Abstract
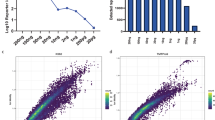
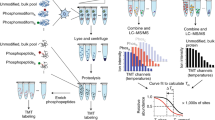
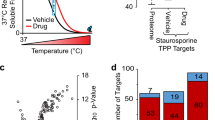
Isobaric labeling empowers proteome-wide expression measurements simultaneously across multiple samples. Here an expanded set of 16 isobaric reagents based on an isobutyl-proline immonium ion reporter structure (TMTpro) is presented. These reagents have similar characteristics to existing tandem mass tag reagents but with increased fragmentation efficiency and signal. In a proteome-scale example dataset, we compared eight common cell lines with and without Torin1 treatment with three replicates, quantifying more than 8,800 proteins (mean of 7.5 peptides per protein) per replicate with an analysis time of only 1.1 h per proteome. Finally, we modified the thermal stability assay to examine proteome-wide melting shifts after treatment with DMSO, 1 or 20 µM staurosporine with five replicates. This assay identified and dose-stratified staurosporine binding to 228 cellular kinases in just one, 18-h experiment. TMTpro reagents allow complex experimental designs—all with essentially no missing values across the 16 samples and no loss in quantitative integrity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The MS data have been deposited in the ProteomeXchange Consortium with the dataset identifier PXD014369, PXD016491 and PXD016940. The list of human kinases was downloaded from UniProt on 9 June 2019 (https://www.uniprot.org/docs/pkinfam). Affinity data of 442 human kinases to staurosporine were downloaded from the database ‘The IUPHAR/BPS Guide to Pharmacology’ on 14 May 2019 (https://www.guidetopharmacology.org/).
References
Li, H. et al. Current trends in quantitative proteomics—an update. J. Mass Spectrom. 52, 319–341 (2017).
Pappireddi, N., Martin, L. & Wuhr, M. A review on quantitative multiplexed proteomics. Chem. Bio. Chem. 20, 1210–1224 (2019).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Rauniyar, N. & Yates, J. R. 3rd Isobaric labeling-based relative quantification in shotgun proteomics. J. Proteome Res. 13, 5293–5309 (2014).
Vasaikar, S. et al. Proteogenomic analysis of human colon cancer reveals new therapeutic opportunities. Cell 177, 1035–1049 e1019 (2019).
Chick, J. M. et al. Defining the consequences of genetic variation on a proteome-wide scale. Nature 534, 500–505 (2016).
Dayon, L. et al. Relative quantification of proteins in human cerebrospinal fluids by MS/MS using 6-plex isobaric tags. Anal. Chem. 80, 2921–2931 (2008).
Paulo, J. A. et al. Effects of MEK inhibitors GSK1120212 and PD0325901 in vivo using 10-plex quantitative proteomics and phosphoproteomics. Proteomics 15, 462–473 (2015).
Stepanova, E., Gygi, S. P. & Paulo, J. A. Filter-based protein digestion (FPD): a detergent-free and scaffold-based strategy for TMT workflows. J. Proteome Res. 17, 1227–1234 (2018).
McAlister, G. C. et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Anal. Chem. 84, 7469–7478 (2012).
Thompson, A. et al. TMTpro: design, synthesis, and initial evaluation of a proline-based isobaric 16-plex tandem mass tag reagent set. Anal. Chem. 91, 15941–15950 (2019).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Erickson, B. K. et al. Active instrument engagement combined with a real-time database search for improved performance of sample multiplexing workflows. J. Proteome Res. 18, 1299–1306 (2019).
Schweppe, D. K. et al. Full-featured, real-time database searching platform enables fast and accurate multiplexed quantitative proteomics. Preprint at bioRxiv https://doi.org/10.1101/668533 (2019).
Kim, J. & Guan, K. L. mTOR as a central hub of nutrient signalling and cell growth. Nat. Cell Biol. 21, 63–71 (2019).
Saxton, R. A. & Sabatini, D. M. mTOR Signaling in growth, metabolism, and disease. Cell 169, 361–371 (2017).
Thoreen, C. C. et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J. Biol. Chem. 284, 8023–8032 (2009).
Navarrete-Perea, J., Yu, Q., Gygi, S. P. & Paulo, J. A. Streamlined tandem mass tag (SL-TMT) protocol: an efficient strategy for quantitative (phospho)proteome profiling using tandem mass tag-synchronous precursor selection-MS3. J. Proteome Res. 17, 2226–2236 (2018).
Mizushima, N. A brief history of autophagy from cell biology to physiology and disease. Nat. Cell Biol. 20, 521–527 (2018).
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018).
An, H. et al. TEX264 Is an endoplasmic reticulum-resident ATG8-interacting protein critical for ER remodeling during nutrient stress. Mol. Cell. 74, 891–908 e810 (2019).
Wu, R. et al. Correct interpretation of comprehensive phosphorylation dynamics requires normalization by protein expression changes. Mol. Cell Proteom. 10, M111 009654 (2011).
Ben-Sahra, I., Howell, J. J., Asara, J. M. & Manning, B. D. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science 339, 1323–1328 (2013).
Barilari, M. et al. ZRF1 is a novel S6 kinase substrate that drives the senescence programme. EMBO J. 36, 736–750 (2017).
Gaetani, M. et al. Proteome integral solubility alteration: a high-throughput proteomics assay for target deconvolution. J. Proteome Res. 18, 4027–4037 (2019).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Harding, S. D. et al. The IUPHAR/BPS Guide to PHARMACOLOGY in 2018: updates and expansion to encompass the new guide to IMMUNOPHARMACOLOGY. Nucleic Acids Res. 46, D1091–D1106 (2018).
Becher, I. et al. Thermal profiling reveals phenylalanine hydroxylase as an off-target of panobinostat. Nat. Chem. Biol. 12, 908–910 (2016).
Dai, L. et al. Horizontal cell biology: monitoring global changes of protein interaction states with the proteome-wide cellular thermal shift assay (CETSA). Annu. Rev. Biochem. 88, 383–408 (2019).
Savitski, M. M. et al. Multiplexed proteome dynamics profiling reveals mechanisms controlling protein homeostasis. Cell 173, 260–274 e225 (2018).
Zecha, J. et al. Peptide level turnover measurements enable the study of proteoform dynamics. Mol. Cell Proteom. 17, 974–992 (2018).
Dephoure, N. & Gygi, S. P. Hyperplexing: a method for higher-order multiplexed quantitative proteomics provides a map of the dynamic response to rapamycin in yeast. Sci. Signal. 5, rs2 (2012).
Hebert, A. S. et al. Neutron-encoded mass signatures for multiplexed proteome quantification. Nat. Methods 10, 332–334 (2013).
Mulvey, C. M. et al. Using hyperLOPIT to perform high-resolution mapping of the spatial proteome. Nat. Protoc. 12, 1110–1135 (2017).
Hughes, C. S. et al. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat. Protoc. 14, 68–85 (2019).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Savitski, M. M., Wilhelm, M., Hahne, H., Kuster, B. & Bantscheff, M. A scalable approach for protein false discovery rate estimation in large proteomic data sets. Mol. Cell Proteom. 14, 2394–2404 (2015).
Beausoleil, S. A., Villen, J., Gerber, S. A., Rush, J. & Gygi, S. P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, 1285–1292 (2006).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank members of the Gygi Laboratory, particularly R. Rad, at Harvard Medical School. This work was funded in part by NIH grant nos. 1R01GM132129 (to J.A.P.) and 5R01GM067945 (to S.P.G), and the Mark Foundation for Cancer Research Fellow of the Damon Runyon Cancer Research Foundation DRG 2359-19 (to J.G.V.V.).
Author information
Authors and Affiliations
Contributions
J.L. prepared the cell line samples treated with Torin1 for MS analysis, performed the data analyses, prepared the figures and wrote the manuscript. J.G.V.V. prepared and conducted the PISA experiments and prepared associated figures. L.P.V. proposed the eight cell lines, then grew and treated the lines for Torin1 experiments and performed western blotting experiments. D.K.S. developed and implemented the real-time online searching tool. E.L.H. advised on data analyses. R.V. and A.M.R. provided additional reagent characterization and advice. C.E., P.N., R.D.B. and J.C.R. further characterized the TMTpro reagents, initiated the collaboration and provided input and oversight for the project. Reagents were conceived, developed, synthesized and characterized by K.K., A.H.T. and I.P. S.P.G. oversaw the project and edited the manuscript. J.A.P. oversaw the project, performed the experiment for comparison of TMT0 and TMTpro0, ran the MS analysis, performed the data analyses and wrote the manuscript. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The TMTpro reagents were commercialized by ThermoFisher Scientific in September 2019. C.E., P.N., R.V., A.M.R., R.D.B. and J.C.R. are employees of ThermoFisher Scientific. A.H.T. and I.P. were employees of Proteome Sciences. K.K. is an employee of Proteome Sciences. S.P.G. is a member of the scientific advisory board for ThermoFisher Scientific.
Additional information
Peer review information Allison Doerr was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12.
Supplementary Table 1
Protein quantifications in 48 cell line samples treated with DMSO and Torin1 (RTS–SPS–MS3).
Supplementary Table 2
Protein quantifications in replicate1 of the 48 cell line samples treated with DMSO and Torin1 (hrMS2 and SPS–MS3).
Supplementary Table 3
Phosphorylation quantifications in 48 cell line samples treated with DMSO and Torin1.
Supplementary Table 4
Protein quantifications in the PISA assay (RTS–SPS–MS3).
Supplementary Table 5
Protein quantifications in the PISA assay (hrMS2).
Supplementary Table 6
Correction factors of the TMTpro reagents used in this work.
Rights and permissions
About this article
Cite this article
Li, J., Van Vranken, J.G., Pontano Vaites, L. et al. TMTpro reagents: a set of isobaric labeling mass tags enables simultaneous proteome-wide measurements across 16 samples. Nat Methods 17, 399–404 (2020). https://doi.org/10.1038/s41592-020-0781-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-0781-4
This article is cited by
-
Monocarboxylate transporters facilitate succinate uptake into brown adipocytes
Nature Metabolism (2024)
-
A complex interplay of intra- and extracellular factors regulates the outcome of fetal- and adult-derived MLL-rearranged leukemia
Leukemia (2024)
-
Functionalizing tandem mass tags for streamlining click-based quantitative chemoproteomics
Communications Chemistry (2024)
-
Mitochondrial ATP generation is more proteome efficient than glycolysis
Nature Chemical Biology (2024)
-
Quantitative proteomic analysis of human serum using tandem mass tags to predict cardiovascular risks in patients with psoriasis
Scientific Reports (2023)