Abstract
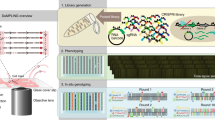
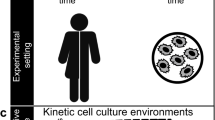
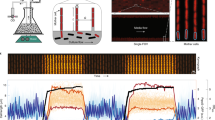
Single-cell genetic screens can be incredibly powerful, but current high-throughput platforms do not track dynamic processes, and even for non-dynamic properties they struggle to separate mutants of interest from phenotypic outliers of the wild-type population. Here we introduce SIFT, single-cell isolation following time-lapse imaging, to address these limitations. After imaging and tracking individual bacteria for tens of consecutive generations under tightly controlled growth conditions, cells of interest are isolated and propagated for downstream analysis, free of contamination and without genetic or physiological perturbations. This platform can characterize tens of thousands of cell lineages per day, making it possible to accurately screen complex phenotypes without the need for barcoding or genetic modifications. We applied SIFT to identify a set of ultraprecise synthetic gene oscillators, with circuit variants spanning a 30-fold range of average periods. This revealed novel design principles in synthetic biology and demonstrated the power of SIFT to reliably screen diverse dynamic phenotypes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding authors upon request.
Code availability
ImageJ macros for image segmentation and MATAB scripts for cell tracking and signal processing are available from the corresponding authors upon request.
References
Schmitz, J., Noll, T. & Grünberger, A. Heterogeneity studies of mammalian cells for bioproduction: from tools to application. Trends Biotechnol. 37, 645–660 (2019).
Daugherty, P. S., Iverson, B. L. & Georgiou, G. Flow cytometric screening of cell-based libraries. J. Immunol. Methods 243, 211–227 (2000).
Kim, H. S., Guzman, A. R., Thapa, H. R., Devarenne, T. P. & Han, A. A droplet microfluidics platform for rapid microalgal growth and oil production analysis. Biotechnol. Bioeng. 113, 1691–1701 (2016).
Han, Y., Gu, Y., Zhang, A. C. & Lo, Y.-H. Review: imaging technologies for flow cytometry. Lab Chip 16, 4639–4647 (2016).
Denervaud, N. et al. A chemostat array enables the spatio-temporal analysis of the yeast proteome. Proc. Natl. Acad. Sci. USA 110, 15842–15847 (2013).
Emanuel, G., Moffitt, J. R. & Zhuang, X. High-throughput, image-based screening of pooled genetic-variant libraries. Nat. Methods 14, 1159–1162 (2017).
Lawson, M. J. et al. In situ genotyping of a pooled strain library after characterizing complex phenotypes. Mol. Syst. Biol. 13, 947 (2017).
Wang, P. et al. Robust growth of Escherichia coli. Curr. Biol. 20, 1099–1103 (2010).
Potvin-Trottier, L., Lord, N. D., Vinnicombe, G. & Paulsson, J. Synchronous long-term oscillations in a synthetic gene circuit. Nature 538, 514–517 (2016).
Park, J. et al. Molecular time sharing through dynamic pulsing in single cells. Cell Syst. 6, 216–229 (2018).
Lugagne, J.-B. et al. Balancing a genetic toggle switch by real-time feedback control and periodic forcing. Nat. Commun. 8, 1671 (2017).
Taheri-Araghi, S., Bradde, S., Vergassola, M. & Jun, S. Cell-size control and homeostasis in bacteria. Curr. Biol. 25, 385–391 (2015).
Bergmiller, T. et al. Biased partitioning of the multidrug efflux pump AcrAB-TolC underlies long-lived phenotypic heterogeneity. Science 356, 311–315 (2017).
Robert, L. et al. Mutation dynamics and fitness effects followed in single cells. Science 359, 1283–1286 (2018).
Norman, T. M., Lord, N. D., Paulsson, J. & Losick, R. Memory and modularity in cell-fate decision making. Nature 503, 481–486 (2013).
Kaiser, M. et al. Monitoring single-cell gene regulation under dynamically controllable conditions with integrated microfluidics and software. Nat. Commun. 9, 212 (2018).
Unger, M. A., Chou, H. P., Thorsen, T., Scherer, A. & Quake, S. R. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000).
Ashkin, A., Dziedzic, J. M. & Yamane, T. Optical trapping and manipulation of single cells using infrared laser beams. Nature 330, 769–771 (1987).
Wakamoto, Y., Umehara, S., Matsumura, K., Inoue, I. & Yasuda, K. Development of non-destructive, non-contact single-cell based differential cell assay using on-chip microcultivation and optical tweezers. Sens. Actuators B Chem. 96, 693–700 (2003).
Probst, C., Grünberger, A., Wiechert, W. & Kohlheyer, D. Microfluidic growth chambers with optical tweezers for full spatial single-cell control and analysis of evolving microbes. J. Microbiol. Methods 95, 470–476 (2013).
Stricker, J. et al. A fast, robust and tunable synthetic gene oscillator. Nature 456, 516–519 (2008).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Ayano, S., Wakamoto, Y., Yamashita, S. & Yasuda, K. Quantitative measurement of damage caused by 1064-nm wavelength optical trapping of Escherichia coli cells using on-chip single cell cultivation system. Biochem. Biophys. Res. Commun. 350, 678–684 (2006).
Dirla, S., Chien, J. Y.-H. & Schleif, R. Constitutive mutations in the Escherichia coli AraC protein. J. Bacteriol. 191, 2668–2674 (2009).
Wang, Z., Xiang, L., Shao, J., Wegrzyn, A. & Wegrzyn, G. Effects of the presence of ColE1 plasmid DNA in Escherichia coli on the host cell metabolism. Microb. Cell Fact. 5, 34 (2006).
Berens, C., Altschmied, L. & Hillen, W. The role of the N terminus in Tet repressor for tet operator binding determined by a mutational analysis. J. Biol. Chem. 267, 1945–1952 (1992).
Farrell, C. M., Grossman, A. D. & Sauer, R. T. Cytoplasmic degradation of ssrA-tagged proteins. Mol. Microbiol. 57, 1750–1761 (2005).
Schaerli, Y. & Isalan, M. Building synthetic gene circuits from combinatorial libraries: screening and selection strategies. Mol. Biosyst. 9, 1559–1567 (2013).
Bashor, C. J. & Collins, J. J. Understanding biological regulation through synthetic biology. Annu. Rev. Biophys. 47, 399–423 (2018).
Levine, J. H., Lin, Y. & Elowitz, M. B. Functional roles of pulsing in genetic circuits. Science 342, 1193–1200 (2013).
Riglar, D. T. et al. Bacterial variability in the mammalian gut captured by a single-cell synthetic oscillator. Nat. Commun. 10, 4665 (2019).
Brewster, R. C. et al. The transcription factor titration effect dictates level of gene expression. Cell 156, 1312–1323 (2014).
Buchler, N. E. & Cross, F. R. Protein sequestration generates a flexible ultrasensitive response in a genetic network. Mol. Syst. Biol. 5, 272 (2009).
Betz, J. L. Cloning and characterization of several dominant-negative and tight-binding mutants of lac repressor. Gene 42, 283–292 (1986).
Teng, S.-W., Mukherji, S., Moffitt, J. R., de Buyl, S. & O’Shea, E. K. Robust circadian oscillations in growing cyanobacteria require transcriptional feedback. Science 340, 737–740 (2013).
Yuan, A. H. & Hochschild, A. A bacterial global regulator forms a prion. Science 355, 198–201 (2017).
Balleza, E., Kim, J. M. & Cluzel, P. Systematic characterization of maturation time of fluorescent proteins in living cells. Nat. Methods 15, 47–51 (2018).
Landgraf, D., Okumus, B., Chien, P., Baker, T. A. & Paulsson, J. Segregation of molecules at cell division reveals native protein localization. Nat. Methods 9, 480–482 (2012).
Spivey, E. C., Xhemalce, B., Shear, J. B. & Finkelstein, I. J. 3D-Printed microfluidic microdissector for high-throughput studies of cellular aging. Anal. Chem. 86, 7406–7412 (2014).
Li, Y. et al. Multigenerational silencing dynamics control cell aging. Proc. Natl. Acad. Sci. USA 114, 11253–11258 (2017).
Pearl Mizrahi, S., Gefen, O., Simon, I. & Balaban, N. Q. Persistence to anti-cancer treatments in the stationary to proliferating transition. Cell Cycle 15, 3442–3453 (2016).
Eriksson, E. et al. Optical manipulation and microfluidics for studies of single cell dynamics. J. Opt. A Pure Appl. Opt. 9, S113–S121 (2007).
Okumus, B. et al. Single-cell microscopy of suspension cultures using a microfluidics-assisted cell screening platform. Nat. Protoc. 13, 170–194 (2017).
Lu, Y., Flaherty, C. & Hendrickson, W . AraC protein contacts asymmetric sites in the Escherichia coli araFGH promoter. J. Biol. Chem. 267, 24848–24857 (1992).
Betz, J. L., Sasmor, H. M., Buck, F., Insley, M. Y. & Caruthers, M. H. Base substitution mutants of the lac operator: in vivo and in vitro affinities for lac repressor. Gene 50, 123–132 (1986).
Espah Borujeni, A., Channarasappa, A. S. & Salis, H. M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res 42, 2646–2659 (2014).
Madsen, M. H., Feidenhans’l, N. A., Hansen, P.-E., Garnæs, J. & Dirscherl, K. Accounting for PDMS shrinkage when replicating structures. J. Micromech. Microeng. 24, 127002 (2014).
Block, S. M. Construction of Optical Tweezers. in Cells: A Laboratory Manual - Vol. II, Sec. 7 (Cold Spring Harbor Laboratory Press, 1997).
Lee, W. M., Reece, P. J., Marchington, R. F., Metzger, N. K. & Dholakia, K. Construction and calibration of an optical trap on a fluorescence optical microscope. Nat. Protoc. 2, 3226–3238 (2007).
Neuman, K. C., Chadd, E. H., Liou, G. F., Bergman, K. & Block, S. M. Characterization of photodamage to Escherichia coli in optical traps. Biophys. J. 77, 2856–2863 (1999).
Seemann, T. Snippy: rapid haploid variant calling and core genome alignment. (2015).
Altshuler, D. et al. An SNP map of the human genome generated by reduced representation shotgun sequencing. Nature 407, 513–516 (2000).
Acknowledgements
We thank R. Yuan for substantial help designing figure illustrations and S. Bakshi for sharing core image-analysis software and providing technical microscopy guidance. All photolithography was performed at the Center for Nanoscale Systems at Harvard University, a member of the National Nanotechnology Coordinated Infrastructure Network, which is supported by the National Science Foundation under award 1541959. All soft lithography was performed at the Microfluidics/Microfabrication Core Facility at Harvard Medical School. This work was supported by the Defense Advanced Research Projects Agency (HR0011-16-2-0049), National Science Foundation (1615487), and National Institutes of Health (R01GM081563).
Author information
Authors and Affiliations
Contributions
S.L., L.P.-T., B.O. and J.P. conceived the study. S.L. designed and fabricated the screening chip, constructed and screened the genetic libraries, and performed the data analysis. S.L. and J.P wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
S.L. and J.P. are listed as inventors on a provisional patent application covering the SIFT technology.
Additional information
Peer review information Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Phenotypic classification following transport via optical trapping.
The trapped cell cohort was transported by optical trap for a range of times (Fig. 3d) and re-seeded as new mother cells in empty growth trenches. Trapped ancestors were mother cells within the same growth trench as transported cells were taken from. Cells that were not trapped included all other mother cells that were imaged during the recovery but were neither themselves nor their progeny exposed to the optical trap. General phenotypic classifications include mother cell death (no growth observed over a 12-h recovery and monitoring period), filamentation, undocumented loss (mother cell absence without observing an abnormal increase of cell length in the prior frame) and continual growth. Similar results were observed across three independent screening runs.
Supplementary Figure 2 Dual-feedback library amplitude to period noise relationship.
Statistical estimates of period CV and amplitude CV of all library variants with oscillatory-like signals over a 24-h period using SIFT (Nosc = 7,803). Variants selected to highlight phenotypic diversity (grey dots with roman numerals) have time-traces shown in Fig. 4c. Red circles represent variants that were isolated by SIFT, some of which were individually characterized in a follow-up mother machine run. Mean generation time was 25 min. Data were gathered from a single screening run.
Supplementary Figure 3 Sequences of mutated regions for all isolated cells from the dual-feedback library.
Numbering with ‘SL’ prefixes mark cells collected from the same lineage (that is, isolated from the same growth trench). Letterings A–C designate individual cells taken from the same clonal lineages. Three individual cells were isolated from each lineage, unless fewer than three cells were present at the time of isolation (for example, cells lost due to filamentation prior to collection). Mutated regulatory regions included araI1, the I1 binding site of AraC; lacO1, the binding site of LacI; and the ribosomal-binding site (RBS). The consensus sequence is shown at the top of each table. The top panel shows promoter variants controlling araC, while the bottom panel corresponds to lacI regulation. Asterisks denote agreement with non-degenerate sequences and dashes signify deletions.
Supplementary Figure 4 Timing performance of dual-feedback oscillators.
Top panel shows period histograms of the original (SL126) and top-performing screened (SL278) dual-feedback circuits without chemical inducers. 6,170 periods are displayed from the original circuit and 3,054 periods from the isolate. Lower panel displays the autocorrelation functions (ACF) of the circuits from the top panel. Given the low signal-to-noise and limited fluctuation amplitudes of the original dual-feedback oscillator (see Fig. 4d), the autocovariance function (ACVF; not normalized by variance) is also shown for each of the two circuits in the inset.
Supplementary Figure 5 Timing performance of other dual-feedback oscillators.
Top panel shows period histograms of the original dual-feedback oscillator (SL126) with the optimized drug combination (0.7% arabinose, 2 mM IPTG); the circuit with constitutive araC mutation (Y13H; SL304) without drugs; and the top-performing screened (isolate; SL278) circuit without drugs. Histograms include 2,830 periods from the original circuit, 7,149 periods from the Y13H circuit, and 3,054 periods from the isolated circuit. Bottom panel displays the autocorrelation functions (ACF) of the circuits from the top panel.
Supplementary Figure 6 Period ranges of oscillator libraries.
All three libraries with period CV and mean period plotted on the same axes. The dominant-negative repressilator library with protein degradation (that is, with ClpXP present) was only characterized and was not screened. Variants with period CVs (CVT) below 20% are qualitatively deemed regular oscillators. Scatter plot displays 7,803 dual-feedback variants with degradation, 3,437 dominant-negative repressilator variants with degradation and 1,277 dominant-negative repressilator variants without degradation. Data for each library were gathered from a single screening run.
Supplementary Figure 7 Dominant-negative repressilator library amplitude to period noise relationship.
Statistical estimates of period CV and amplitude CV of all library variants with oscillatory-like signals over a 24-h period using SIFT (Nosc = 1,277). Variants selected to highlight phenotypic diversity (grey dots with roman numerals) have time-traces shown in Fig. 5d. Red circles represent variants that were isolated, some of which were individually characterized in a follow-up mother machine run. Mean generation time was 24 min. Data were gathered from a single screening run.
Supplementary Figure 8 Sequences of mutated regions for all isolated cells from the dominant-negative repressilator library.
Numbering with ‘SL’ prefixes mark cells collected from the same lineage (that is, isolated from the same growth trench). Letterings A–C designate individual cells taken from the same clonal lineages. Three individual cells were isolated from each lineage, unless fewer than three cells were present at the time of isolation (for example, cells lost due to filamentation prior to collection). Mutated regulatory regions included the –35 and –10 hexamers of the core promoter elements and the ribosomal-binding site (RBS) controlling expression of dominant-negative TetR. The consensus sequence is shown at the top of the table. Asterisks denote agreement with non-degenerate sequences and dashes signify deletions.
Supplementary Figure 9 Period CVs as a function of sampled periods.
The mother machine data for characterizing a top-performing dominant-negative isolate (SL229 with a true period CV of 0.099) were parsed by individual time-traces and not pooled. Each trench-wise CV calculation was then sampled for 2 to 10 consecutive periods (only considering traces with 10 or more periods present; 141 traces). The trend in the CV variance as a function of the number of sampled periods highlights the chance of getting ‘false’ outlier CVs when just one lineage is monitored for oscillators with extremely long periods, as was the case for the dominant-negative repressilator screen, stressing the need for isolation followed by thorough characterization of these variants.
Supplementary Figure 10 Timing performance of an isolated dominant-negative repressilator with long-term oscillations.
Top panel shows period histograms of the original integrated repressilator (SL305) and a dominant-negative repressilator library isolate with lengthy coherent oscillations (SL224). Histograms include 8,161 periods from the original circuit and 886 periods from the isolated circuit. Bottom panel displays the autocorrelation functions (ACF) of the circuits from the top panel.
Supplementary information
Supplementary Information
Supplementary Figures 1–10
Supplementary Video 1
Cell transport for screening. Three cells (third transit contains two cells stuck together) were individually transported via optical trapping from a single culture trench in the growth lane, moved across the perforated median (collection valves in open state) and dropped off in the collection lane. The movie was captured at 100× optical magnification and compiled at 15 frames per second. Cell transfers were performed more than 200 times across 5 independent screening runs with similar results.
Supplementary Video 2
Cell transport for re-seeding. Three cells were individually transported via optical trapping from a single culture trench in the growth lane, across the perforated median (collection valves in open state), and seeded within three new culture trenches in the collection lane. The movie was captured at 100× optical magnification and compiled at 15 frames per second. Cell re-seedings within collection lanes were performed more than 10 times across three independent screening runs with similar results.
Rights and permissions
About this article
Cite this article
Luro, S., Potvin-Trottier, L., Okumus, B. et al. Isolating live cells after high-throughput, long-term, time-lapse microscopy. Nat Methods 17, 93–100 (2020). https://doi.org/10.1038/s41592-019-0620-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0620-7
This article is cited by
-
Emerging tools for uncovering genetic and transcriptomic heterogeneities in bacteria
Biophysical Reviews (2024)
-
An optogenetic toolkit for light-inducible antibiotic resistance
Nature Communications (2023)
-
Synthetic Micrographs of Bacteria (SyMBac) allows accurate segmentation of bacterial cells using deep neural networks
BMC Biology (2022)
-
Cell region fingerprints enable highly precise single-cell tracking and lineage reconstruction
Nature Methods (2022)
-
Capture and reagent exchange (CARE) wells for cell isolation, labeling, and characterization
Microfluidics and Nanofluidics (2022)