Abstract
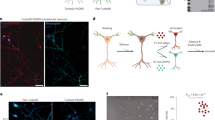
Spontaneous and sensory-evoked activity propagates across varying spatial scales in the mammalian cortex, but technical challenges have limited conceptual links between the function of local neuronal circuits and brain-wide network dynamics. We present a method for simultaneous cellular-resolution two-photon calcium imaging of a local microcircuit and mesoscopic widefield calcium imaging of the entire cortical mantle in awake mice. Our multi-scale approach involves a microscope with an orthogonal axis design where the mesoscopic objective is oriented above the brain and the two-photon objective is oriented horizontally, with imaging performed through a microprism. We also introduce a viral transduction method for robust and widespread gene delivery in the mouse brain. These approaches allow us to identify the behavioral state-dependent functional connectivity of pyramidal neurons and vasoactive intestinal peptide-expressing interneurons with long-range cortical networks. Our imaging system provides a powerful strategy for investigating cortical architecture across a wide range of spatial scales.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon request.
Code availability
All code used for analyses is available through Code Ocean.
References
Jiang, X. et al. Principles of connectivity among morphologically defined cell types in adult neocortex. Science 350, aac9462 (2015).
Wall, N. R. et al. Brain-wide maps of synaptic input to cortical interneurons. J. Neurosci. 36, 4000–4009 (2016).
Wertz, A. et al. PRESYNAPTIC NETWORKS. Single-cell-initiated monosynaptic tracing reveals layer-specific cortical network modules. Science 349, 70–74 (2015).
Crochet, S., Lee, S. H. & Petersen, C. C. H. Neural circuits for goal-directed sensorimotor transformations. Trends Neurosci. 42, 66–77 (2018).
Kim, E. J., Juavinett, A. L., Kyubwa, E. M., Jacobs, M. W. & Callaway, E. M. Three types of cortical layer 5 neurons that differ in brain-wide connectivity and function. Neuron 88, 1253–1267 (2015).
Chen, J. L., Carta, S., Soldado-Magraner, J., Schneider, B. L. & Helmchen, F. Behaviour-dependent recruitment of long-range projection neurons in somatosensory cortex. Nature 499, 336–340 (2013).
Han, Y. et al. The logic of single-cell projections from visual cortex. Nature 556, 51–56 (2018).
Lur, G., Vinck, M. A., Tang, L., Cardin, J. A. & Higley, M. J. Projection-specific visual feature encoding by layer 5 cortical subnetworks. Cell Rep. 14, 2538–2545 (2016).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Yang, H., Kwon, S. E., Severson, K. S. & O’Connor, D. H. Origins of choice-related activity in mouse somatosensory cortex. Nat. Neurosci. 19, 127–134 (2016).
Jun, J. J. et al. Fully integrated silicon probes for high-density recording of neural activity. Nature 551, 232–236 (2017).
Kim, C. K. et al. Simultaneous fast measurement of circuit dynamics at multiple sites across the mammalian brain. Nat. Methods 13, 325–328 (2016).
Sofroniew, N. J., Flickinger, D., King, J. & Svoboda, K. A large field of view two-photon mesoscope with subcellular resolution for in vivo imaging. eLife 5, e14472 (2016).
Stirman, J. N., Smith, I. T., Kudenov, M. W. & Smith, S. L. Wide field-of-view, multi-region, two-photon imaging of neuronal activity in the mammalian brain. Nat. Biotechnol. 34, 857–862 (2016).
Clancy, K., Orsolic, I. & Mrsic-Flogel, T. D. Locomotion-dependent remapping of distributed cortial networks. Nat. Neurosci. 22, 778–786 (2019).
Xiao, D. et al. Mapping cortical mesoscopic networks of single spiking cortical or sub-cortical neurons. eLife 6, e19976 (2017).
Lake, E. M. et al. Spanning spatiotemporal scales with simultaneous mesoscopic Ca2+imaging and functional MRI. Preprint at bioRxiv https://doi.org/10.1101/464305 (2018).
Lecoq, J. et al. Visualizing mammalian brain area interactions by dual-axis two-photon calcium imaging. Nat. Neurosci. 17, 1825–1829 (2014).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Ma, Y. et al. Wide-field optical mapping of neural activity and brain haemodynamics: considerations and novel approaches. Philos. Trans. R. Soc. Lond. B 371, 20150360 (2016).
Allen, W. E. et al. Global representations of goal-directed behavior in distinct cell types of mouse neocortex. Neuron 94, 891–907 (2017).
Ackman, J. B., Burbridge, T. J. & Crair, M. C. Retinal waves coordinate patterned activity throughout the developing visual system. Nature 490, 219–225 (2012).
Ackman, J. B., Zeng, H. & Crair, M. C. Structured dynamics of neural activity across developing neocortex. Preprint at bioRxiv https://doi.org/10.1101/012237 (2014).
Dubbs, A., Guevara, J. & Yuste, R. moco: fast motion correction for calcium imaging. Front. Neuroinform. 10, 6 (2016).
Pachitariu, M. et al. Suite2p: beyond 10,000 neurons with standard two-photon microscopy. Preprint at bioRxiv https://doi.org/10.1101/061507 (2017).
Podgorski, K. & Ranganathan, G. Brain heating induced by near-infrared lasers during multiphoton microscopy. J. Neurophysiol. 116, 1012–1023 (2016).
Franklin, T. B., Krueger-Naug, A. M., Clarke, D. B., Arrigo, A. P. & Currie, R. W. The role of heat shock proteins Hsp70 and Hsp27 in cellular protection of the central nervous system. Int J. Hyperth. 21, 379–392 (2005).
Batista-Brito, R. et al. Developmental dysfunction of vip interneurons impairs cortical circuits. Neuron 95, 884–895 (2017).
Vinck, M., Batista-Brito, R., Knoblich, U. & Cardin, J. A. Arousal and locomotion make distinct contributions to cortical activity patterns and visual encoding. Neuron 86, 740–754 (2015).
DeNardo, L. A., Berns, D. S., DeLoach, K. & Luo, L. Connectivity of mouse somatosensory and prefrontal cortex examined with trans-synaptic tracing. Nat. Neurosci. 18, 1687–1697 (2015).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Shen, X., Papademetris, X. & Constable, R. T. Graph-theory based parcellation of functional subunits in the brain from resting-state fMRI data. NeuroImage 50, 1027–1035 (2010).
Daigle, T. L. et al. A suite of transgenic driver and reporter mouse lines with enhanced brain-cell-type targeting and functionality. Cell 174, 465–480 (2018).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Steinmetz, N. A. et al. Aberrant cortical activity in multiple GCaMP6-expressing transgenic mouse lines. eNeuro 4, ENEURO.0207-17.2017 (2017).
Foust, K. D. et al. Intravascular AAV9 preferentially targets neonatal neurons and adult astrocytes. Nat. Biotechnol. 27, 59–65 (2009).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Hamodi, A. S., Sabino, A. M., Fitzgerald, N. D. & Crair, M. C. Transverse sinus injections: A novel method for whole-brain vector-driven gene delivery. Preprint at bioRxiv https://doi.org/10.1101/579730 (2019).
Tremblay, R., Lee, S. & Rudy, B. GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91, 260–292 (2016).
Fu, Y. et al. A cortical circuit for gain control by behavioral state. Cell 156, 1139–1152 (2014).
Pfeffer, C. K., Xue, M., He, M., Huang, Z. J. & Scanziani, M. Inhibition of inhibition in visual cortex: the logic of connections between molecularly distinct interneurons. Nat. Neurosci. 16, 1068–1076 (2013).
Lenschow, C. & Brecht, M. Barrel cortex membrane potential dynamics in social touch. Neuron 85, 718–725 (2015).
Klingler, E. et al. Single-cell molecular connectomics of intracortically-projecting neurons. Preprint at bioRxiv https://doi.org/10.1101/378760 (2018).
Tang, L. & Higley, M. J. Layer 5 circuits in V1 differentially control visuomotor behavior. Preprint at bioRxiv https://doi.org/10.1101/540807 (2019).
Lee, S., Kruglikov, I., Huang, Z. J., Fishell, G. & Rudy, B. A disinhibitory circuit mediates motor integration in the somatosensory cortex. Nat. Neurosci. 16, 1662–1670 (2013).
O’Connor, D. H., Peron, S. P., Huber, D. & Svoboda, K. Neural activity in barrel cortex underlying vibrissa-based object localization in mice. Neuron 67, 1048–1061 (2010).
Sriram, B., Li, L.,Cruz-Martin, A., & Ghosh, A. A sparse probabilistic code underlies the limits of behavioral discrimination. Preprint at bioRxiv https://doi.org/10.1101/424713 (2018).
Chavarha, M. et al. Fast two-photon volumetric imaging of an improved voltage indicator reveals electrical activity in deeply located neurons in the awake brain. Preprint at bioRxiv https://doi.org/10.1101/445064 (2018).
Kannan, M. et al. Fast, in vivo voltage imaging using a red fluorescent indicator. Nat. Methods 15, 1108–1116 (2018).
Harris, J. A. et al. Anatomical characterization of Cre driver mice for neural circuit mapping and manipulation. Front. Neural Circuits 8, 76 (2014).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Ringach, D. L., Shapley, R. M. & Hawken, M. J. Orientation selectivity in macaque V1: diversity and laminar dependence. J. Neurosci. 22, 5639–5651 (2002).
Shen, X., Tokoglu, F., Papademetris, X. & Constable, R. T. Groupwise whole-brain parcellation from resting-state fMRI data for network node identification. Neuroimage 82, 403–415 (2013).
Acknowledgements
The authors thank all members of the Multiscale Imaging of Spontaneous Activity in Cortex collaboration at Yale University for input during all stages of this project. We thank X. Ge, E. Lake, L. Tang, D. Scheinost and E. Mohns for input on data analysis, K. Zhang for assistance with data preprocessing, Y. Zhang for assistance with animal husbandry and maintenance and members of the Cardin, Constable, Crair and Higley laboratories for helpful comments during the preparation of this manuscript. We thank D. Kim and the GENIE Project (Janelia Research Campus) for GCaMP6s and GCaMP6f plasmids. We thank H. Zeng (Allen Institute) for the TIGRE2 transgenic mice. This work was supported by funding from the NIH (grant nos. MH099045 to M.J.H., EY022951 to J.A.C., NS094358 to M.C.C. and R.T.C., EY026878 to M.C.C., MH111424 to R.T.C. and M.C.C., EY029581 to D.B., GM007205 to D.B., NS007224 to D.B., EY028869 to A.S.H.). M.C.C. was supported by the William Ziegler III family.
Author information
Authors and Affiliations
Contributions
All authors contributed to the overall study design. D.B., A.S.H. and J.A.C. collected the data. D.B., A.S.H., X.S. and J.A.C. analyzed the data. D.B., A.S.H., X.S., M.C.C. and M.J.H. wrote the manuscript. R.T.C., J.A.C., M.C.C. and M.J.H. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nina Vogt was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Two-photon imaging resolution through a glass microprism.
a, Example images of fluorescent beads imaged using either a 1.0 NA water objective through cover glass (Zeiss, left), an 0.4 NA air objective through cover glass (Mitutoyo, center), or the same 0.4 NA objective through the microprism (right). b, Point-spread function sizes, measured as the full-width at half-maximum amplitude, for the three imaging approaches shown in (a). Lateral (left) and axial (right) data are shown. Lateral mean±SEM: water/CG: 0.81 ± 0.01, air/CG: 1.08 ± 0.01, air/prism: 1.00 ± 0.02; axial mean±SEM: water/CG: 6.76 ± 0.21, air/CG: 9.55 ± 0.36, air/prism: 13.67 ± 0.35; n = 22, 24, and 20 beads, respectively. c, Distribution of signal-to-noise ratios for cells imaged under the same conditions. Mean±SEM: water/CG: 0.72 ± 0.02, air/CG: 0.87 ± 0.02, air/prism: 1.05 ± 0.02, p < 0.001 for all comparisons, Kolmogorov-Smirnov test, n = 142, 131, and 113 cells, respectively. d, Distribution of orientation selectivity indexes for cells imaged under the same conditions. Mean±SEM: water/CG: 0.38 ± 0.02, air/CG: 0.34 ± 0.02, air/prism: 0.34 ± 0.02, p > 0.05 for all comparisons, Kolmogorov-Smirnov test, n = 142, 131, and 113 cells, respectively.
Supplementary Figure 2 Comparison of methods for extracting fluorescent time-series.
a, Example field of view showing GCaMP6f-expressing neurons imaged via two-photon microscopy. Scale bar is 20 µm b, Distribution of neurons (regions of interest, ROIs) identified following rigid motion correction with manual selection (left) or non-rigid motion correction with automated selection (right). Color-matched cells are shown in (a). c, Relationship between correlations of single neuron time series versus correlations of CCNs, for the two different methods of ROI selection, for 176 matched cells in 1 mouse. d, Relationship between t-SNE 1 and t-SNE 2 values for all CCNs, illustrating that maps identified following either manual or automated ROI selection are similarly distributed, for 238 cells identified manually and 244 cells identified by Suite2p in 1 mouse. Colors are as in (b).
Supplementary Figure 3 Analysis of tissue damage beneath an implanted microprism.
a, Example coronal section of mouse brain, stained for Hsp70/72, a marker for cell damage. The prism was implanted over the right hemisphere (dashed box). Scale bar is 0.5 mm. b, Population data showing no change in Hsp70/72 expression relative to the control hemisphere for four animals.
Supplementary Figure 4 Comparison of parcellation methods with sensory-evoked activity.
Three example mice are shown, with activity evoked by whisker deflection (red) or auditory stimulation (blue) illustrated. Parcellations highlighted in white are for either the Allen CCFv3 atlas (upper images) or functional parcellation derived from spontaneous activity in the same animal (lower images). Scale bar is 2 mm.
Supplementary Figure 5 Additional examples of functional neuronal connectivity with cortical networks.
a, Illustration of the functional parcellation with regions labelled based on correspondence with the anatomical parcellation. Color code as in Fig. 3e. b, Activity index calculated from all significance maps for a single animal using the functional parcellation. c, Averages from the 3 clusters in (b). d, Schematized two-photon field-of-view with individual cells colored by cluster membership. Scale bar is 20 μm. e-h, Same as (a-d), but for a different mouse.
Supplementary Figure 6 Example outlier cell-centered networks for the clusters in Fig. 3.
Example CCNs displayed with the activity index of functionally-defined parcels from the same mouse shown in Fig. 3. The clusters to which these cells belong are indicated to the top left of each CCN. CCNs are roughly grouped based on which anatomical regions are most active, indicated to the left.
Supplementary Figure 7 Comparison of methods for calculating cortex-wide network membership.
a, Schematic of the spike-triggered average map (STM) procedure16, adapted for two-photon data. A spike-triggered average is calculated for all frames in a 2 second window around each ‘spike.’ For our analysis, we threshold each cell’s spike probability at four standard deviations above 0 (or a minimum of 0.2) to derive spike times. The STM is the maximum value for each pixel within the 2 second spike-triggered average. Permutation testing is performed by calculating a null distribution of STMs using 1000 random shuffles of spike times. Non-significant (p>0.05) pixels are transparent. b, Schematic of the spike-triggered average map (SpAM) procedure15, adapted for two-photon data. SpAMs are calculated by taking the partial correlation of cell ΔF/F and each mesoscopic pixel ΔF/F with respect to mean ΔF/F for all pixels across cortex. P values for each correlation are Benjamini-Hochberg corrected for multiple comparisons (5% false positive rate). Non-significant (p>0.05) pixels are transparent. c, CCN for the same cell (cell 3 from Fig. 3) as in (a, b). Non-significant (p > 0.05) pixels are transparent. d, Correlation between CCNs, SpAMs, and STMs for all analyzed cells (same as in Fig. 3) across 3 mice. Box-and-whisker plots of cell correlations show median, interquartile, and 5th-95th percentile values.
Supplementary Figure 8 Viral or transgenic expression of GCaMP6 does not disrupt cortical or hippocampal physiology.
a, Mean firing rate of regular spiking, putative pyramidal neurons in the visual cortex for the three cohorts of mice (wild-type mean±SEM: 1.1±0.2, n=3 mice, AAV9-hSyn-GCaMP6s mean±SEM: 1.2±0.2, n=3 mice, Slc17a7-Cre;CaMKIIa-tTa;TITL-GCaMP6f mean±SEM: 1.6±0.3, n=3 mice). b, Example local field potential (LFP) recordings from the three cohorts. c, Normalized LFP power spectra for the three cohorts, measured in visual cortex (upper traces) or hippocampus (lower traces). Shaded areas indicate SEM. N = 3 mice for each group. d, Overlaid power spectra from (c).
Supplementary Figure 9 Additional examples of CCNs varying by neuronal class and sensitivity to arousal.
a, Activity index calculated from all significance maps for all cells recorded in a single animal. Cells are clustered into six groups. Columns to the right indicate cell type (red indicates VIP-IN, black indicates pyramidal cell) and whether cells are correlated with whisking (colors as in Fig. 5d). b, Averages of the six clusters in (a) with parcels colored by their activity index. c, Fractional distribution of cells into each cluster, separated by type or modulation by whisking. d-f, g-i, Same as (a-c) for two additional mice.
Supplementary Figure 10 Two broad motifs for cortical network organization.
Schematic illustration showing the general patterns of long-range connectivity for cells in the primary somatosensory cortex as determined by simultaneous two-photon and mesoscopic imaging. Individual neurons tend to belong to either a fronto-lateral or postero-medial network, with both categories also involving retrosplenial areas.
Supplementary Figure 11 CCNs for VIP-INs and pyramidal cells are stable across days.
a, Schematized two-photon field-of-view of cells imaged over multiple days. Red indicates VIP-INs and black indicates pyramidal cells. Lighter shades indicate cells that were active in only one imaging session. Scale bar is 20 μm. b, CCNs for pyramidal cells and VIP-IN indicated by arrows in (a) over two days. c, Pearson’s correlation of CCNs calculated for individual pyramidal cells and VIP-INs from data collected over two days.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Table 1.
Rights and permissions
About this article
Cite this article
Barson, D., Hamodi, A.S., Shen, X. et al. Simultaneous mesoscopic and two-photon imaging of neuronal activity in cortical circuits. Nat Methods 17, 107–113 (2020). https://doi.org/10.1038/s41592-019-0625-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0625-2
This article is cited by
-
Multimodal measures of spontaneous brain activity reveal both common and divergent patterns of cortical functional organization
Nature Communications (2024)
-
Global spatiotemporal synchronizing structures of spontaneous neural activities in different cell types
Nature Communications (2024)
-
Rapid fluctuations in functional connectivity of cortical networks encode spontaneous behavior
Nature Neuroscience (2024)
-
The Cousa objective: a long-working distance air objective for multiphoton imaging in vivo
Nature Methods (2024)
-
Simulations approaching data: cortical slow waves in inferred models of the whole hemisphere of mouse
Communications Biology (2023)