Abstract
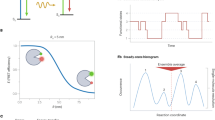
The applications of Förster resonance energy transfer (FRET) grow with each year. However, different FRET techniques are not applied consistently, nor are results uniformly presented, which makes implementing and reproducing FRET experiments challenging. We discuss important considerations for designing and evaluating ensemble FRET experiments. Alongside a primer on FRET basics, we provide guidelines for making experimental design choices such as the donor-acceptor pair, instrumentation and labeling chemistries; selecting control experiments to unambiguously demonstrate FRET and validate that the experiments provide meaningful data about the biomolecular process in question; analyzing raw data and assessing the results; and reporting data and experimental details in a manner that easily allows for reproducibility. Some considerations are also given for FRET assays and FRET imaging, especially with fluorescent proteins. Our goal is to motivate and empower all biologists to consider FRET for the powerful research tool it can be.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Web of Science (Clarivate Analytics, accessed 3 June 2019); https://apps.webofknowledge.com
Medintz, I. L. & Hildebrandt, N. (eds) FRET – Förster Resonance Energy Transfer from Theory to Applications (Wiley-VCH, 2013).
McNutt, M. Journals unite for reproducibility. Science 346, 679–679 (2014).
Lerner, E. et al. Toward dynamic structural biology: two decades of single-molecule Forster resonance energy transfer. Science 359, eaan1133 (2018).
Hellenkamp, B. et al. Precision and accuracy of single-molecule FRET measurements—a multi-laboratory benchmark study. Nat. Meth. 15, 669–676 (2018).
Kalinin, S. et al. A toolkit and benchmark study for FRET-restrained high-precision structural modeling. Nat. Meth. 9, 1218–1225 (2012).
Stryer, L. & Haugland, R. P. Energy transfer – a spectroscopic ruler. Proc. Natl. Acad. Sci. USA 58, 719–726 (1967).
Marx, V. Probes: FRET sensor design and optimization. Nat. Methods 14, 949–953 (2017).
Ai, H. W., Hazelwood, K. L., Davidson, M. W. & Campbell, R. E. Fluorescent protein FRET pairs for ratiometric imaging of dual biosensors. Nat. Methods 5, 401–403 (2008).
Greenwald, E. C., Mehta, S. & Zhang, J. Genetically encoded fluorescent biosensors illuminate the spatiotemporal regulation of signaling networks. Chem. Rev. 118, 11707–11794 (2018).
Miyawaki, A. & Niino, Y. Molecular spies for bioimaging-fluorescent protein-based probes. Mol. Cell 58, 632–643 (2015).
Bajar, B. T. et al. Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Sci. Rep. 6, 20889 (2016).
Hochreiter, B., Garcia, A. P. & Schmid, J. A. Fluorescent proteins as genetically encoded FRET biosensors in life sciences. Sensors 15, 26281–26314 (2015).
Sahl, S. J., Hell, S. W. & Jakobs, S. Fluorescence nanoscopy in cell biology. Nat. Rev. Mol. Cell Bio. 18, 685–701 (2017).
Boeneman, K. et al. Quantum dot DNA bioconjugates: attachment chemistry strongly influences the resulting composite architecture. ACS Nano. 4, 7253–7266 (2010).
Sapsford, K. E., Berti, L. & Medintz, I. L. Materials for fluorescence resonance energy transfer: beyond traditional ‘dye to dye’ combinations. Angew. Chem. Int. Ed. 45, 4562–4588 (2006).
Sapsford, K. E., Tyner, K. M., Dair, B. J., Deschamps, J. R. & Medintz, I. L. Analyzing nanomaterial bioconjugates: a review of current and emerging purification and characterization techniques. Anal. Chem. 83, 4453–4488 (2011).
Hermanson, G. T. (ed.) Bioconjugate Techniques 3rd edn (Acad. Press, 2013).
Byrne, A. G. et al. in FRET - Förster Resonance Energy Transfer from Theory to Applications (eds Medintz, I. L. & Hildebrandt, N.) 657–756 (Wiley-VCH, 2013).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy (Springer, 2006).
Bhuckory, S., Lefebvre, O., Qiu, X., Wegner, K. D. & Hildebrandt, N. Evaluating quantum dot performance in homogeneous FRET immunoassays for prostate specific antigen. Sensors 16, 197 (2016).
Wegner, K. D. et al. Three-dimensional solution-phase Förster resonance energy transfer analysis of nanomolar quantum dot bioconjugates with subnanometer resolution. Chem. Mat. 26, 4299–4312 (2014).
Gorris, H. H. & Resch-Genger, U. Perspectives and challenges of photon-upconversion nanoparticles - Part II: bioanalytical applications. Anal. Bioanal. Chem. 409, 5875–5890 (2017).
Wurth, C., Grabolle, M., Pauli, J., Spieles, M. & Resch-Genger, U. Relative and absolute determination of fluorescence quantum yields of transparent samples. Nat. Protoc. 8, 1535–1550 (2013).
Stennett, E. M. S., Ma, N., van der Vaart, A. & Levitus, M. Photophysical and dynamical properties of doubly linked Cy3-DNA constructs. J. Phys. Chem. B 118, 152–163 (2014).
Melinger, J. S. et al. FRET from multiple pathways in fluorophore-labeled DNA. ACS Photonics 3, 659–669 (2016).
Seidel, C. A. M., Schulz, A. & Sauer, M. H. M. Nucleobase-specific quenching of fluorescent dyes. 1. Nucleobase one-electron redox potentials and their correlation with static and dynamic quenching efficiencies. J. Phys. Chem. 100, 5541–5553 (1996).
Wozniak, A. K., Schroder, G. F., Grubmuller, H., Seidel, C. A. M. & Oesterhelt, F. Single-molecule FRET measures bends and kinks in DNA. Proc. Natl. Acad. Sci. USA 105, 18337–18342 (2008).
Van der Meer, B. W., Van der Meer, D. M. & Vogel, S. S. in FRET - Förster Resonance Energy Transfer: From Theory to Applications (eds Medintz, I. L. & Hildebrandt, N.) Ch. 4 (Wiley-VCH, 2013).
Dos Santos, M. C. & Hildebrandt, N. Recent developments in lanthanide-to-quantum dot FRET using time-gated fluorescence detection and photon upconversion. Trac. Trends Anal. Chem. 84, 60–71 (2016).
Hildebrandt, N., Wegner, K. D. & Algar, W. R. Luminescent terbium complexes: superior Forster resonance energy transfer donors for flexible and sensitive multiplexed biosensing. Coord. Chem. Rev. 273, 125–138 (2014).
Wild, D. The Immunoassay Handbook 3rd edn (Elsevier, 2013).
Wegner, K. D., Jin, Z. W., Linden, S., Jennings, T. L. & Hildebrandt, N. Quantum-dot-based Forster resonance energy transfer immunoassay for sensitive clinical diagnostics of low-volume serum samples. ACS Nano 7, 7411–7419 (2013).
Jalink, K. & Rheenen, J.v. in Laboratory Techniques in Biochemistry and Molecular Biology Vol. 33. (ed. Gadella, T. W. J.) 289–349 (Elsevier, 2009).
Chen, H., Puhl, H. L., Koushik, S. V., Vogel, S. S. & Ikeda, S. R. Measurement of FRET efficiency and ratio of donor to acceptor concentration in living cells. Biophys. J. 91, L39–L41 (2006).
Hoppe, A., Christensen, K. & Swanson, J. A. Fluorescence resonance energy transfer-based stoichiometry in living cells. Biophys. J. 83, 3652–3664 (2002).
Zal, T. & Gascoigne, N. R. J. Photobleaching-corrected FRET efficiency imaging of live cells. Biophys. J. 86, 3923–3939 (2004).
Vogel, S. S., Blank, P. S., Koushik, S. V. & Thaler, C. in Laboratory Techniques in Biochemistry and Molecular Biology (ed. Gadella, T. W. J.) (Elsevier, 2009).
Neher, R. A. & Neher, E. Applying spectral fingerprinting to the analysis of FRET images. Microsc. Res. Tech. 64, 185–195 (2004).
Thaler, C., Koushik, S. V., Blank, P. S. & Vogel, S. S. Quantitative multiphoton spectral imaging and its use for measuring resonance energy transfer. Biophys. J. 89, 2736–2749 (2005).
Wlodarczyk, J. et al. Analysis of FRET signals in the presence of free donors and acceptors. Biophys. J. 94, 986–1000 (2008).
Zimmermann, T., in Advances in Biochemical Engineering-Biotechnology (ed. Rietdorf, J.) 245–265 (Springer Verlag, 2005).
Bastiaens, P. I. H. & Squire, A. Fluorescence lifetime imaging microscopy: spatial resolution of biochemical processes in the cell. Trends Cell Bio. 9, 48–52 (1999).
Chen, Y. E. & Periasamy, A. Characterization of two-photon excitation fluorescence lifetime imaging microscopy for protein localization. Microsc. Res. Tech. 63, 72–80 (2004).
Wallrabe, H. & Periasamy, A. Imaging protein molecules using FRET and FLIM microscopy. Curr. Opin. Biotech. 16, 19–27 (2005).
Wouters, F. S. & Bastiaens, P. I. H. Fluorescence lifetime imaging of receptor tyrosine kinase activity in cells. Curr. Bio. 9, 1127–1130 (1999).
Kenworthy, A. K. Imaging protein-protein interactions using fluorescence resonance energy transfer microscopy. Methods 24, 289–296 (2001).
Bader, A. N., Hofman, E. G., Voortman, J., Henegouwen, P. & Gerritsen, H. C. Homo-FRET imaging enables quantification of protein cluster sizes with subcellular resolution. Biophys. J. 97, 2613–2622 (2009).
Clayton, A. H. A., Hanley, Q. S., Arndt-Jovin, D. J., Subramaniam, V. & Jovin, T. M. Dynamic fluorescence anisotropy imaging microscopy in the frequency domain (rFLIM). Biophys. J. 83, 1631–1649 (2002).
Lidke, D. S. et al. Imaging molecular interactions in cells by dynamic and static fluorescence anisotropy (rFLIM and emFRET). Biochem. Soc. Trans. 31, 1020–1027 (2003).
Tramier, M. & Coppey-Moisan, M. Fluorescence anisotropy imaging microscopy for homo-FRET in living cells. Meth. Cell Bio. 85, 395–414 (2008).
Yeow, E. K. L. & Clayton, A. H. A. Enumeration of oligomerization states of membrane proteins in living cells by homo-FRET spectroscopy and microscopy: theory and application. Biophys. J. 92, 3098–3104 (2007).
Gadella, T. W. J. (ed.) FRET and FLIM Techniques 1st edn (Elsevier Science, 2008).
Periasamy, A., Mazumder, N., Sun, Y., Christopher, K. G. & Day, R. N. in Advanced Time-Correlated Single Photon Counting Applications (ed. W. Becker) 249–276 (Springer, 2015).
Kim, Y. et al. Venus(A206) dimers behave coherently at room temperature. Biophys. J. 116, 1918–1930 (2019).
Vogel, S. S., Nguyen, T. A., van der Meer, B. W. & Blank, P. S. The impact of heterogeneity and dark acceptor states on FRET: Implications for using fluorescent protein donors and acceptors. PLoS ONE 7, e49593 (2012).
Zacharias, D. A., Violin, J. D., Newton, A. C. & Tsien, R. Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Chandrasekhar, S. Stochastic problems in physics and astronomy. Rev. Mod. Phys. 15, 1–89 (1943).
Anikovsky, M., Dale, L., Ferguson, S. & Petersen, N. Resonance energy transfer in cells: a new look at fixation effect and receptor aggregation on cell membrane. Biophys. J. 95, 1349–1359 (2008).
King, C., Sarabipour, S., Byrne, P., Leahy, D. J. & Hristova, K. The FRET signatures of noninteracting proteins in membranes: Simulations and experiments. Biophys. J. 106, 1309–1317 (2014).
Loura, L. M. S. & Prieto, M. FRET in membrane biophysics: an overview. Front. Physiol. 2, 82 (2011).
Ma, Y. Q. et al. An intermolecular FRET sensor detects the dynamics of T cell receptor clustering. Nat. Comm. 8, 15100 (2017).
Humpolickova, J. et al. Inhibition of the precursor and mature forms of HIV-1 protease as a tool for drug evaluation. Sci. Rep. 8, 10438 (2018).
Dou, J. Y. et al. De novo design of a fluorescence-activating beta-barrel. Nature 561, 485–491 (2018).
Mastop, M. et al. Characterization of a spectrally diverse set of fluorescent proteins as FRET acceptors for mTurquoise2. Sci. Rep. 7, 11999 (2017).
Murakoshi, H. & Shibata, A. C. E. ShadowY: a dark yellow fluorescent protein for FLIM-based FRET measurement. Sci. Rep. 7, 6791 (2017).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Piatkevich, K. D. & Verkhusha, V. V. in Recent Advances in Cytometry, Part A: Instrumentation, Methods 5th edn (eds Darzynkiewicz, Z. et al.) 431–461 (2011).
Lam, A. J. et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nat. Methods 9, 1005–1012 (2012).
Abraham, B. G. et al. Fluorescent protein based FRET pairs with improved dynamic range for fluorescence lifetime measurements. PLoS ONE 10, e0134436 (2015).
Bajar, B. T., Wang, E. S., Zhang, S., Lin, M. Z. & Chu, J. A guide to fluorescent protein FRET pairs. Sensors 16, E1488 (2016).
Zipfel, W. R., Williams, R. M. & Webb, W. W. Nonlinear magic: multiphoton microscopy in the biosciences. Nat. Biotechnol. 21, 1368–1376 (2003).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Bio. 190, 165–175 (2010).
Valeur, B. & Berberan-Santos, M. N. (eds) Molecular Fluorescence: Principles and Applications (Wiley-VCH, 2012).
Clegg, R. M. & Sener, M., Govindjee. From Forster resonance energy transfer to coherent resonance energy transfer and back. Opt. Biopsy 7561, 75610C (2010).
Clegg, R. M. in FRET and FLIM Techniques (ed. Gadella, T. W. J.) Ch. 1 (Elsevier, 2009).
Sapsford, K. E., Wildt, B., Mariani, A., Yeatts, A. B. & Medintz, I. L. in FRET – Förster Resonance Energy Transfer From Theory to Applications (eds Medintz, I. L. & Hildebrandt, N.) 165–268 (Wiley-VCH, 2013).
Jameson, D. M. Introduction to Fluorescence (CRC Press, 2014).
Cranfill, P. J. et al. Quantitative assessment of fluorescent proteins. Nat. Methods 13, 557–562 (2016).
Fritz, R. D. et al. A versatile toolkit to produce sensitive FRET biosensors to visualize signaling in time and space. Sci. Signal. 6, rs12 (2013).
Acknowledgements
This Perspective originates from the International Discussion Meeting on Förster Resonance Energy Transfer (FRET) in the Life Sciences II 2016 (http://fret.uni-duesseldorf.de/cms/), held at the Max Planck Institute for Biophysical Chemistry in Göttingen, Germany — where Förster first developed his seminal FRET theory. I.L.M. acknowledges the Office of Naval Research, the NRL Nanosciences Institute and a LUCI project in support of the VBFF through the OSD. N.H. acknowledges the Institut Universitaire de France (IUF). W.R.A. acknowledges the Natural Sciences and Engineering Research Council (NSERC) of Canada, the Canada Foundation for Innovation (CFI), BCKDF, a Canada Research Chair (Tier 2), a Michael Smith Foundation for Health Research Scholar Award and an Alfred P. Sloan Foundation Research Fellowship. S.S.V. acknowledges funding by the intramural program of the National Institutes of Health, National Institute on Alcohol Abuse and Alcoholism, Bethesda, MD. The mention of commercial and other websites in this article does not constitute any endorsement by the authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests
Additional information
Peer review information: Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–4, Supplementary Tables 1–4 and Supplementary Notes 1–33
Rights and permissions
About this article
Cite this article
Algar, W.R., Hildebrandt, N., Vogel, S.S. et al. FRET as a biomolecular research tool — understanding its potential while avoiding pitfalls. Nat Methods 16, 815–829 (2019). https://doi.org/10.1038/s41592-019-0530-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0530-8
This article is cited by
-
Assessment of the FRET-based Teen sensor to monitor ERK activation changes preceding morphological defects in a RASopathy zebrafish model and phenotypic rescue by MEK inhibitor
Molecular Medicine (2024)
-
Sequestration within peptide coacervates improves the fluorescence intensity, kinetics, and limits of detection of dye-based DNA biosensors
Communications Chemistry (2024)
-
Multi-step FRET systems based on discrete supramolecular assemblies
Communications Chemistry (2024)
-
Decoding the Cellular Trafficking of Prion-like Proteins in Neurodegenerative Diseases
Neuroscience Bulletin (2024)
-
Utilizing dual-pathway energy transfer in upconversion nanoconjugates for reinforced photodynamic therapy
Nano Research (2024)