 is highly conserved 0 is not conserved. a-e n = 9 bars are mean error bars are s.d.
Extended Data Fig. 5 Examples for AA-identity score correlation with tiled data set.
Extension to data shown in Fig 3d) showing the dropout of tiled sgRNAs targeting multiple genes with a color code for AA identity scores derived from amino acid biases shown in Fig 3d). sgRNAs targeting regions with high AA identity scores (mostly hydrophobic amino acids) in red show a tendency to higher depletion (y-axis) than sgRNAs targeting regions with low AA identity scores.
Extended Data Fig. 6 Amino acid identity scoring matrix across 6 libraries.
Extended data to Fig 3c and d a, Scoring matrix for amino acid identity around the CRISPR cutsite. Shown are data for screens with 6 different libraries using tracr_v2 backbone. b, linear regression model Predictive power (R-score) of various amino acids in a variable window size around the cutsite. The optimal window size was determined to be 13 aa. Bars show mean with s.d. of one data set n = 9 times run each time independently split into training and testing sets. For the VBC score derivation we did not use any data from Vienna human library.
Extended Data Fig. 7 Amino acid identity score in sense and antisense and independence to nucleotide-sequence based scores.
a, Extended data to Fig 3c and 3d. Amino acid identity score calculated separately for screens from different tracr_v2 libraries that target antisense or sense to the gene orientation. b, Predictive power (R-score) of individual and combined features using linear regression to predict strongly depleting sgRNA within essential genes in the tiled gene dataset9. In grey nucleotide-based scores Doench 2016 and TracrV2, in light blue the AA-identity score. The red box (81%) markes the additive improvement in R-score that is gained by the AA-identity score over nucleotide scores deduced from panel c) c, n-1 combination of individual features as in b). Reduced combined predictive power when leaving out AA-identiy score shows independence of AA-identiy score from nucleotide scores. Red box marks the fraction of predictive power of the AA-identity score that remains even in presence of sgRNA restrictions due to nucleotide matrices. b and c bars are mean with s.d.
Extended Data Fig. 8 Gene architecture components, independence of Bioscore features and residual RNA and protein in frameshift clones.
a, Predictive power (R-score) of individual and combined features using linear regression to predict strongly depleting sgRNA within essential genes in a test set of tiled gene datasets Munoz 201623. Individual parameters for feature ‘gene structure’. n = 9 runs bars are mean with s.d. b and c Re-analyzed data from Smits et al.23. Every dot represents a KO-clone containing a frameshift mutation. b, Residual RNA level c, residual protein level is shown in dependence of positon along the transcript. Consistenly targeting genes within the first 5% of the coding DNA Sequence resulted in significantly higher number of clones with residual RNA and protein. b and c left panel n = 150, right panel bin1 n = 24, bin2 n = 55, bin3 n = 55, bin4 n = 16. p-values are heteroscedastic two-sided t-test. bars are mean with s.d. d, Predictive value of the Bioscore leaving out one feature category at a time. n = 9 runs bars are mean with s.d.
Extended Data Fig. 9 VBC-score performance on tiled gene depletion data, dAUC screen quality parameter and Scorer performance based on ddAUC.
a, Additional examples as in Fig. 3i. Shown is the performance of the VBC score on its training data set Munoz et al.9. Every dot marks an sgRNA along the gene body on the x-axis, versus log2 fold change depletion on the y-axis. Colored dots represent VBC sgRNA score. b, Illustration of dAUC parameter example cell line AML2 (from Behan and Iorio et al.16) The difference between red curve (non-essential gene set18) and green curve (core essential gene set17) quantifies the core essential gene recall. c, ddAUC is the difference in dAUC Parameter of the better and worse half of sgRNAs (defined by different sgRNA scoring algorithms). The parameter is normalized to the separation ddAUC that would be achieved by a theoretically perfect sgRNA score (defining the better half of sgRNA as those which show more depletion). Every dot represents a different tracrV2 library including Munoz 2016 (genome wide library, distinct library from tiled training dataset)9, Behan and Iorio et al.16, Tzelepis et al.8, Sanson et al.13 (Brunello library), Vienna Hm library, Vienna Ms library and CRISPR-UMI library Michlits 201729.
Extended Data Fig. 10 Correlation of screen performance versus predictive power of Bioscore and sgRNA activity score. Forward validation of top 2 VBC-score sgRNAs versus top 2 Doench2016 sgRNAs.
a, Every point represents a CRISPR-screen with tracrV1 or tracrV2 backbone. The y-axis shows the Pearson R correlation between sgRNA Bioscore and absolute sgRNA LFC for all depleting genes (that is predictive power of Bioscore within that screen). The x-axis shows the median LFC for all genes of the core essential genes (CEG) defined by Hart et al.17 b, as a, showing correlation with combined sgRNA activity score and median CEG depletion. c, Forward validation. For every tested gene the top 2 VBC-Score sgRNA and the top 2 Doench 2016-Score sgRNAs were selected. Genes are grouped into 3 classes CEG (core essential genes – positive controls), NEG (non-essential genes – negative controls), and test-Genes (359 Genes previously identified to have moderate median depletion with at least one sgRNA showing strong depletion). The sgRNAs were grouped to compare best VBC-score sgRNA with best Doench2016-score sgRNA and second best VBC-Score sgRNA with second best Doench2016-score sgRNA. For pairs with different sgRNAs the ratio of sgRNAs with superior depletion of the VBC-score sgRNA is shown. d, Forward validation of sgRNA dropout of top VBC-Score sgRNAs versus top Doench2016 sgRNAs. Every dot represents a sgRNA targeting CEG (core essential genes) n = 28, NEG (non-essential genes) n = 90, test-gene set of 359 previously weakly depleting genes with 2 sgRNAs per gene n = 718. p-values are heteroscedastic two-sided t-test. center line is mean with s.d.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 and Note 1.
Supplementary Table 1
Vienna_Hm_screens_sgRNAs; contains sgRNAs and raw read counts for fitness screens in KBM7, MiaPaca2 and RKO cells.
Supplementary Table 2
Vienna_Ms_screens_sgRNAs; contains sgRNAs and raw data for haploid and diploid mESC fitness screens. Using the genome-wide Vienna_ms_library.
Supplementary Table 3
CRISPR-UMI_mESC_screens; contains sgRNAs and raw data for haploid and diploid mESC fitness screens. Using mostly the nuclear CRISPR-UMI library.
Supplementary Table 4
Indel_Mapping_sgRNAs_PCR primers; contains a list of PCR primers used for indel mapping experiments.
Supplementary Table 5
sgRNA_nt_distributions_libraries; contains tables with the frequency of each basepair at each sgRNA position as they were used in various published libraries.
Supplementary Table 6
Species_for_AA_cons_align; contains a list of species that were used for alignments to derive a.a.-based conservation scores.
Supplementary Table 7
Benchmarking_detailed_results contains results of scorer performance comparisons using three different benchmarking methods.
Supplementary Table 8
Libraries_sgRNA_Scorers; contains lists of sgRNAs used in various libraries with sgRNA properties.
Supplementary Table 9
FWvalidation_sgRNAs_read.
Supplementary Table 10
Mean bioscore for GO terms; lists the mean bioscore of all genes present in a GO term.
Supplementary Table 11
hg38_all_sgRNAs list of >4 million human sgRNAs with properties.
Supplementary Table 12
mm10_all_sgRNAs; list of >4 million mouse sgRNAs with properties.
Supplementary Table 13
hg38_top6_sgRNAs; list of top 6 sgRNAs per gene for human genes.
Supplementary Table 14
mm10_top6_sgRNAs; list of top 6 sgRNAs per gene for mouse genes.
Source data
Rights and permissions
About this article
Cite this article
Michlits, G., Jude, J., Hinterndorfer, M. et al. Multilayered VBC score predicts sgRNAs that efficiently generate loss-of-function alleles. Nat Methods 17, 708–716 (2020). https://doi.org/10.1038/s41592-020-0850-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0850-8
This article is cited by
-
ABCC1 and glutathione metabolism limit the efficacy of BCL-2 inhibitors in acute myeloid leukemia
Nature Communications (2023)
-
Single-cell brain organoid screening identifies developmental defects in autism
Nature (2023)
-
Discovery of a Drug-like, Natural Product-Inspired DCAF11 Ligand Chemotype
Nature Communications (2023)
-
Efficient combinatorial targeting of RNA transcripts in single cells with Cas13 RNA Perturb-seq
Nature Methods (2023)
-
STING inhibits the reactivation of dormant metastasis in lung adenocarcinoma
Nature (2023)