Abstract
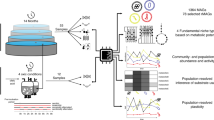
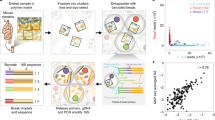
Metagenomic sequencing has enabled detailed investigation of diverse microbial communities, but understanding their spatiotemporal variability remains an important challenge. Here, we present decomposition of variance using replicate sampling (DIVERS), a method based on replicate sampling and spike-in sequencing. The method quantifies the contributions of temporal dynamics, spatial sampling variability, and technical noise to the variances and covariances of absolute bacterial abundances. We applied DIVERS to investigate a high-resolution time series of the human gut microbiome and a spatial survey of a soil bacterial community in Manhattan’s Central Park. Our analysis showed that in the gut, technical noise dominated the abundance variability for nearly half of the detected taxa. DIVERS also revealed substantial spatial heterogeneity of gut microbiota, and high temporal covariances of taxa within the Bacteroidetes phylum. In the soil community, spatial variability primarily contributed to abundance fluctuations at short time scales (weeks), while temporal variability dominated at longer time scales (several months).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequencing data are available at NCBI SRA under PRJNA541083.
Code availability
MATLAB scripts to perform all variance and covariance decomposition analyses from original OTU abundance tables are available on GitHub at https://github.com/brianwji/DIVERS. Implementation of DIVERS in R is available on GitHub at https://github.com/hym0405/DIVERS.
References
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Thompson, L. R. et al. A communal catalogue reveals earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded human microbiome project. Nature 550, 61–68 (2017).
Hunt, D. E. et al. Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science 320, 1081–1085 (2008).
Faust, K., Lahti, L., Gonze, D., de Vos, W. M. & Raes, J. Metagenomics meets time series analysis: unraveling microbial community dynamics. Curr. Opin. Microbiol. 25, 56–66 (2015).
Martin-Platero, A. M. et al. High resolution time series reveals cohesive but short-lived communities in coastal plankton. Nat. Commun. 9, 266 (2018).
Vandeputte, D. et al. Quantitative microbiome profiling links gut community variation to microbial load. Nature 551, 507–511 (2017).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Stammler, F. et al. Adjusting microbiome profiles for differences in microbial load by spike-in bacteria. Microbiome 4, 28 (2016).
Tkacz, A., Hortala, M. & Poole, P. S. Absolute quantitation of microbiota abundance in environmental samples. Microbiome 6, 110 (2018).
Zmora, N. et al. Personalized gut mucosal colonization resistance to empiric probiotics is associated with unique host and microbiome features. Cell 174, 1388–1405 e1321 (2018).
Gohl, D. M. et al. Systematic improvement of amplicon marker gene methods for increased accuracy in microbiome studies. Nat. Biotechnol. 34, 942–949 (2016).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Faust, K. & Raes, J. Microbial interactions: from networks to models. Nat. Rev. Microbiol 10, 538–550 (2012).
Sonnenburg, E. D. et al. Diet-induced extinctions in the gut microbiota compound over generations. Nature 529, 212–215 (2016).
Sonnenburg, E. D. et al. Specificity of polysaccharide use in intestinal Bacteroides species determines diet-induced microbiota alterations. Cell 141, 1241–1252 (2010).
Rakoff-Nahoum, S., Foster, K. R. & Comstock, L. E. The evolution of cooperation within the gut microbiota. Nature 533, 255–259 (2016).
Rakoff-Nahoum, S., Coyne, M. J. & Comstock, L. E. An ecological network of polysaccharide utilization among human intestinal symbionts. Curr. Biol. 24, 40–49 (2014).
Ramirez, K. S. et al. Biogeographic patterns in below-ground diversity in New York City’s Central Park are similar to those observed globally. Proc. Biol. Sci. 281, 20141988 (2014).
O’Brien, S. L. et al. Spatial scale drives patterns in soil bacterial diversity. Environ. Microbiol. 18, 2039–2051 (2016).
Carini, P. et al. Unraveling the effects of spatial variability and relic DNA on the temporal dynamics of soil microbial communities. Preprint at https://www.biorxiv.org/content/10.1101/402438v1 (2018).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc. Natl Acad. Sci. USA 103, 626–631 (2006).
Contijoch, E. J. et al. Gut microbiota density influences host physiology and is shaped by host and microbial factors. eLife 8, e40553 (2019).
Wargo, J. A., Reddy, S. M., Reuben, A. & Sharma, P. Monitoring immune responses in the tumor microenvironment. Curr. Opin. Immunol. 41, 23–31 (2016).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Baym, M. et al. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS ONE 10, e0128036 (2015).
Kilpatrick, A. M. & Ives, A. R. Species interactions can explain Taylor’s power law for ecological time series. Nature 422, 65–68 (2003).
Grun, D., Kester, L. & van Oudenaarden, A. Validation of noise models for single-cell transcriptomics. Nat. Methods 11, 637–640 (2014).
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008).
Williams, C. R. CE Gaussian Processes for Machine Learning (MIT Press, 2006).
Sala, C. et al. Stochastic neutral modelling of the gut microbiota’s relative species abundance from next generation sequencing data. BMC Bioinforma. 17, S16 (2016).
Acknowledgements
H.H.W. acknowledges funding from the NIH (grant nos. R01AI132403, R01DK118044), Burroughs Wellcome Fund (no. PATH 1016691), Bill & Melinda Gates Foundation (no. INV-000609), and the Schaefer Research Scholars Program for this work. R.U.S. is supported by a Fannie and John Hertz Foundation Fellowship and a NSF Graduate Research Fellowship (no. DGE-1644869). B.W.J. is supported in part by the NIH under Ruth L. Kirschstein National Research Service Award (NRSA) Institutional Research Training Grant (no. T32GM007367) and by the MD-PhD program at Columbia University. D.V. acknowledges funding from the NIH (grant nos. R01GM079759, R01DK118044).
Author information
Authors and Affiliations
Contributions
B.W.J. and R.U.S. conceived the study, designed the data collection workflow and performed all data analysis. B.W.J. and P.D.D. developed the variance and covariance decomposition models. R.U.S. performed all experiments with assistance from A.K. Y.H. assisted with data analysis and code implementation in R. H.H.W. and D.V. oversaw the project, and guided experiments and data analysis. All authors wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nicole Rusk was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–16, Supplementary Tables 1–3 and Supplementary Note
Source data
Rights and permissions
About this article
Cite this article
Ji, B.W., Sheth, R.U., Dixit, P.D. et al. Quantifying spatiotemporal variability and noise in absolute microbiota abundances using replicate sampling. Nat Methods 16, 731–736 (2019). https://doi.org/10.1038/s41592-019-0467-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0467-y
This article is cited by
-
EMBED: Essential MicroBiomE Dynamics, a dimensionality reduction approach for longitudinal microbiome studies
npj Systems Biology and Applications (2023)
-
High-throughput microbial culturomics using automation and machine learning
Nature Biotechnology (2023)
-
An improved workflow for accurate and robust healthcare environmental surveillance using metagenomics
Microbiome (2022)
-
High-throughput identification and quantification of single bacterial cells in the microbiota
Nature Communications (2022)
-
Growth promotion and antibiotic induced metabolic shifts in the chicken gut microbiome
Communications Biology (2022)