Abstract
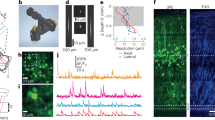
We designed a head-mounted three-photon microscope for imaging deep cortical layer neuronal activity in a freely moving rat. Delivery of high-energy excitation pulses at 1,320 nm required both a hollow-core fiber whose transmission properties did not change with fiber movement and dispersion compensation. These developments enabled imaging at >1.1 mm below the cortical surface and stable imaging of layer 5 neuronal activity for >1 h in freely moving rats performing a range of behaviors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Three-photon imaging datasets acquired from awake and freely moving animals are available upon reasonable request. Owing to the size of the datasets, they are not available on a public server. Optical designs for custom parts are available at https://www.caesar.de/en/our-research/current-groups/behavior-and-brain-organization/designs.html. Segments of custom fiber are available at a nominal cost to cover production of the fiber upon request.
Code availability
The code supporting the plots and other findings in the manuscript are available from the corresponding author upon reasonable request.
References
Helmchen, F., Fee, M. S., Tank, D. W. & Denk, W. A miniature head-mounted two-photon microscope. High-resolution brain imaging in freely moving animals. Neuron 31, 903–912 (2001).
Sawinski, J. et al. Visually evoked activity in cortical cells imaged in freely moving animals. Proc. Natl Acad. Sci. USA 106, 19557–19562 (2009).
Zong, W. et al. Fast high-resolution miniature two-photon microscopy for brain imaging in freely behaving mice. Nat. Methods 14, 713–719 (2017).
Horton, N. G. et al. In vivo three-photon microscopy of subcortical structures within an intact mouse brain. Nat. Photonics 7, 205–209 (2013).
Wang, T. et al. Three-photon imaging of mouse brain structure and function through the intact skull. Nat. Methods 15, 789–792 (2018).
Yildirim, M., Sugihara, H., So, P. T. C. & Sur, M. Functional imaging of visual cortical layers and subplate in awake mice with optimized three-photon microscopy. Nat. Commun. 10, 177 (2019).
Russell, P. Photonic crystal fibers. Science 299, 358–362 (2003).
Frosz, M. H., Roth, P., Günendi, M. C. & Russell, P. S. J. Analytical formulation for the bend loss in single-ring hollow-core photonic crystal fibers. Photonics Res. 5, 88–91 (2017).
Ouzounov, D. G., Wang, T., Wu, C. & Xu, C. GCaMP6 ΔF/F dependence on the excitation wavelength in 3-photon and 2-photon microscopy of mouse brain activity. Biomed. Opt. Express 10, 3343–3352 (2019).
Bouwmans, G. et al. Properties of a hollow-core photonic bandgap fiber at 850 nm wavelength. Opt. Express 11, 1613–1620 (2003).
Proctor, B. & Wise, F. Quartz prism sequence for reduction of cubic phase in a mode-locked TiAl2O3 laser. Opt. Lett. 17, 1295–1297 (1992).
Horton, N. G. & Xu, C. Dispersion compensation in three-photon fluorescence microscopy at 1,700 nm. Biomed. Opt. Express 6, 1392–1397 (2015).
Piyawattanametha, W. et al. Fast-scanning two-photon fluorescence imaging based on a microelectromechanical systems two-dimensional scanning mirror. Opt. Lett. 31, 2018–2020 (2006).
Mittmann, W. et al. Two-photon calcium imaging of evoked activity from L5 somatosensory neurons in vivo. Nat. Neurosci. 14, 1089–1093 (2011).
Girman, S. V., Sauve, Y. & Lund, R. D. Receptive field properties of single neurons in rat primary visual cortex. J. Neurophysiol. 82, 301–311 (1999).
Wallace, D. J. et al. Rats maintain an overhead binocular field at the expense of constant fusion. Nature 498, 65–69 (2013).
Hopt, A. & Neher, E. Highly nonlinear photodamage in two-photon fluorescence microscopy. Biophys. J. 80, 2029–2036 (2001).
Koester, H. J., Baur, D., Uhl, R. & Hell, S. W. Ca2+ fluorescence imaging with pico- and femtosecond two-photon excitation: signal and photodamage. Biophys. J. 77, 2226–2236 (1999).
Harvey, C. D., Coen, P. & Tank, D. W. Choice-specific sequences in parietal cortex during a virtual-navigation decision task. Nature 484, 62–68 (2012).
Nimmerjahn, A., Kirchhoff, F., Kerr, J. N. & Helmchen, F. Sulforhodamine 101 as a specific marker of astroglia in the neocortex in vivo. Nat. Methods 1, 31–37 (2004).
Brainard, D. H. The Psychophysics Toolbox. Spat. Vis. 10, 433–436 (1997).
Pelli, D. G. The VideoToolbox software for visual psychophysics: transforming numbers into movies. Spat. Vis. 10, 437–442 (1997).
Euler, T. et al. Eyecup scope—optical recordings of light stimulus-evoked fluorescence signals in the retina. Pflügers Arch. 457, 1393–1414 (2009).
Strickland, D. & Mourou, G. Compression of amplified chirped optical pulses. Opt. Commun. 55, 447–449 (1985).
Pachitariu, M. et al. Suite2p: beyond 10,000 neurons with standard two-photon microscopy. Preprint at bioRxiv https://doi.org/10.1101/061507 (2017).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Acknowledgements
We thank K. Briggman for comments on an earlier version of this manuscript, C. Xu (Cornell University) for his generous gift of an initial fiber to test, and both him and W. Denk for insightful discussions. We thank R. Pohle for assistance with editing. We thank U. Czubayko, J. Klesing and R. Beck for help with histology. We thank M. Bräuer, R. Honnef, M. Strauβfeld and Z. Amir from the mechanical workshop for production of microscope parts. Funding was obtained from Stiftung Caesar and the Max Planck Society and from Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under SFB 1089-227953431.
Author information
Authors and Affiliations
Contributions
Experimental design: A.K., D.J.W., J.N.D.K. Microscope hardware and software development: A.K., J.S. Fiber design, fabrication and characterization: M.H.F., R.Z., P.S.J.R. Animal and indicator preparation: D.J.W., J.S., V.P. Data collection: A.K., D.J.W., J.S., K.-M.V., J.N.D.K. Analysis design and implementation: A.K., D.J.W., K.-M.V., J.N.D.K. Manuscript preparation: A.K., D.J.W., J.S., J.N.D.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Properties of the hollow core photonic bandgap crystal fiber (HC-PBGF).
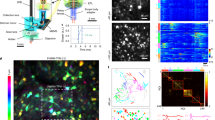
a, average image of pollen grains acquired from the miniature microscope using a single-ring hollow-core photonic crystal fiber with a 25 µm core diameter, showing the region of interest (ROI, green) in which the fluorescence trace in b was quantified. b, fluorescence trace before and during (gray box) bending of the fiber approximating that experienced during freely moving experiments. Insert shows a micrograph of a cross section through the fiber. c, pollen grains imaged using a HC-PBGF with a 14 µm core-diameter. ROI in which the fluorescence trace in d shown in green. d, fluorescence trace before and during (gray box) bending of the fiber. Insert shows a micrograph of a cross section through the fiber. e, spectral phase of the pulses propagating through 1.1 m of fiber, with compensation of group velocity dispersion (GVD) only (green), third-order dispersion (TOD) and GVD compensated (blue) and the flat phase corresponding to the transform-limited pulse (red) in Fig. 1d. The insert shows the same curves with magnified Y-axis scaling and the normalized spectrum used for the measurements of the spectral phases (black). f, transmission losses of the fiber as a function of wavelength. Plot shows mean (dark blue) and measurement uncertainty, calculated as described previously13. g, full laser pulse temporal profile after the fiber before dispersion compensation (green) and after compensation (blue) compared to calculated transform-limited pulse with the spectrum measured after the fiber (red). h, example laser spectra recorded after transmission through the HC-PBGF for incident power giving 0.5 (blue), 15.7 (red) and 87.5 mW (yellow) transmitted power at the sample. i, laser spectrum center of mass wavelength recorded after transmission through the HC-PBGF as a function of incident power (power measured at the sample). Panels a-d representative of 2 separate experiments.
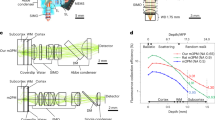
Extended Data Fig. 2 Microscope optical pathway and parts list.
a, optical design for miniature microscope with parts listing.
Extended Data Fig. 3 Three-photon excitation through the HC-PBGF and head-mounted microscope, integration window gating and subcellular structures visible at different cortical depths.
a, log-log plot of gray scale value (GSV) of fluorescence intensity as a function of laser power demonstrating 3-photon excitation. Least squares fit has a slope of 2.99. b, probability distribution of photon counts after the laser pulse trigger. Shaded region shows time window for laser pulse triggered data acquisition. Insert shows raw data from 2609 individual laser pulses. c, example images of the same pollen grain acquired with standard (left) and gated (right) acquisition. e, box and whisker plot of image signal-to-noise ratio for gated and standard image acquisition. Red line indicates data median, box shows 25th and 75th percentile of data and whiskers represent an extension to 1.5 times the interquartile range (outliers are plotted as red markers). f–h, overview images showing dendrites (arrowheads) visible at the indicated depths from the cortical surface. Scale bar in g 20 µm, applies to all panels. Panels f–h are representative of >6 datasets from 6 animals.
Extended Data Fig. 4 Statistics of animals’ behavior, image displacement and stability of neuropil fluorescence recorded during freely moving imaging datasets.
a, overlay of movement paths for all datasets from all animals, each path represented in a different color (n=7 datasets from 5 animals). Track outline shown in black. Paths can leave the track boundary when the animal reaches its head over the track edge to peer around. b, distributions showing animals’ velocity (left) and acceleration (right) during the imaging files for which the behavioral paths are shown in a. Distributions for individual animals shown in gray, mean in black. c, distributions of head roll (left), pitch (middle) and azimuth (right) over all datasets shown in a, during periods when the animals’ velocity was greater than 0.05 ms-1. Positive roll is roll to the animal’s right, positive pitch is up and azimuth 0° is up in a. Distributions for individual animals shown in cyan, mean in black. d, same as for c, but including all data, including extended periods where the animal was grooming or peering in one direction off the track. e, distribution of image displacement (mean ± SD) for different animal velocities for all datasets (7 datasets from 5 animals). Colors denote different datasets. f, mean image displacement (mean ± SD) as a function of animal acceleration. Datasets and colors as in e.
Extended Data Fig. 5 Analysis of fast axial motion.
a, raw fluorescence trace (black) and low-pass filtered fluorescence signal (Gaussian filter, 150 ms time constant, red). Blue crosses show the negative transients selected by the following analysis. b, high-pass filtered trace obtained by subtracting a low-pass filtered fluorescence from the raw trace. c, high-pass filtered signal from b scaled with the square root of the low-pass filtered fluorescence signal and the chosen threshold (red line). d, fluorescence trace obtained by low-pass filtering the raw signal (Gaussian filter, 30 ms). Black crosses show timings, when events were selected. e, fluorescence trace obtained by applying a rolling-average of 2 frames on the raw data. Black crosses show timings, when events were selected. f, individual examples (blue) and mean (red) of relative changes of fluorescence centered around the selected events from one neuron over 200 s of imaging data (<20 events from 6000 frames). Mean from rolling 2 frame average filtered data for the same data segments shown in black. g, average event trace (red) and the corresponding filtered traces with a rolling average of 2 frames filter (black), 30 ms Gaussian filter (green), 50 ms Gaussian filter (cyan) and 75 ms Gaussian filter (magenta). h, sequence of 7 consecutive images from the frame sequence averaged around detected events. Frame 4 corresponds to the average detected event. Neuron of interest outlined in red in frame 4. i, heat map of the probability of occurrence of events over the track. j, histograms of probabilities of the occurrence of events over the range measured during imaging for acceleration (upper left), velocity (lower left), angular acceleration (upper right) and angular velocity (lower right).
Extended Data Fig. 6 Headplate implant and microscope mounting plates.
a, individual diagrams of headplate and associated microscope mounting plates. i, implanted titanium headplate. ii, intermediate plate. iii, objective mount plate. iv, objective thread. v, miniature objective with threaded brass sleeve. vi, removable 30 × 5 mm2 tab for anesthetized experiments (alternative 15 × 5 mm2 tab used for awake experiments). b, cross section through headplate and microscope mounting plates as assembled during imaging experiments.
Extended Data Fig. 7 Microscope detector system design and excitation at 1320 nm wavelength of sulforhodamine 101.
a, detector system design and parts listing. b, example multi-channel green, red and blue data acquisition using the miniature microscope, with merged image overlay. Green labelling is GCaMP6s labelled neurons, red from SR101 labelled astrocytes and blue from third-harmonic signal, all in rat posterior parietal cortex. c, dependence of red fluorescence emission as a function of excitation laser power at 1320 nm. Data in panel b representative of 4 experiments in 4 animals. Data in panel c from a single measurement.
Supplementary information
Supplementary Information
Supplementary Note
41592_2020_817_MOESM3_ESM.mp4
Supplementary Video 1 Side imaging of fluorescence distribution on a phantom with fluorescent beads excited with two or three photons with increasing depth. The phantom was prepared as described previously in ref. 1. In brief, the phantom consisted of agarose (0.5%) and contained 1-µm non-fluorescent polystyrene beads at a concentration of 5.4 × 109 beads per ml. A concentration of 2.8 × 108 beads per ml for 0.5-µm yellow-green beads was used to provide fluorescence. Two-photon excitation was provided with a conventional two-photon microscope and a long-working-distance (4-mm) high-NA (NA 1.05) objective. Side imaging was performed with a minimal imaging system consisting of an imaging lens with f = 30 mm and a camera. Note that a sharp focal point is not observed because of the thickness of the phantom. The focusing depth was gradually increased and the excitation laser power was increased to keep the signal intensity approximately constant, until strong out-of-focus fluorescence was obtained. The same phantom was then imaged under a three-photon microscope with the same objective lens. The same side imaging system and sequence were used. The two videos were combined in this final video. The approximate imaging depth for the two-photon excitation condition was 1 mm. For the two-photon measurement, the laser power maximum was 200 mW at 920 nm (80 MHz), while for the three-photon measurement it was 150 mW at 1,300 nm (300 kHz) at maximum. Note that the phantom has been designed to mimic two-photon scattering and was not tailored specifically to reproduce actual brain scattering under 3PE conditions. The video was generated from a single measurement.
41592_2020_817_MOESM4_ESM.avi
Supplementary Video 2 Imaging with the head-mounted three-photon microscope from the surface of the cortex down to 1,120 µm. Image stack to 1,120 µm below the cortical surface showing GCaMP6s-labeled neurons (green) and astrocytes (red). Axial spacing was 2 µm, and individual frames are an average of 20 images. Data are representative of four experiments from four animals.
41592_2020_817_MOESM5_ESM.mp4
Supplementary Video 3 Imaging with the head-mounted three-photon microscope in a freely moving animal. Synchronized overview image (upper left), animal position tracking (upper. right), raw 3PE imaging data (lower left) and Ca2+ kinetic traces from example neurons in the field of view (lower right). Red markers in animal position trace represent the position of the center of the microscope at the beginning of the 3PE imaging frames. Red synchronization line on the Ca2+ kinetic traces indicates the location on the traces of the currently displayed images. Data representative of 8 datasets from 6 animals.
Rights and permissions
About this article
Cite this article
Klioutchnikov, A., Wallace, D.J., Frosz, M.H. et al. Three-photon head-mounted microscope for imaging deep cortical layers in freely moving rats. Nat Methods 17, 509–513 (2020). https://doi.org/10.1038/s41592-020-0817-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0817-9
This article is cited by
-
Large-scale cranial window for in vivo mouse brain imaging utilizing fluoropolymer nanosheet and light-curable resin
Communications Biology (2024)
-
Deep brain imaging on the move
Nature Methods (2023)
-
Large depth-of-field ultra-compact microscope by progressive optimization and deep learning
Nature Communications (2023)
-
A three-photon head-mounted microscope for imaging all layers of visual cortex in freely moving mice
Nature Methods (2023)
-
Miniature three-photon microscopy maximized for scattered fluorescence collection
Nature Methods (2023)