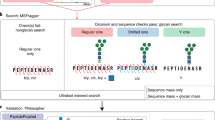
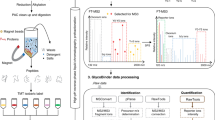
Abstract
We report a liquid chromatography coupled to tandem mass spectrometry O-glycoproteomics strategy using data-independent acquisition (DIA) mode for direct analysis of O-glycoproteins. This approach enables characterization of glycopeptides and structures of O-glycans on a proteome-wide scale with quantification of stoichiometries (though it does not allow for direct unambiguous glycosite identification). The method relies on a spectral library of O-glycopeptides; the Glyco-DIA library contains sublibraries obtained from human cell lines and human serum, and it currently covers 2,076 O-glycoproteins (11,452 unique glycopeptide sequences) and the 5 most common core1 O-glycan structures. Applying the Glyco-DIA library to human serum without enrichment for glycopeptides enabled us to identify and quantify 269 distinct glycopeptide sequences bearing up to 5 different core1 O-glycans from 159 glycoproteins in a SingleShot analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All of the Glyco-DIA libraries are available on our O-Glycoproteome database (http://glycoproteomics.somee.com) and https://github.com/CCGMS/Glyco-DIA, and can be directly imported to Spectronaut. All of the raw data including DDA and DIA runs have been deposited to the ProteomeXchange Consortium35, via the PRIDE partner repository, with the data set identifier PXD011063.
Code availability
All of the R scripts are available at https://github.com/CCGMS/Glyco-DIA.
References
Goth, C. K. et al. A systematic study of modulation of ADAM-mediated ectodomain shedding by site-specific O-glycosylation. Proc. Natl Acad. Sci. USA 112, 14623–14628 (2015).
Ohtsubo, K. & Marth, J. D. Glycosylation in cellular mechanisms of health and disease. Cell 126, 855–867 (2006).
Levery, S. B. et al. Advances in mass spectrometry driven O-glycoproteomics. Biochim. Biophys. Acta 1850, 33–42 (2015).
Darula, Z., Sherman, J. & Medzihradszky, K. F. How to dig deeper? Improved enrichment methods for mucin core-1 type glycopeptides. Mol. Cell. Proteom. 11, 016774 (2012). mcp. O111.
King, S. L. et al. Characterizing the O-glycosylation landscape of human plasma, platelets, and endothelial cells. Blood Adv. 1, 429–442 (2017).
Medzihradszky, K. F., Kaasik, K. & Chalkley, R. J. Tissue-specific glycosylation at the glycopeptide level. Mol. Cell. Proteom. 14, 050393 (2015). mcp. M115.
Hoffmann, M., Marx, K., Reichl, U., Wuhrer, M. & Rapp, E. Site-specific O-glycosylation analysis of human blood plasma proteins. Mol. Cell. Proteom. 15, 624–641 (2016).
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through simple cell technology. EMBO J. 32, 1478–1488 (2013).
Hintze, J. et al. Probing the contribution of individual polypeptide GalNAc-transferase isoforms to the O-glycoproteome by inducible expression in isogenic cell lines. J. Biol. Chem. 293, 19064–19077 (2018).
Venable, J. D., Dong, M.-Q., Wohlschlegel, J., Dillin, A. & Yates, J. R. III Automated approach for quantitative analysis of complex peptide mixtures from tandem mass spectra. Nat. Methods 1, 39 (2004).
Chapman, J. D., Goodlett, D. R. & Masselon, C. D. Multiplexed and data-independent tandem mass spectrometry for global proteome profiling. Mass Spectrom. Rev. 33, 452–470 (2014).
Gillet, L. C. et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol. Cell. Proteom. 11, O111.016717 (2012).
Ludwig, C. et al. Data‐independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Mol. Syst. Biol. 14, e8126 (2018).
Liu, Y. et al. Glycoproteomic analysis of prostate cancer tissues by SWATH mass spectrometry discovers N-acylethanolamine acid amidase and protein tyrosine kinase 7 as signatures for tumor aggressiveness. Mol. Cell. Proteom. 13, 1753–1768 (2014).
Pan, K.-T., Chen, C.-C., Urlaub, H. & Khoo, K.-H. Adapting data-independent acquisition for mass spectrometry-based protein site-specific N-glycosylation analysis. Anal. Chem. 89, 4532–4539 (2017).
Couto, N., Davlyatova, L., Evans, C. A. & Wright, P. C. Application of the broadband collision-induced dissociation (bbCID) mass spectrometry approach for protein glycosylation and phosphorylation analysis. Rapid Commun. Mass Spectrom. 32, 75–85 (2018).
Zacchi, L. F. & Schulz, B. L. SWATH-MS glycoproteomics reveals consequences of defects in the glycosylation machinery. Mol. Cell. Proteom. 15, 2435–2447 (2016).
Lin, C.-H., Krisp, C., Packer, N. H. & Molloy, M. P. Development of a data independent acquisition mass spectrometry workflow to enable glycopeptide analysis without predefined glycan compositional knowledge. J. Proteom. 172, 68–75 (2018).
Steentoft, C. et al. Mining the O-glycoproteome using zinc-finger nuclease-glycoengineered simple cell lines. Nat. Methods 8, 977–982 (2011).
Bruderer, R. et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol. Cell. Proteom. 14, 1400–1410 (2015).
Furukawa, J.-i et al. Quantitative O-glycomics by microwave-assisted β-elimination in the presence of pyrazolone analogues. Anal. Chem. 87, 7524–7528 (2015).
Fujitani, N. et al. Total cellular glycomics allows characterizing cells and streamlining the discovery process for cellular biomarkers. Proc. Natl Acad. Sci. USA 110, 2105–2110 (2013).
Wuhrer, M., Catalina, M. I., Deelder, A. M. & Hokke, C. H. Glycoproteomics based on tandem mass spectrometry of glycopeptides. J. Chromatogr. B 849, 115–128 (2007).
Nilsson, J. Liquid chromatography-tandem mass spectrometry-based fragmentation analysis of glycopeptides. Glycoconj. J. 33, 261–272 (2016).
Rosenberger, G. et al. Statistical control of peptide and protein error rates in large-scale targeted data-independent acquisition analyses. Nat. Methods 14, 921–927 (2017).
Chalkley, R. J., Thalhammer, A., Schoepfer, R. & Burlingame, A. Identification of protein O-GlcNAcylation sites using electron transfer dissociation mass spectrometry on native peptides. Proc. Natl Acad. Sci. USA 106, 8894–8899 (2009).
Liu, Y. et al. Quantitative variability of 342 plasma proteins in a human twin population. Mol. Syst. Biol. 11, 786 (2015).
Ting, Y. S. et al. PECAN: library-free peptide detection for data-independent acquisition tandem mass spectrometry data. Nat. Methods 14, 903 (2017).
Darula, Z. & Medzihradszky, K. F. Carbamidomethylation side reactions may lead to glycan misassignments in glycopeptide analysis. Anal. Chem. 87, 6297–6302 (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Katrine, T.-B. S. et al. Probing isoform-specific functions of polypeptide GalNAc-transferases using zinc finger nuclease glycoengineered SimpleCells. Proc. Natl Acad. Sci. USA 109, 9893–9898 (2012).
R Core Team. R: A language and environment for statistical computing v3.5.1 (R, 2013); https://www.r-project.org/
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Reiter, L. et al. mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat. Methods 8, 430 (2011).
Vizcaíno, J. A. et al. ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 32, 223 (2014).
Acknowledgements
This work was supported by The Lundbeck Foundation, The Mizutani Foundation and the Danish National Research Foundation (DNRF107). We thank L. Kristensen and C. Dauly (Thermo Fisher Scientific) for their help with this study.
Author information
Authors and Affiliations
Contributions
Z.Y., Y.M., H.C. and S.Y.V. conceived and designed the study. Z.Y. and S.Y.V. contributed with experimental data and interpretation. Z.Y., H.C. and S.Y.V. wrote the manuscript. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Allison Doerr was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Comparison between Glyco-DIA libraries.
(a) Number of glycoproteins in Tn-DIA libraries from HEK293, HepG2, OVCAR3 and M3T4 SC lines. (b) Number of glycopeptides in Tn-DIA libraries from HEK293, HepG2, OVCAR3 and M3T4 SC lines. (c) Number of glycoproteins in T-DIA libraries from HEK293 WT and HepG2 WT cell lines and human blood serum. (d) Number of glycopeptides in T-DIA libraries from HEK293 WT and HepG2 WT cell lines and human blood serum.
Supplementary Figure 2 Comparison of glycopeptides in serum library with and without depletion of most abundant proteins.
Top 12 abundant proteins were depleted with spin columns (85164, Pierce, Thermo Scientific), including α1-acid glycoprotein, α1-antitrypsin, α1-macroglubulin, albumin, apolipoprotein A-I, apolipoprotein A-II, fibrinogen, haptoglobin, IgA, IgG, IgM, transferrin.
Supplementary Figure 3 Window width distribution of DIA method in Glyco-DIA.
The DIA method was designed based on the precursor distribution in HEK293 WT T-DIA library (histogram in the background). It covers precursor range from m/z 400 to m/z 1,200 and a total of 41 windows are listed in the method, including 3 windows of 100 m/z isolation width, 12 windows of 20 m/z isolation width, and 26 windows of 10 m/z isolation width.
Supplementary Figure 4
Comparison of glycopeptides in the HepG2 WT SSL and the comprehensive HepG2 T-DIA library.
Supplementary Figure 5
Comparison of identified glycopeptides in the HepG2 WT Jacalin enriched non-diluted sample with SSL and the HepG2 T-DIA library.
Supplementary Figure 6 ShapQualityScore distributions of sLWAC enriched HepG2 WT samples processed with the HepG2 T-DIA library.
A total of 541 T-glycopeptides were identified and 279 with ShapeQualityScore higher than 0.7.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6 and Supplementary Notes 1–8.
Supplementary Table 1
Overview of the generated spectral libraries and the figures they were used in
Supplementary Table 2
Combined list of all the identifications in the SingleShot analysis
Supplementary Data Set 1
ETD-MS2 spectra and extracted ion chromatograms of all the positional isoforms from the ETD/HCD DDA run for the site-specific library.
Supplementary Data Set 2
Comparison of serum T-DIA library with published data, related to Supplementary Note 3.
Rights and permissions
About this article
Cite this article
Ye, Z., Mao, Y., Clausen, H. et al. Glyco-DIA: a method for quantitative O-glycoproteomics with in silico-boosted glycopeptide libraries. Nat Methods 16, 902–910 (2019). https://doi.org/10.1038/s41592-019-0504-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0504-x
This article is cited by
-
Prediction of glycopeptide fragment mass spectra by deep learning
Nature Communications (2024)
-
Hybrid-DIA: intelligent data acquisition integrates targeted and discovery proteomics to analyze phospho-signaling in single spheroids
Nature Communications (2023)
-
Oxonium ion scanning mass spectrometry for large-scale plasma glycoproteomics
Nature Biomedical Engineering (2023)
-
Recent trends in glycoproteomics by characterization of intact glycopeptides
Analytical and Bioanalytical Chemistry (2023)
-
Glycoproteomics
Nature Reviews Methods Primers (2022)