Abstract
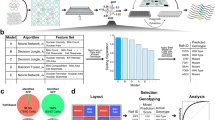
Genetic screens using pooled CRISPR-based approaches are scalable and inexpensive, but restricted to standard readouts, including survival, proliferation and sortable markers. However, many biologically relevant cell states involve cellular and subcellular changes that are only accessible by microscopic visualization, and are currently impossible to screen with pooled methods. Here we combine pooled CRISPR–Cas9 screening with microraft array technology and high-content imaging to screen image-based phenotypes (CRaft-ID; CRISPR-based microRaft followed by guide RNA identification). By isolating microrafts that contain genetic clones harboring individual guide RNAs (gRNA), we identify RNA-binding proteins (RBPs) that influence the formation of stress granules, the punctate protein–RNA assemblies that form during stress. To automate hit identification, we developed a machine-learning model trained on nuclear morphology to remove unhealthy cells or imaging artifacts. In doing so, we identified and validated previously uncharacterized RBPs that modulate stress granule abundance, highlighting the applicability of our approach to facilitate image-based pooled CRISPR screens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Sequencing data available under GEO accession GSE139815. RBP CRISPR plasmid library is available on Addgene (141438). Protein–protein interaction data used in this study are curated from Mentha (v.2018-01-08) (https://mentha.uniroma2.it/doDownload.php?file=2018-01-08.zip) and BioPlex v.2.0 (https://bioplex.hms.harvard.edu/data/BioPlex_interactionList_v2.tsv). Any additional data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
CRaft-ID software available at https://github.com/YeoLab/CRaftID.
References
Shalem, O. et al. Genome-scale CRISPR–cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
DeJesus, R. et al. Functional CRISPR screening identifies the ufmylation pathway as a regulator of SQSTM1/p62. eLife 5, e17290 (2016).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-Seq. Cell 167, 1883–1896.e15 (2016).
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882.e21 (2016).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866.e17 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Link, W. et al. Chemical interrogation of FOXO3a nuclear translocation identifies potent and selective inhibitors of phosphoinositide 3-kinases. J. Biol. Chem. 284, 28392–28400 (2009).
de Groot, R., Luthi, J., Lindsay, H., Holtackers, R. & Pelkmans, L. Large-scale image-based profiling of single-cell phenotypes in arrayed CRISPR-Cas9 gene perturbation screens. Mol. Syst. Biol. 14, e8064 (2018).
Maharana, S. et al. RNA buffers the phase separation behavior of prion-like RNA binding proteins. Science 360, 918–921 (2018).
Caicedo, J. C., Singh, S. & Carpenter, A. E. Applications in image-based profiling of perturbations. Curr. Opin. Biotechnol. 39, 134–142 (2016).
Kiger, A. A. et al. A functional genomic analysis of cell morphology using RNA interference. J. Biol. 2, 27 (2003).
Liu, T., Sims, D. & Baum, B. Parallel RNAi screens across different cell lines identify generic and cell type-specific regulators of actin organization and cell morphology. Genome Biol. 10, R26 (2009).
Feldman, D. et al. Optical pooled screens in human cells. Cell 179, 787–799.e17 (2019).
Camsund, D. et al. Time-resolved imaging-based CRISPRi screening. Nat. Methods 17, 86–92 (2020).
Wang, C., Lu, T., Emanuel, G., Babcock, H. P. & Zhuang, X. Imaging-based pooled CRISPR screening reveals regulators of lncRNA localization. Proc. Natl Acad. Sci. USA 116, 10842–10851 (2019).
Wang, Y. et al. Micromolded arrays for separation of adherent cells. Lab Chip 10, 2917–2924 (2010).
DiSalvo, M., Smiddy, N. M. & Allbritton, N. L. Automated sensing and splitting of stem cell colonies on microraft arrays. APL Bioeng. 3, 036106 (2019).
Gach, P. C., Wang, Y., Phillips, C., Sims, C. E. & Allbritton, N. L. Isolation and manipulation of living adherent cells by micromolded magnetic rafts. Biomicrofluidics 5, 032002 (2011).
Kedersha, N. & Anderson, P. Mammalian stress granules and processing bodies. Methods Enzymol. 431, 61–81 (2007).
Anderson, P., Kedersha, N. & Ivanov, P. Stress granules, P-bodies and cancer. Biochim. Biophys. Acta 1849, 861–870 (2015).
Grabocka, E. & Bar-Sagi, D. Mutant KRAS enhances tumor cell fitness by upregulating stress granules. Cell 167, 1803–1813.e12 (2016).
Wolozin, B. & Ivanov, P. Stress granules and neurodegeneration. Nat. Rev. Neurosci. 20, 649–666 (2019).
Murakami, T. et al. ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88, 678–690 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Boeynaems, S. et al. Drosophila screen connects nuclear transport genes to DPR pathology in c9ALS/FTD. Sci. Rep. 6, 20877 (2016).
Lee, K. H. et al. C9orf72 dipeptide repeats impair the assembly, dynamics, and function of membrane-less organelles. Cell 167, 774–788.e17 (2016).
Lin, Y. et al. Toxic PR poly-dipeptides encoded by the C9orf72 repeat expansion target LC domain polymers. Cell 167, 789–802.e12 (2016).
Martinez, F. J. et al. Protein-RNA networks regulated by normal and ALS-associated mutant HNRNPA2B1 in the nervous system. Neuron 92, 780–795 (2016).
Mackenzie, I. R. et al. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron 95, 808–816.e9 (2017).
Fang, M. Y. et al. Small-molecule modulation of TDP-43 recruitment to stress granules prevents persistent TDP-43 accumulation in ALS/FTD. Neuron 103, 802–819.e11 (2019).
Markmiller, S. et al. Context-dependent and disease-specific diversity in protein interactions within stress granules. Cell 172, 590–604.e13 (2018).
Jain, S. et al. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell 164, 487–498 (2016).
Youn, J. Y. et al. High-density proximity mapping reveals the subcellular organization of mRNS-associated granules and bodies. Mol. Cell 69, 517–532.e11 (2018).
McEwen, E. et al. Heme-regulated inhibitor kinase-mediated phosphorylation of eukaryotic translation initiation factor 2 inhibits translation, induces stress granule formation, and mediates survival upon arsenite exposure. J. Biol. Chem. 280, 16925–16933 (2005).
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. G3 7, 2719–2727 (2017).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Schneider-Poetsch, T. et al. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nat. Chem. Biol. 6, 209–217 (2010).
Tourriere, H. et al. The RasGAP-associated endoribonuclease G3BP assembles stress granules. J. Cell Biol. 160, 823–831 (2003).
Ohn, T., Kedersha, N., Hickman, T., Tisdale, S. & Anderson, P. A functional RNAi screen links O-GlcNAc modification of ribosomal proteins to stress granule and processing body assembly. Nat. Cell Biol. 10, 1224–1231 (2008).
Low, K. J. et al. PUF60 variants cause a syndrome of ID, short stature, microcephaly, coloboma, craniofacial, cardiac, renal and spinal features. Eur. J. Hum. Genet 25, 552–559 (2017).
Zhang, X. et al. Contribution of SNRNP200 sequence variations to retinitis pigmentosa. Eye 27, 1204–1213 (2013).
Handrigan, G. R. et al. Deletions in 16q24.2 are associated with autism spectrum disorder, intellectual disability and congenital renal malformation. J. Med. Genet. 50, 163–173 (2013).
Chung, J. et al. Genome-wide association study of cerebral small vessel disease reveals established and novel loci. Brain 142, 3176–3189 (2019).
Cirillo, L. et al. UBAP2L forms distinct cores that act in nucleating stress granules upstream of G3BP1. Curr. Biol. 30, 698–707.e96 (2020).
Huttlin, E. L. et al. The bioplex network: a systematic exploration of the human interactome. Cell 162, 425–440 (2015).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Tan, F. E. et al. A transcriptome-wide translational program defined by LIN28B expression level. Mol. Cell 73, 304–313.e3 (2019).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
DiSalvo, M. et al. Characterization of tensioned PDMS membranes for imaging cytometry on microraft arrays. Anal. Chem. 90, 4792–4800 (2018).
Cao, Q. et al. CRISPR-FOCUS: a web server for designing focused CRISPR screening experiments. PLoS ONE 12, e0184281 (2017).
Attayek, P. J. et al. Array-based platform to select, release, and capture Epstein–Barr virus-infected cells based on intercellular adhesion. Anal. Chem. 87, 12281–12289 (2015).
Baltz, A. G. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 46, 674–690 (2012).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Castello, A. et al. Comprehensive identification of RNA-binding domains in human cells. Mol. Cell 63, 696–710 (2016).
Beckmann, B. M. et al. The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs. Nat. Commun. 6, 10127 (2015).
Conrad, T. et al. Serial interactome capture of the human cell nucleus. Nat. Commun. 7, 11212 (2016).
Sundararaman, B. et al. Resources for the comprehensive discovery of functional RNA elements. Mol. Cell 61, 903–913 (2016).
Trendel, J. et al. The human RNA-binding proteome and its dynamics during translational arrest. Cell 176, 391–403.e19 (2019).
Queiroz, R. M. L. et al. Comprehensive identification of RNA–protein interactions in any organism using orthogonal organic phase separation (OOPS). Nat. Biotechnol. 37, 169–178 (2019).
Acknowledgements
We thank S. Gebhart and N. Trotta from Cell Microsystems for extensive consultation and troubleshooting support on this project. We thank Yeo laboratory members S. Markmiller for the HEK293T-G3BP1-GFP cell line and F. Tan for the PiggyBAC shuttle vectors. We acknowledge Yeo laboratory members S. Markmiller, M. Perelis, J. Nussbacher, A. Smargon, M. Corley and E. Boyle for critical reading of the manuscript. We thank the members of the Nikon Imaging Center at UC San Diego for help with imaging experiments. E.C.W. and A.Q.V were supported by the National Science Foundation Graduate Research Fellowship. E.C.W. and N.A. were supported in part by a Ruth L. Kirschstein Institutional National Research Award from the National Institute for General Medical Sciences, T32 GM008666. J.M.E. is supported by the Ruth L. Kirschstein F31 National Research Service Award (F31 CA217173) and Cancer Systems Biology Training Program (P50 GM085764 and U54 CA209891). M.D. is supported by the Ruth L. Kirschstein F31 National Research Service Award (F31 CA206233). E.L.V. is supported by the National Human Genome Research Institute (K99HG009530). This work is partially supported by NIH grants HG004659 and NS103172 to G.W.Y and NIH grant EY024556 to N.L.A.
Author information
Authors and Affiliations
Contributions
E.C.W., A.Q.V. and G.W.Y conceptualized the project. E.L.V. designed the CRISPR library. J.M.E. cloned the CRISPR library and performed viral infections. A.Q.V. optimized cell plating on microraft arrays. E.C.W. wrote analysis software and performed targeted library preparation. M.D. assisted with confocal imaging and fabricated microraft arrays. A.A.S. and E.L.V. designed the bulk CRISPR library preparation method. N.A. and A.Q.V. implemented neural network analysis. W.J. performed PPI analyses. A.Q.V. and E.C.W. performed validation experiments. E.C.W., A.Q.V. and G.W.Y. wrote the manuscript. N.L.A. and G.W.Y supervised the project.
Corresponding author
Ethics declarations
Competing interests
G.W.Y. is co-founder, member of the Board of Directors, on the SAB, equity holder and paid consultant for Locana and Eclipse BioInnovations. G.W.Y is a visiting professor at the National University of Singapore. E.L.V. is co-founder, member of the Board of Directors, on the SAB, equity holder and paid consultant for Eclipse BioInnovations. The interests of G.W.Y. and E.L.V. have been reviewed and approved by the University of California, San Diego in accordance with its conflict of interest policies. N.L.A. is a co-founder, on the SAB, equity holder and paid consultant for Altis Biosystems and a co-founder and equity holder in Cell Microsystems. The interests of N.L.A. have been reviewed and approved by the University of North Carolina, Chapel Hill through 1 November 2019 and by University of Washington, Seattle as of 1 November 2019 in accordance with their conflict of interest policies. The authors declare no other competing interests.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of microraft arrays for stress granule quantification.
a, Uncropped Western blots measuring EIF2AK1 protein expression in cells infected with sg-NTC (nontargeting control), sg-EIF2AK1, or uninfected control cells (293 T). (n = 1) b, Scatterplot of mCherry and mCitrine area measured in the nuclei of all colonies detected on a microraft array. Colonies that contain fluorescent signal from both channels in more than 10% of the total nuclei area are determined as doublets (gray). c, Time-course analysis of stress granule formation in HEK293T cells under multiple sodium arsenite concentrations measured in 30-minute intervals. Stress granule area is quantified as G3BP1(+) cytoplasmic puncta across n = 1 image. d, Top, schematic of microraft array without glass-back support. Orthogonal view of autofluorescence (green) in microrafts across PDMS array after imaging with high laser power. Bottom, diagram of microraft array with 1 mm glass support with orthogonal view of autofluorescent microrafts after imaging with high laser power (green). e, Random sampling to estimate plating frequency of sgRNAs on rafts in this screen. Given the relative abundances of sgRNAs on day 7 and the total number of colonies plated (~120,000), random sampling was used to estimate the number of rafts that contain each sgRNA (x-axis), binned in counts of 5. Bars are the average of n = 10 random samplings with error bars displaying standard deviation.
Extended Data Fig. 2 Performance of classifiers in image filtering model.
a, Learning curves for each binary classifier for 10,000 epochs of training. b, Top, confusion matrix for 365 test images comparing the overall model’s predicted classifications for each image with its ground-truth. Bottom, average precision rate, recall rate (true positive rate), F1-scores (harmonic mean of precision and recall), and number of images (n) for each binary classifier.
Extended Data Fig. 3 Library preparation scheme to sequence sgRNA infected in colonies.
a, Schematic of PCR barcoding design targeting common regions flanking the sgRNA insert. b, Agarose gel of PCR products with increasing cycle numbers to determine the minimum number of PCR cycles required to amplify a product for sequencing. Input material from bulk sample is used as a positive control. All rafts sequenced in this study were amplified with 22 cycles for PCR1 and 10 cycles for PCR2 (n = 213 total, 173 successful). c, TapeStation results of PCR products for one representative library containing four pooled microrafts. d, Agarose gel of PCR2 product for three representative libraries, each containing four pooled microrafts. Gel extraction was used to isolate the product of interest (red box) from a total of n = 32 libraries. e, Summary of the number of sgRNAs identified from each isolated cell colony. f, Bar chart of the total number of rafts picked, sequenced, and confirmed by siRNA depletion.
Extended Data Fig. 4 Uncropped Western blots for siRNA knock-down experiments.
a, Samples with knock-down of each protein (labeled above) compared to nontargeting control and untransfected sample. 1 - si-KD targeting, 2 - si-Nontargeting control, 3 - Untransfected HEK293T cells. b, GAPDH blots for each respective sample tested in panel a using antibodies of the opposite species on the same membrane.
Extended Data Fig. 5 Depletion of stress-granule regulatory proteins also reduces UBAP2L puncta formation.
a, siRNA depletion of target RBPs. UBAP2L(+) granule/nuclei area was normalized to the nontargeting control (NTC) for each experiment. RBPs are ordered in order of appearance in Fig. 3c. *RBPs that had significant reduction (P < 0.05, unpaired two-tailed t test, d.f. = 4, 95% confidence interval) of UBAP2L (+) granule area relative to NTC in at least 3 of the 4 biological replicates. Data are mean ± s.d. across n = 3 wells/condition (4 images/well). b, G3BP1(+) granule/nuclei area from respective wells measured in panel a. Values are normalized to nontargeting control (NTC) for each experiment. *RBPs that had significant reduction (P < 0.05, unpaired two-tailed t test, d.f. = 4, 95% confidence interval) of UBAP2L (+) granule area relative to NTC in at least 3 of the 4 biological replicates. Data are mean ± s.d. across n = 3 wells/condition (4 images/well).
Supplementary information
Supplementary Information
Step-by-step experimental protocol required to perform a screen with CRaft-ID.
Supplementary Tables
Excel workbook with Supplementary Tables 1–3.
Rights and permissions
About this article
Cite this article
Wheeler, E.C., Vu, A.Q., Einstein, J.M. et al. Pooled CRISPR screens with imaging on microraft arrays reveals stress granule-regulatory factors. Nat Methods 17, 636–642 (2020). https://doi.org/10.1038/s41592-020-0826-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0826-8
This article is cited by
-
Apurinic/apyrimidinic endodeoxyribonuclease 1 (APE1) promotes stress granule formation via YBX1 phosphorylation in ovarian cancer
Cellular and Molecular Life Sciences (2024)
-
Time-resolved proteomic profiling reveals compositional and functional transitions across the stress granule life cycle
Nature Communications (2023)
-
A brief guideline for studies of phase-separated biomolecular condensates
Nature Chemical Biology (2022)
-
Illuminating RNA biology through imaging
Nature Cell Biology (2022)
-
Pooled image-base screening of mitochondria with microraft isolation distinguishes pathogenic mitofusin 2 mutations
Communications Biology (2022)