Abstract
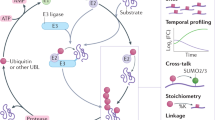
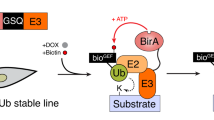
Ubiquitin (Ub) conjugation is an essential post-translational modification that affects nearly all proteins in eukaryotes. The functions and mechanisms of ubiquitination are areas of extensive study, and yet the dynamics and regulation of even free (that is, unconjugated) Ub are poorly understood. A major impediment has been the lack of simple and robust techniques to quantify Ub levels in cells and to monitor Ub release from conjugates. Here, we describe avidity-based fluorescent sensors that address this need. The sensors bind specifically to free Ub, have dissociation constant Kd values down to 60 pM and, together with a newly developed workflow, allow us to distinguish and quantify the pools of free, protein-conjugated and thioesterified forms of Ub from cell lysates. Alternatively, free Ub in fixed cells can be visualized microscopically by staining with a sensor. Real-time assays using the sensors afford unprecedented flexibility and precision to measure deubiquitination of virtually any (poly)Ub conjugate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this paper are available from the corresponding author on reasonable request.
References
Komander, D. & Rape, M. The ubiquitin code. Annu Rev. Biochem. 81, 203–229 (2012).
Park, C. W. & Ryu, K. Y. Cellular ubiquitin pool dynamics and homeostasis. BMB Rep. 47, 475–482 (2014).
Ryu, K. Y., Garza, J. C., Lu, X. Y., Barsh, G. S. & Kopito, R. R. Hypothalamic neurodegeneration and adult-onset obesity in mice lacking the Ubb polyubiquitin gene. Proc. Natl Acad. Sci. USA 105, 4016–4021 (2008).
Ryu, K. Y. et al. The mouse polyubiquitin gene UbC is essential for fetal liver development, cell-cycle progression and stress tolerance. EMBO J. 26, 2693–2706 (2007).
Ryu, K. Y. et al. The mouse polyubiquitin gene Ubb is essential for meiotic progression. Mol. Cell Biol. 28, 1136–1146 (2008).
Kimura, Y. et al. An inhibitor of a deubiquitinating enzyme regulates ubiquitin homeostasis. Cell 137, 549–559 (2009).
Wang, C. H. et al. USP5/Leon deubiquitinase confines postsynaptic growth by maintaining ubiquitin homeostasis through Ubiquilin. eLife 6, e26886 (2017).
Crimmins, S. et al. Transgenic rescue of ataxia mice with neuronal-specific expression of ubiquitin-specific protease 14. J. Neurosci. 26, 11423–11431 (2006).
Chen, P. C. et al. The proteasome-associated deubiquitinating enzyme Usp14 is essential for the maintenance of synaptic ubiquitin levels and the development of neuromuscular junctions. J. Neurosci. 29, 10909–10919 (2009).
Oh, C., Park, S., Lee, E. K. & Yoo, Y. J. Downregulation of ubiquitin level via knockdown of polyubiquitin gene Ubb as potential cancer therapeutic intervention. Sci. Rep. 3, 2623 (2013).
Kedves, A. T. et al. Recurrent ubiquitin B silencing in gynecological cancers establishes dependence on ubiquitin C. J. Clin. Invest 127, 4554–4568 (2017).
Hallengren, J., Chen, P. C. & Wilson, S. M. Neuronal ubiquitin homeostasis. Cell Biochem. Biophys. 67, 67–73 (2013).
Dantuma, N. P., Groothuis, T. A., Salomons, F. A. & Neefjes, J. A dynamic ubiquitin equilibrium couples proteasomal activity to chromatin remodeling. J. Cell Biol. 173, 19–26 (2006).
Yau, R. & Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 18, 579–586 (2016).
Kaiser, S. E. et al. Protein standard absolute quantification (PSAQ) method for the measurement of cellular ubiquitin pools. Nat. Methods 8, 691–696 (2011).
Reyes-Turcu, F. E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208 (2006).
Swanson, K. A., Kang, R. S., Stamenova, S. D., Hicke, L. & Radhakrishnan, I. Solution structure of Vps27 UIM-ubiquitin complex important for endosomal sorting and receptor downregulation. EMBO J. 22, 4597–4606 (2003).
Lee, S. et al. Structural basis for ubiquitin recognition and autoubiquitination by Rabex-5. Nat. Struct. Mol. Biol. 13, 264–271 (2006).
Penengo, L. et al. Crystal structure of the ubiquitin binding domains of rabex-5 reveals two modes of interaction with ubiquitin. Cell 124, 1183–1195 (2006).
Maupin-Furlow, J. A. Ubiquitin-like proteins and their roles in archaea. Trends Microbiol. 21, 31–38 (2013).
Choi, Y. S., Jeon, Y. H., Ryu, K. S. & Cheong, C. 60th residues of ubiquitin and Nedd8 are located out of E2-binding surfaces, but are important for K48 ubiquitin-linkage. FEBS Lett. 583, 3323–3328 (2009).
Zhang, D., Raasi, S. & Fushman, D. Affinity makes the difference: nonselective interaction of the UBA domain of Ubiquilin-1 with monomeric ubiquitin and polyubiquitin chains. J. Mol. Biol. 377, 162–180 (2008).
Sokratous, K. et al. Probing affinity and ubiquitin linkage selectivity of ubiquitin-binding domains using mass spectrometry. J. Am. Chem. Soc. 134, 6416–6424 (2012).
Swaney, D. L., Rodriguez-Mias, R. A. & Villen, J. Phosphorylation of ubiquitin at Ser65 affects its polymerization, targets, and proteome-wide turnover. EMBO Rep. 16, 1131–1144 (2015).
Kane, L. A. et al. PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J. Cell Biol. 205, 143–153 (2014).
Ordureau, A. et al. Defining roles of PARKIN and ubiquitin phosphorylation by PINK1 in mitochondrial quality control using a ubiquitin replacement strategy. Proc. Natl Acad. Sci. USA 112, 6637–6642 (2015).
Koyano, F. et al. Ubiquitin is phosphorylated by PINK1 to activate parkin. Nature 510, 162–166 (2014).
Harper, J. W., Ordureau, A. & Heo, J. M. Building and decoding ubiquitin chains for mitophagy. Nat. Rev. Mol. Cell Biol. 19, 93–108 (2018).
Wauer, T. et al. Ubiquitin Ser65 phosphorylation affects ubiquitin structure, chain assembly and hydrolysis. EMBO J. 34, 307–325 (2015).
Dang, L. C., Melandri, F. D. & Stein, R. L. Kinetic and mechanistic studies on the hydrolysis of ubiquitin C-terminal 7-amido-4-methylcoumarin by deubiquitinating enzymes. Biochemistry 37, 1868–1879 (1998).
Geurink, P. P. et al. Development of diubiquitin-based FRET probes to quantify ubiquitin linkage specificity of deubiquitinating enzymes. Chem. Bio. Chem. 17, 816–820 (2016).
Yao, T. & Cohen, R. E. Ubiquitin-ovomucoid fusion proteins as model substrates for monitoring degradation and deubiquitination by proteasomes. Methods Enzym. 398, 522–540 (2005).
Wang, T. et al. Evidence for bidentate substrate binding as the basis for the K48 linkage specificity of otubain 1. J. Mol. Biol. 386, 1011–1023 (2009).
Wiener, R. et al. E2 ubiquitin-conjugating enzymes regulate the deubiquitinating activity of OTUB1. Nat. Struct. Mol. Biol. 20, 1033–1039 (2013).
Gates, Z. P., Stephan, J. R., Lee, D. J. & Kent, S. B. Rapid formal hydrolysis of peptide-alphathioesters. Chem. Commun. 49, 786–788 (2013).
Shahnawaz, M., Thapa, A. & Park, I. S. Stable activity of a deubiquitylating enzyme (Usp2-cc) in the presence of high concentrations of urea and its application to purify aggregation-prone peptides. Biochem. Biophys. Res. Commun. 359, 801–805 (2007).
Kim, W. et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell 44, 325–340 (2011).
Yang, X. et al. Absolute quantification of E1, ubiquitin-like proteins and Nedd8-MLN4924 adduct by mass spectrometry. Cell Biochem. Biophys. 67, 139–147 (2013).
Chen, J. J. et al. Mechanistic studies of substrate-assisted inhibition of ubiquitin-activating enzyme by adenosine sulfamate analogues. J. Biol. Chem. 286, 40867–40877 (2011).
Bianchi, M. et al. Dynamic transcription of ubiquitin genes under basal and stressful conditions and new insights into the multiple UBC transcript variants. Gene 573, 100–109 (2015).
Liu, Z. et al. Noncovalent dimerization of ubiquitin. Angew. Chem. Int. Ed. Engl. 51, 469–472 (2012).
Joo, H. Y. et al. Regulation of cell cycle progression and gene expression by H2A deubiquitination. Nature 449, 1068–1072 (2007).
Jencks, W. P. On the attribution and additivity of binding energies. Proc. Natl Acad. Sci. USA 78, 4046–4050 (1981).
Scott, D., Oldham, N. J., Strachan, J., Searle, M. S. & Layfield, R. Ubiquitin-binding domains: mechanisms of ubiquitin recognition and use as tools to investigate ubiquitin-modified proteomes. Proteomics 15, 844–861 (2015).
Hicke, L., Schubert, H. L. & Hill, C. P. Ubiquitin-binding domains. Nat. Rev. Mol. Cell Biol. 6, 610–621 (2005).
Morrow, M. E. et al. Active site alanine mutations convert deubiquitinases into high-affinity ubiquitin-binding proteins. EMBO Rep. 19, e45680 (2018).
Harrigan, J. A., Jacq, X., Martin, N. M. & Jackson, S. P. Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat. Rev. Drug Disco. 17, 57–78 (2018).
Majetschak, M. Extracellular ubiquitin: immune modulator and endogenous opponent of damage-associated molecular pattern molecules. J. Leukoc. Biol. 89, 205–219 (2011).
Raasi, S. & Pickart, C. M. Ubiquitin chain synthesis. Methods Mol. Biol. 301, 47–55 (2005).
Whitby, F. G., Xia, G., Pickart, C. M. & Hill, C. P. Crystal structure of the human ubiquitin-like protein NEDD8 and interactions with ubiquitin pathway enzymes. J. Biol. Chem. 273, 34983–34991 (1998).
Wilkinson, K. D. et al. Metabolism of the polyubiquitin degradation signal: structure, mechanism, and role of isopeptidase T. Biochemistry 34, 14535–14546 (1995).
Wilkinson, K. D. Quantitative analysis of protein–protein interactions. Methods Mol. Biol. 261, 15–32 (2004).
Pasupala, N. et al. OTUB1 non-catalytically stabilizes the E2 ubiquitin-conjugating enzyme UBE2E1 by preventing its autoubiquitination. J. Biol. Chem. 293, 18285–18295 (2018).
Acknowledgements
We thank A. Ma (University of California, San Francisco) for MEF cells, C. Wolberger (Johns Hopkins University) for OTUB1 protein and B. Brasher (Boston Biochem) for phosphoubiquitin. We also thank O. Peersen for assistance with the rapid-kinetics experiments and use of the Bio-Logic stopped-flow spectrofluorometer, and R. Handa for use of the Imaris image analysis software. This research was supported by NIH-NIGMS grant no. R01 GM115997 (to R.E.C.) and NIH-NIEHS grant no. R21 ES029150 (to R.E.C. and T.Y.).
Author information
Authors and Affiliations
Contributions
Y.S.C. and R.E.C. conceived and designed the ubiquitin sensor reagents. Y.S.C. produced the sensors and characterized them in vitro. S.A.B., T.Y. and R.E.C. conceived the cell-based studies, which were done by S.A.B., L.F.P. and F.S. All authors contributed to writing or commenting on the manuscript.
Corresponding author
Ethics declarations
Competing interests
U.S. patent no. 10,018,634 has been awarded to Colorado State University Research Foundation (R.E.C. and Y.C., inventors) for ubiquitin sensors and assays described in this paper.
Additional information
Peer review information: Rita Strack was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Primary sequences of the free Ub sensors.
The bold, underlined residues indicate mutated residues, and the cysteines in red indicate fluorophore conjugation sites. The residues highlighted in cyan are linkers introduced to connect the domains, and the yellow and green highlighted sequences show peptides added to provide 6-His and HA-epitope tags, respectively.
Supplementary Figure 2 Sensor binding assays.
a, Atto532-Ub(S20C) titrated with tIVR. b, Atto532-tIVR titrated with Ub. c, Alexa488-Ub(S20C) titrated to tISR. d, tISR competition binding assay performed by titration of Ub in presence of Alexa488-Ub(S20C). In each titration, the fluorescence intensity change from ligand binding was measured. Error bars in a, which are mostly hidden by the point symbols, show standard deviations of the assay repeated two times independently; titrations in b - d were performed once. Kd values ± s.d. shown in each panel were determined from non-linear fits to the data.
Supplementary Figure 3 Screening for an optimal fluorescent dye and conjugation site on the tIVR sensor.
a, Cys residue in the linker connecting IsoTBuz and Vps27UIM; b, N46C; and c, M105C in IsoTBuz were selected to conjugate various fluorescent dyes. In each panel, the arrow shows the conjugation site on the tIVR model and the histograms show performance with different fluorophores installed at that site. To determine which site and fluorescent dye gave the largest fluorescence change upon Ub binding, fluorescence intensities of the labeled sensors (2 nM) were measured with and without saturating Ub. F0 is fluorescence intensity of the labeled sensors without Ub, and ∆F is the intensity change upon addition of 200 µM Ub.
Supplementary Figure 4 Kinetics of Atto532-tUI binding to Ub.
a, To determine the dissociation rate of the Atto532-tUI•Ub complex, Atto532-tUI (1.0 nM) was pre-incubated at 25 °C with 1.0 nM Ub, and then excess unlabeled tUI (1 µM) was rapidly added and the fluorescence was measured every 0.25 s using a stopped-flow fluorimeter. Three independent reactions were performed, and all the data were fit with a single-exponential decay model to determine koff. The experimental data (blue) are shown superimposed with the fitted curve (red). b, The change in fluorescence intensity of 50 pM Atto532-tUI immediately after addition of 0.5, 1.0, or 2.0 nM Ub was monitored at 25 °C. The fluorescence was measured every 0.25 s. From each curve, an observed rate (kobs) was determined from the best-fitting exponential curve, and from these kon was determined from the equation kobs = kon * [Ub] + koff. Each reaction to generate the association data was performed once. c, The experimentally determined association and dissociation rates ± s.d. for the Atto532-tUI•Ub complex are shown together with the Kd ± s.d. calculated from their ratio ± s.d.
Supplementary Figure 5 Analysis of the DUB reaction products by SDS-PAGE.
Samples from the end-points of the real-time DUB reactions in Fig. 2 were separated by SDS-PAGE together with the following control samples: Atto532-tIVR in lane 1, Ub5-OM(LY) in lane 2, Ub1-OM(LY) in lane 3, and the Usp2cc-digested product in lane 7. Fluorescent gel bands were detected with a Typhoon laser scanner using excitation at either (a) 488 nm, (b) 532 nm, or (c) both to visualize Lucifer Yellow-labeled ovomucoid protein (OM(LY)) or Atto532-labeled tIVR. d, The bar graph compares the Ub released in the DUB reactions in Fig. 2 quantified by either the protein band fluorescence (panel a) or the sensor fluorescence (Fig. 2). e, Atto532-tIVR (2 nM) was titrated with Ub (2–32 nM) and the data were fit with a 1:1 binding model as described in Methods. The fitted equation was used to convert fluorescence intensities from the real-time DUB reaction curves (Fig. 2, left panel) to free Ub concentrations (Fig. 2, right panel).
Supplementary Figure 6 Generation of Ub-hydrazide from Ub thioesters.
a, SDS-PAGE was used to monitor E2-Ub thioester formation and subsequent hydrazinolysis. Ub was incubated with E1 and E2 (UbcH5c) enzymes without (lane 1) or with (lane 2) ATP to form Ub~UbcH5c thioester. After treatment with NEM (lane 3) to inactivate enzymatic activities, hydrazine was added at the concentrations indicated; lanes 4–6 show that hydrazinolysis appeared to be complete (i.e., E2-Ub thioester was gone) under all three conditions. b, Upper scheme shows the reaction to generate Ub-hydrazide (see Methods) used in Fig. 1b-d titrations. Lower panel shows overlayed MALDI-TOF mass spectra of Ub and products (i.e., Ub–MESNA and Ub–hydrazide) from each step in the synthesis. The results in a and b are representative of 3 independent experiments.
Supplementary Figure 7 Processing cell lysates with Usp2cc releases most conjugated Ub.
SDS-PAGE and immunoblotting with anti-Ub antibody (P4G7) shows the levels of conjugated Ub in HeLa cell lysates prepared as described in the Methods for in-solution measurements of Ub pools. The lysate was generated and incubated without or with Usp2cc for 1 h at 37 °C as described in Methods. This analysis was performed once.
Supplementary Figure 8 Ub pools in proteasome-inhibited HeLa cells do not change significantly after 1 h.
In-solution quantification of Ub pools in lysates after treatment with vehicle (DMSO) or the proteasome inhibitor, BTZ (1 µM). Statistical analysis of the sample mean was by one-way ANOVA with Bonferroni’s adjustment; error bars represent ± s.d. (n = 3).
Supplementary Figure 9 Competition by free Ub for cell staining by HA-tUI.
Quantitation of mean fluorescence in HeLa cells (maximum projection images) after incubation with HA-tUI with or without excess free Ub. For 100 nM HA-tUI, n = 131; for 100 nM HA-tUI + 100 μM Ub, n = 28. AU, arbitrary units; error bars indicate mean ± s.d. Statistical analysis used two-tailed unpaired Student's t-test with Welch's correction.
Supplementary Figure 10 Free Ub staining by HA-tUI shows a different subcellular distribution than K48-linked polyUb or total conjugated Ub.
HeLa cells were fixed with 4% PFA and stained with DAPI (blue), anti-K48 polyUb antibody (green), and HA-tUI (magenta) in panel a, or DAPI (blue), anti-Ub FK2 antibody (green), and HA-tUI (magenta) in panel b. For each cell, the images are from a single z-stack plane that shows both nucleus and cytoplasm. The cells shown are representative of more than 10 cells analyzed on each of two coverslips.
Supplementary Figure 11 Competition binding assays to measure tUI affinity for Ub.
Competition binding assays were performed with a, 70 pM Atto532-tUI or b, 50 pM Atto532-tUI in the presence of 80 pM Ub. tUI was titrated from 0.063 nM to 32 nM, from which a mean Ki of 194 pM was determined from fittings to the two independent data sets.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11
Rights and permissions
About this article
Cite this article
Choi, YS., Bollinger, S.A., Prada, L.F. et al. High-affinity free ubiquitin sensors for quantifying ubiquitin homeostasis and deubiquitination. Nat Methods 16, 771–777 (2019). https://doi.org/10.1038/s41592-019-0469-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0469-9