Abstract
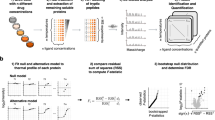
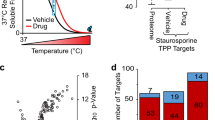
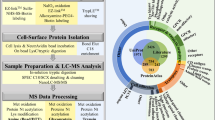
Numerous drugs and endogenous ligands bind to cell surface receptors leading to modulation of downstream signaling cascades and frequently to adaptation of the plasma membrane proteome. In-depth analysis of dynamic processes at the cell surface is challenging due to biochemical properties and low abundances of plasma membrane proteins. Here we introduce cell surface thermal proteome profiling for the comprehensive characterization of ligand-induced changes in protein abundances and thermal stabilities at the plasma membrane. We demonstrate drug binding to extracellular receptors and transporters, discover stimulation-dependent remodeling of T cell receptor complexes and describe a competition-based approach to measure target engagement of G-protein-coupled receptor antagonists. Remodeling of the plasma membrane proteome in response to treatment with the TGFB receptor inhibitor SB431542 leads to partial internalization of the monocarboxylate transporters MCT1/3 explaining the antimetastatic effects of the drug.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All source data are available in the main text or the supplementary materials. The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE59 partner repository with the dataset identifier PXD016249. Annotations of proteins were based on the UniProt database (14 December 2016, https://www.uniprot.org/).
Code availability
An implementation of the above described CS-TPP analysis procedure can be found at https://github.com/mathiaskalxdorf/RTSA or at https://doi.org/10.5281/zenodo.4274152.
References
Yin, H. & Flynn, A. D. Drugging membrane protein interactions. Annu. Rev. Biomed. Eng. 18, 51–76 (2016).
Guan, Y. et al. Kinetics of small molecule interactions with membrane proteins in single cells measured with mechanical amplification. Sci. Adv. 1, e1500633 (2015).
Lomenick, B. et al. Target identification using drug affinity responsive target stability (DARTS). Proc. Natl Acad. Sci. USA 106, 21984–21989 (2009).
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Bantscheff, M., Schirle, M., Sweetman, G., Rick, J. & Kuster, B. Quantitative mass spectrometry in proteomics: a critical review. Anal. Bioanal. Chem. 389, 1017–1031 (2007).
Rutkowska, A. et al. A modular probe strategy for drug localization, target identification and target occupancy measurement on single cell level. ACS Chem. Biol. 11, 2541–2550 (2016).
Frei, A. P., Moest, H., Novy, K. & Wollscheid, B. Ligand-based receptor identification on living cells and tissues using TRICEPS. Nat. Protoc. 8, 1321–1336 (2013).
Reinhard, F. B. M. et al. Thermal proteome profiling monitors ligand interactions with cellular membrane proteins. Nat. Methods 12, 1129–1131 (2015).
Kawatkar, A. et al. CETSA beyond soluble targets: a broad application to multipass transmembrane proteins. ACS Chem. Biol. 14, 1913–1920 (2019).
Kalxdorf, M., Gade, S., Eberl, H. C. & Bantscheff, M. Monitoring cell surface N-glycoproteome dynamics by quantitative proteomics reveals mechanistic insights into macrophage differentiation. Mol. Cell. Proteomics 16, 770–785 (2017).
Zeng, Y., Ramya, T. N. C., Dirksen, A., Dawson, P. E. & Paulson, J. C. High-efficiency labeling of sialylated glycoproteins on living cells. Nat. Methods 6, 207–209 (2009).
Cvjetkovic, A. et al. Detailed analysis of protein topology of extracellular vesicles–evidence of unconventional membrane protein orientation. Sci. Rep. 6, 36338 (2016).
Leuenberger, P. et al. Cell-wide analysis of protein thermal unfolding reveals determinants of thermostability. Science 355, eaai7825 (2017).
Haltia, T. & Freire, E. Forces and factors that contribute to the structural stability of membrane proteins. Biochim. Biophys. Acta 1228, 1–27 (1995).
Gagnon, K. B. & Delpire, E. Physiology of SLC12 transporters: lessons from inherited human genetic mutations and genetically engineered mouse knockouts. Am. J. Physiol. Cell Physiol. 304, C693–C714 (2013).
Falivelli, G. et al. Attenuation of eph receptor kinase activation in cancer cells by coexpressed ephrin ligands. PLoS ONE 8, e81445 (2013).
Kanatani, Y. et al. Role of CD14 expression in the differentiation-apoptosis switch in human monocytic leukemia cells treated with 1alpha,25-dihydroxyvitamin D3 or dexamethasone in the presence of transforming growth factor beta1. Cell Growth Differ. Mol. Biol. J. Am. Assoc. Cancer Res. 10, 705–712 (1999).
Nyhan, K. C. et al. Jagged/Notch signalling is required for a subset of TGFβ1 responses in human kidney epithelial cells. Biochim. Biophys. Acta 1803, 1386–1395 (2010).
Romero, M. F., Chen, A.-P., Parker, M. D. & Boron, W. F. The SLC4 family of bicarbonate (HCO3−) transporters. Mol. Asp. Med. 34, 159–182 (2013).
Fransvea, E., Angelotti, U., Antonaci, S. & Giannelli, G. Blocking transforming growth factor-beta up-regulates E-cadherin and reduces migration and invasion of hepatocellular carcinoma cells. Hepatol. 47, 1557–1566 (2008).
Zhang, Q. et al. LY2157299 monohydrate, a TGF-βR1 inhibitor, suppresses tumor growth and ascites development in ovarian cancer. Cancers https://doi.org/10.3390/cancers10080260 (2018).
Halder, S. K., Beauchamp, R. D. & Datta, P. K. A specific inhibitor of TGF-beta receptor kinase, SB-431542, as a potent antitumor agent for human cancers. Neoplasia 7, 509–521 (2005).
Miranda-Gonçalves, V. et al. Hypoxia-mediated upregulation of MCT1 expression supports the glycolytic phenotype of glioblastomas. Oncotarget 7, 46335–46353 (2016).
Xu, R.-G. et al. MCT1 promotes tumor progression through regulating epithelial-mesenchymal transition in pancreatic cancer. Int. J. Clin. Exp. Pathol. 10, 3243–3250 (2017).
Gray, A. L., Coleman, D. T., Shi, R. & Cardelli, J. A. Monocarboxylate transporter 1 contributes to growth factor-induced tumor cell migration independent of transporter activity. Oncotarget 7, 32695–32706 (2016).
Payen, V. L. et al. Monocarboxylate transporter MCT1 promotes tumor metastasis independently of its activity as a lactate transporter. Cancer Res. 77, 5591–5601 (2017).
Gaetke, L. M., Chow-Johnson, H. S. & Chow, C. K. Copper: toxicological relevance and mechanisms. Arch. Toxicol. 88, 1929–1938 (2014).
Virginio, C., Church, D., North, R. A. & Surprenant, A. Effects of divalent cations, protons and calmidazolium at the rat P2X7 receptor. Neuropharmacology 36, 1285–1294 (1997).
Gómez, M., González, A., Sáez, C. A. & Moenne, A. Copper-induced membrane depolarizations involve the induction of mosaic TRP channels, which activate VDCC leading to calcium increases in ulva compressa. Front. Plant Sci. 7, 754 (2016).
Matsuzaki, S. et al. Annexin A4-conferred platinum resistance is mediated by the copper transporter ATP7A. Int. J. Cancer 134, 1796–1809 (2014).
Trickett, A. & Kwan, Y. L. T cell stimulation and expansion using anti-CD3/CD28 beads. J. Immunol. Methods 275, 251–255 (2003).
Pozzi, N. et al. Defective surface expression of attractin on T cells in patients with common variable immunodeficiency (CVID). Clin. Exp. Immunol. 123, 99–104 (2001).
Tan, C. S. H. et al. Thermal proximity coaggregation for system-wide profiling of protein complex dynamics in cells. Science 359, 1170–1177 (2018).
Birnbaum, M. E. et al. Molecular architecture of the αβ T cell receptor-CD3 complex. Proc. Natl Acad. Sci. USA 111, 17576–17581 (2014).
Morra, M., Zubiaur, M., Terhorst, C., Sancho, J. & Malavasi, F. CD38 is functionally dependent on the TCR/CD3 complex in human T cells. FASEB J. 12, 581–592 (1998).
Voisinne, G., Gonzalez de Peredo, A. & Roncagalli, R. CD5, an undercover regulator of TCR signaling. Front. Immunol. 9, 2900 (2018).
Stillwell, R. & Bierer, B. E. T cell signal transduction and the role of CD7 in costimulation. Immunol. Res. 24, 31–52 (2001).
Kumar, A. et al. CXCR4 physically associates with the T cell receptor to signal in T cells. Immunity 25, 213–224 (2006).
Muhammad, A. et al. Sequential cooperation of CD2 and CD48 in the buildup of the early TCR signalosome. J. Immunol. 182, 7672–7680 (2009).
Lioudyno, M. I. et al. Orai1 and STIM1 move to the immunological synapse and are up-regulated during T cell activation. Proc. Natl Acad. Sci. USA 105, 2011–2016 (2008).
Dragovich, M. A. et al. SLAMF6 clustering is required to augment T cell activation. PLoS ONE 14, e0218109 (2019).
Yu, M. et al. Regulation of T cell receptor signaling by activation-induced zinc influx. J. Exp. Med. 208, 775–785 (2011).
Okiyoneda, T., Apaja, P. M. & Lukacs, G. L. Protein quality control at the plasma membrane. Curr. Opin. Cell Biol. 23, 483–491 (2011).
Triantafilou, K., Triantafilou, M. & Dedrick, R. L. A CD14-independent LPS receptor cluster. Nat. Immunol. 2, 338–345 (2001).
Hyun, S. Y. et al. Development of a novel Hsp90 inhibitor NCT-50 as a potential anticancer agent for the treatment of non-small cell lung cancer. Sci. Rep. 8, 1–16 (2018).
Huang, J. & Wang, H. Hsp83/Hsp90 physically associates with insulin receptor to promote neural stem cell reactivation. Stem Cell Rep. 11, 883–896 (2018).
Gaetani, M. et al. Proteome integral solubility alteration: a high-throughput proteomics assay for target deconvolution. J. Proteome Res. 18, 4027–4037 (2019).
Franken, H. et al. Thermal proteome profiling for unbiased identification of direct and indirect drug targets using multiplexed quantitative mass spectrometry. Nat. Protoc. 10, 1567–1593 (2015).
Silva, J. C., Gorenstein, M. V., Li, G.-Z., Vissers, J. P. C. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteom. 5, 144–156 (2006).
Becher, I. et al. Affinity profiling of the cellular kinome for the nucleotide cofactors ATP, ADP, and GTP. ACS Chem. Biol. 8, 599–607 (2013).
Savitski, M. M. et al. Delayed fragmentation and optimized isolation width settings for improvement of protein identification and accuracy of isobaric mass tag quantification on Orbitrap-type mass spectrometers. Anal. Chem. 83, 8959–8967 (2011).
Savitski, M. M. et al. Targeted data acquisition for improved reproducibility and robustness of proteomic mass spectrometry assays. J. Am. Soc. Mass. Spectrom. 21, 1668–1679 (2010).
Savitski, M. M. et al. Measuring and managing ratio compression for accurate iTRAQ/TMT quantification. J. Proteome Res. 12, 3586–3598 (2013).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B. Methodol. 57, 289–300 (1995).
Poole, W., Gibbs, D. L., Shmulevich, I., Bernard, B. & Knijnenburg, T. A. Combining dependent P-values with an empirical adaptation of Brown’s method. Bioinforma. 32, i430–i436 (2016).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 47, D442–D450 (2019).
Acknowledgements
We thank J. Stuhlfauth, N. Garcia-Altrieth, K. Beß and B. Dlugosch for the supporting cell culture production; M. Bösche, T. Rudi, M. Klös-Hudak, K. Kammerer and M. Steidel for assistance with mass spectrometry and C. Boecker and T. Mathieson for the IT and computational support.
Author information
Authors and Affiliations
Contributions
M.K. and H.C.E. designed the experiments. M.K. and I.G. performed the CS-TPP experiments. I.B. and S.K. performed lactate assay experiments. M.K. analyzed the CS-TPP data. N.K performed TPCA analysis. I.B. reviewed the figures. M.M.S. gave scientific advice. M.K., H.C.E. and M.B. wrote the manuscript. M.B. and H.C.E. supervised the work.
Corresponding authors
Ethics declarations
Competing interests
M.K., I.G., S.K., H.C.E. and M.B. are GSK employees. M.M.S. is a GSK shareholder.
Additional information
Peer review information Arunima, Singh was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Supplementary Table 1
Source data for the comparison of conventional TPP and TPP with cell surface enrichment on the example of cellular treatment with 1 µM ouabain.
Supplementary Table 2
Source data for all CS-TPP experiments.
Supplementary Table 3
Source data the gene ontology enrichment analyses comparing proteins with substantially different melting points (ΔTM > ±4 °C) between conventional TPP and TPP with additional cell surface enrichment.
Supplementary Table 4
Source data for the correlation of thermal stability with protein properties.
Supplementary Table 5
Summary of information about additionally affected proteins in CS-TPP experiments.
Supplementary Table 6
Source data for the TPCA analysis.
Rights and permissions
About this article
Cite this article
Kalxdorf, M., Günthner, I., Becher, I. et al. Cell surface thermal proteome profiling tracks perturbations and drug targets on the plasma membrane. Nat Methods 18, 84–91 (2021). https://doi.org/10.1038/s41592-020-01022-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-020-01022-1
This article is cited by
-
A chemical proteomics approach for global mapping of functional lysines on cell surface of living cell
Nature Communications (2024)
-
The Golgi stacking protein GRASP55 is targeted by the natural compound prodigiosin
Cell Communication and Signaling (2023)
-
Improved in situ characterization of protein complex dynamics at scale with thermal proximity co-aggregation
Nature Communications (2023)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)
-
KOPI: Kinase inhibitOr Proteome Impact analysis
Scientific Reports (2022)