Abstract
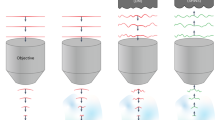
We advance two-photon microscopy for near-diffraction-limited imaging up to 850 µm below the pia in awake mice. Our approach combines direct wavefront sensing of light from a guidestar (formed by descanned fluorescence from Cy5.5-conjugated dextran in brain microvessels) with adaptive optics to compensate for tissue-induced aberrations in the wavefront. We achieve high signal-to-noise ratios in recordings of glutamate release from thalamocortical axons and calcium transients in spines of layer 5b basal dendrites during active tactile sensing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
The system aberration calculation, DM and SHWS calibration and sample-induced wavefront reconstruction and AO correction code are available in the Supplementary Software.
References
Svoboda, K., Denk, W., Kleinfeld, D. & Tank, D. W. Nature 385, 161–165 (1997).
Mittmann, W. et al. Nat. Neurosci. 14, 1089–1093 (2011).
Kondo, M., Kobayashi, K., Ohkura, M., Nakai, J. & Matsuzaki, M. eLife 6, e26839 (2017).
Ji, N. Nat. Methods 14, 374–380 (2017).
Tao, X. et al. Opt. Lett. 36, 1062–1064 (2011).
Wang, K. et al. Nat. Methods 11, 625–628 (2014).
Wang, K. et al. Nat. Commun. 6, 7276 (2015).
Ji, N., Milkie, D. E. & Betzig, E. Nat. Methods 7, 141–147 (2010).
Débarre, D. et al. Opt. Lett. 34, 2495–2497 (2009).
Tang, J., Germain, R. N. & Cui, M. Proc. Natl Acad. Sci. USA 109, 8434–8439 (2012).
Holtmaat, A. et al. Nat. Protoc. 4, 1128–1144 (2009).
Drew, P. J. et al. Nat. Methods 7, 981–984 (2010).
Dana, H. et al. eLife 5, e12727 (2016).
Marvin, J. S. et al. Nat. Methods 15, 936–939 (2018).
Oberlaender, M. et al. Cereb. Cortex 22, 2375–2391 (2011).
Castro-Alamancos, M. A. & Connors, B. W. Prog. Neurobiol. 51, 581–606 (1997).
Hopt, A. & Neher, E. Biophys. J. 80, 2029–2036 (2001).
Laissue, P. P., Alghamdi, R. A., Tomancak, P., Reynaud, E. G. & Shroff, H. Nat. Methods 14, 657–661 (2017).
Yildirim, M., Sugihara, H., So, P. T. C. & Sur, M. Nat. Commun. 11, 177 (2019).
Kleinfeld, D., Mitra, P. P., Helmchen, F. & Denk, W. Proc. Natl Acad. Sci. USA 95, 15741–15746 (1998).
Ouzounov, D. G. et al. Nat. Methods 14, 388–390 (2017).
Driscoll, J. D. et al. J. Neurophysiol. 104, 1803–1811 (2011).
Tsai, P. S. & Kleinfeld D. in Methods for In Vivo Optical Imaging 2nd edn (ed. Frostig, R. D.) 59–115 (CRC Press, 2009).
Noll, R. J. J. Opt. Soc. Am. 66, 207–211 (1979).
Thompson, R. E., Larson, D. R. & Webb, W. W. Biophys. J. 82, 2775–2783 (2002).
Cubalchini, R. J. Opt. Soc. Am. 69, 973–977 (1979).
Gong, S. et al. Nature 425, 917–925 (2003).
Dana, H. et al. PLoS One 13, e0205444 (2018).
Young, P. et al. Nat. Neurosci. 11, 721–728 (2008).
Morii, S., Ngai, A. C. & Winn, H. R. J. Cereb. Blood Flow Metab. 6, 34–41 (1986).
Shih, A. Y., Drew, P. J., Mateo, C., Kleinfeld, D. & Tsai, P. S. J. Vis. Exp. 61, e3742 (2012).
Lam, A. J. et al. Nat. Methods 9, 105–112 (2012).
Shih, A. Y. et al. J. Cereb. Blood Flow Metab. 32, 1277–1309 (2012).
Knutsen, P. M., Mateo, C. & Kleinfeld, D. Philos. Trans. R. Soc. B 371, 20150351 (2016).
Kleinfeld, D., Sachdev, R. N. S., Merchant, L. M., Jarvis, M. R. & Ebner, F. F. Neuron 34, 1021–1034 (2002).
Schindelin, J. et al. Nat. Methods 9, 676–682 (2012).
Guizar-Sicairos, M., Thurman, S. T. & Fienup, J. R. Opt. Lett. 33, 156–158 (2008).
Pnevmatikakis, E. A. & Giovannucci, A. J. Neurosci. Methods 291, 83–94 (2017).
Vercauteren, T., Pennec, X., Perchant, A. & Ayache, N. Neuroimage 45, S61–S72 (2009).
Schmid, B., Schindelin, J., Cardona, A., Longair, M. & Heisenberg, M. BMC Bioinformatics 11, 274 (2010).
Acknowledgements
We acknowledge N. Ji, C. Xu and J. Kubby for detailed advice on deep imaging; T. Komiyama (UCSD) for the gift of transgenic mice; B. Lim (UCSD) for the gift of viruses; M. Thunemann for instruction on synthesis of Cy5.5–dextran; C. Foo for assistance with the motion-correction algorithm; A. Lozada and C. Mateo for advice on surgery; M. Deschênes, M. Häusser, B. Strowbridge and K. Svoboda for discussions; and B. Friedman for comments on an early version of the manuscript. Design and fabrication of the microscope were supported by the National Science Foundation (MRI grant PHY153264) and physiological measurements were additionally supported by the National Institutes of Health (NINDS grant R35 NS097265).
Author information
Authors and Affiliations
Contributions
D.K. and R.L. designed the instrument, planned the experiments and wrote the manuscript; R.L. fabricated the instrument and performed all experiments; Z.L. coded the reconstruction algorithm and assisted with initial experiments; J.S.M. synthesized the glutamate sensor; and D.K. ensured that all university rules and forms that govern environmental health and safety, including the ethical use of animals as well as the use of chemicals, controlled substances, hazardous substances, lasers and viruses were complied with.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The best signal-to-noise ratio (SNR) of in vivo dendritic spine morphological and calcium imaging in previously published work.
(a, b) Comparison of in vivo dendritic spine morphological imaging SNR among previous works using traditional two-photon microscopy or with adaptive optics at different depth. Subgraph in the bottom left corner of a shows how to estimate SNR. The images for SNR estimation are directly imported from the publications. The arrows in panel a indicate the increase of the SNR by adaptive optics. (c) Comparison of in vivo dendritic spine calcium functional imaging SNR among previous works using traditional two-photon microscopy or with adaptive optics at different depth. Subgraph in the bottom left corner shows how to estimate the SNR of dendritic spine calcium signal.
Supplementary Figure 2 Schematic and calibration of adaptive optical two-photon microscope setup.
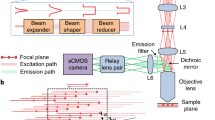
(a) A tunable femtosecond beam is expanded and incident on the deformable mirror (DM). The DM, the X and Y galvo mirrors, and the back focal plane of the objective are all conjugated by the 4-f lens pair between each other. The Shack-Hartmann wavefront sensor (SHWS) is placed in a descan mode. The microlens array of the SHWS is further conjugated with the objective back focal place. During wavefront sensing, the fluorescence emission light is descanned and separated with the guidestar excitation beam by dichroic D4 before arriving at the microlens array of the SHWS. Function signals are collected onto multi-pixel photon counters. (b, c) System AO aberration calibration. Top panels show the changes of Zernike mode coefficients as a function of optimization cycles. Bottom panels show the system aberration represented by Zernike mode index at wavelengths of 1.07 µm and 1.25 µm after 30 cycles optimization. (d) Lateral and axial images and signal profiles of a 200-nm diameter fluorescent bead imaged at 1070-nm wavelength.
Supplementary Figure 3 Design of detection system using multi-pixel photon counting modules.
Part numbers refer to lenses from Thorlabs. (a) Short-arm channel. (b) Long-arm channels.
Supplementary Figure 4 Wavefront sensing at depths up to 850 µm in living mouse cortex.
The signal-to-background ratio (SBR) of the spots image acquired by Shack-Hartmann wavefront sensor decays as the depth increases as a result of tissue scattering. (a) Maximum intensity projections along x axis (Δx = 200 µm) of the cortical vasculature labeled by Cy5.5-dextran. (b) Guide-star microvascular images acquired by two-photon imaging at 14 µm to 850 µm below the pia and their corresponding spot images acquired by the SHWS during direct wavefront sensing. (c) Histograms of the SBR at different depths for the data on panel b with enlarged histograms of the SBR≤1.1 shown as subgraphs from 500-µm to 850-µm depth. (d) Examples of the wavefront image spots with the SBR below the value 1.1. (e) Weight distribution as a function of SBR that is used for wavefront reconstruction.
Supplementary Figure 5 Full AO correction enhance the signal of Cy5.5-dextran labeled vascular and red blood cell flux in line scan at depth.
(a) Imaging of vascular labeled by Cy5.5-dextran at 692 µm depth below the pia with System and Full AO correction. (b) Red blood cell flux shown as line scan signals along the red and green lines in a are compared between System and Full AO. (c, d) Same imaging as panels a and b but at a depth of 806 µm below the pia. (e) Dependence of signal enhancement of red blood cell flux by AO correction as a function of the depth below the pia. Signal enhancement is defined as the ratio of peak-to-valley contrast of the line scan image of Full AO to System AO. 120 measurements over 3 mice were computed and shown in boxplot with maximum, minimum, third quartile, first quartile and median (n ≥ 6).
Supplementary Figure 6 Wavefront patterns, diffraction limited pia image used for k-space analysis in Fig. 2 and signal enhancement analysis.
(a) Images of SHWS spots and corresponding wavefronts used for correction in panels a and d of Fig. 2, respectively. (b,c) Images and corresponding spatial frequencies at 850-µm depth with System AO and Full AO correction used in panel d of Fig. 2. (d) Diffraction limited image of apical dendrites and spines in pia that used in the k-space comparison in panel d of Fig. 2. (e) Dependence of signal enhancement by AO correction as a function of the depth below the pia. Signal enhancement was calculated by the ratio of peak-to-valley contrast of the line profiles across the fine process in the neuronal images in vS1 between Full AO and System AO. About 190 measurements of dendrites and spines at different depth of vS1 cortex in 5 Thy1-YFP or Thy1-jRGECO1a mice were shown in the boxplot with maximum, minimum, third quartile, first quartile and median (n ≥ 5).
Supplementary Figure 7 High-resolution imaging through layer 5b vS1 cortex in a Rbp4-cre mouse labeled by mRuby2.
(a) The three-dimensional image stack from in vivo imaging of mRuby2 labeled layer 5 pyramid neurons in a 150×150×810 µm3 volume of vS1 cortex at an excitation wavelength of λ = 1.07 µm. Full AO corrections were measured and applied every 50 µm of depth. An enlarged view of Fig. 2e. (b) Images at different depths, from 560 to 765 µm below the pia, with System AO and Full AO corrections. The corresponding correcting wavefront is shown.
Supplementary Figure 8 Adaptive optics correction over wide field at a depth of 570 µm below pia.
The subject is a Thy1-YFP mouse. (a, b) Images tiled by 3×3 sub-region scanning with System AO correction and Full AO correction are compared. (c) Zoom-in view of the boxed region of the image in a to compare the images with System and Full correction at different regions over a wide field. (d) Signal profiles of the white lines in a are compared between System and Full AO as a representative of 3 measurements in 2 mice. (e) Scanning path for wide field AO imaging with corresponding subregion wavefronts as shown in f, which were sequentially applied onto the DM and synchronized to sub-region switching by internal triggers. (f) Full AO correcting wavefronts used in panel b.
Supplementary Figure 9 In vivo imaging of glutamate response of thalamocortical axons in vS1 cortex during vibrissa stimulation with full AO.
(a) Coronal section of the mouse brain shows the projection from VPM thalamus to vS1 cortex labeled by SF-Venus-iGluSnFR.A184S. (b) In vivo two-photon imaging of the thalamocortical axons in layer 4 of vS1 cortex is shown as the representative of 4 mice. Red box indicates the imaging region shown in panel d. (c) Air puff whisker stimulation during two-photon imaging. (d) Two-photon imaging of SF-Venus-iGluSnFR.A184S shows the glutamate release from thalamocortical axons and boutons varies as a function of depth as the representative of 7 measurements across 4 mice. Glutamate responses are averaged over the ROI defined by the red box region for all depths. (e, f) Comparison between System AO and Full AO correction on functional imaging of thalamocortical axons and boutons that express SF-Venus-iGluSnFR.A184S at 350 µm below the pia. Corresponding glutamate responses to vibrissa stimulation in the ROIs defined by the dashed circles are shown in panel f. (g, h) Same as panels e and f but at the depth of 650 µm. Experiments in panels e-h were repeated in 2 mice.
Supplementary Figure 10 Full AO correction enables imaging of Cy5.5-dextran labeled vascular and red blood cell flux through the polished and reinforced thinned-skull window at depth in vS1 cortex.
(a) Maximum intensity projections on y axis of the 140 × 140 × 600 µm3 volume images are compared between System (left) and Full (right) AO correction. Wavefronts at four different depths are shown on the right. Experiments were repeated in 2 mice with 2 measurements across depths. (b) Lateral images at 225, 325, 425 and 525 µm depth belong to the volume shown in panel a are compared between System (left) and Full AO (right) correction. (c) Signal profiles along the yellow dashed line in panel b are compared between System (blue) and Full AO (red) correction. (d) Imaging of vascular labeled by Cy5.5-dextran at 383 µm depth through the thinned-skull window with Full AO correction. (e) Red blood cell flux shown as line scan signals along the red and green lines in panel d are compared between System and Full AO. (f, g) Same imaging as panels d and e but at a depth of 541 µm below the pia. (h) Dependence of signal enhancement of red blood cell flux by AO correction as a function of the depth below the pia. Signal enhancement is defined as the ratio of peak-to-valley contrast of the line scan image of Full AO to System AO. 60 measurements over 3 mice were computed and shown in the boxplot with maximum, minimum, third quartile, first quartile and median (n ≥ 6).
Supplementary Figure 11 Wide-field imaging through thinned-skull window with Full AO and signal enhancement analysis.
(a,b) Neuronal images at 320 µm below the pia in S1 cortex of Thy1-YFP mouse with System AO and Full AO correction and corresponding k-space map shown in panel c. (d, e) Vascular images at the same region of panel a with System AO and Full AO corrections. (f) Tiled scanning path is used for wide field AO imaging, same as in Supplementary Fig. 8. (g) Wavefronts used for correction in panels b and e. (h) Signal profiles along the dashed lines in panels a and b respectively, are compared between System AO (blue trace) and Full AO (red trace) correction. (i) Signal enhancement was calculated by the ratio of peak-to-valley contrast of the line signal profiles across the fine process in the neuronal images in vS1 between Full AO and System AO. 128 measurements in 3 Thy1-YFP or Thy1-jRGECO1a mice were shown as the boxplot with maximum, minimum, third quartile, first quartile and median (n ≥ 10).
Supplementary Figure 12 Adaptive optics correction enables neuronal morphological and functional imaging through a thinned-skull window down to 500 µm below the pia.
(a, b) Imaging of apical dendrites and spines labeled by jRGECO1a at the depth of 100 µm and 220 µm below the pia through the thinned skull window with adaptive optics. (c) Functional imaging of barrel cortex at 386 µm below pia in Thy1-jRGECO1a mouse with System and Full AO correction are shown. (d) Spontaneous calcium transients within the ROIs in compared between System AO (blue trace) and Full AO (red trace) corrections. (e, f) Same imaging as panels a and b but at a depth of 520 µm below the pia. Note that for ROI calcium transient calculation, neuropil responses are subtracted by using the surround regions. 3 measurements at similar depth as panels e and f were repeated.
Supplementary Figure 13 Assessing phototoxicity by measuring sensory evoked glutamate response of thalamocortical axons in vS1 L5 during chronic imaging with adaptive optics.
(a) Experiment of 30-minutes continuous imaging with rhythmic air puff whisker stimulation. (b) The experiment as in panel a was repeated at same location for 3 days. Images of the thalamocortical axons labeled by SF-Venus-iGluSnFR.A184S at the same region of B1 barrel at the depth of 660 µm and corresponding wavefronts for each day are shown. (c) Sensory evoked glutamate signals of the axons in the imaging field were averaged every 5 minutes (n = 15 trials) and shown during 30-minutes continuous imaging for each day. (d) Averaged glutamate baseline signal of 3 days based on 3 measurements.
Supplementary Figure 14 Assessing phototoxicity by measuring sensory evoked calcium response of basal dendrites and spines in vS1 L5 during long-term imaging with adaptive optics.
(a, b) Same experiment as in Supplementary Fig. 13a was performed. Images of the dendrites and spines labeled by jRGECO1a in C1 barrel region at 650-µm depth. (c) Sensory evoked calcium signals of the dendrites in ROI7 and spines in ROI1 and ROI3 shown in a and b were averaged every 5 minutes during 30-minutes imaging session (15 trials). (d, e) Normalized amplitudes of sensory evoked calcium signals of 6 spines in ROI1-ROI6 as in panel b and 3 dendrites in ROI7-ROI9 as in panel a. Slopes were calculated by linear regression fit with initial point constraints for each ROI datasets. The standard deviation and the mean of the slope values were calculated and shown in panel d (n = 6) and panel e (n = 3). The black lines indicate the fitting curve using mean slope with constraint initial point.
Supplementary Figure 15 Assessing phototoxicity by measuring red blood cell flux during long-term line scanning on capillary with adaptive optics.
(a) Imaging of the capillary and line scanning along the path shown as red. 30-minutes continuous line scanning was performed at 740-µm depth below the pia. 800-ms red blood cell flux images at the beginning of each 5 minutes were shown during 30 minutes. Imaging power was adjusted minimum that allows to achieve sufficient single to noise for flux measurement. No stalling was observed. (b) Line scanning imaging as a was performed on a capillary at the depth of 732 µm but with maximum power. 400-ms red blood cell flux images at the beginning of each 5 minutes were shown during 30 minutes. No stalling was observed. (c) Line scanning imaging as a was performed on a capillary at the depth of 540 µm with maximum power as in b. 1600-ms red blood cell flux images at the beginning of each 5 minutes were shown during 20 minutes. Stalling was observed after 20-minutes line scanning. Experiments at similar depth below the pia as panels a-c were repeated in 2 mice.
Supplementary information
Supplementary Information
Supplementary Figures 1–15 and Supplementary Tables 1–3
Supplementary Software
Matlab code for the system-aberration calculation, DM and SHWS calibration and sample-induced wavefront reconstruction with full AO correction. Usage instructions, example images and Matlab data are also provided
Supplementary Video 1
Cy5.5–dextran-labeled vasculature. Three-dimensional reconstruction of Cy5.5–dextran-labeled vasculature image down to 1.1-mm depth below the pia acquired by in vivo two-photon imaging.
Supplementary Video 2
Morphological imaging with system AO and full AO. In vivo imaging with system AO and full AO correction from 685 to 720 µm below the pia in a Thy1–YFP mouse.
Supplementary Video 3
Functional imaging with system AO and full AO. In vivo calcium imaging with system AO and full AO correction at 700 µm below the pia in a Rbp4-Cre mouse labeled with jRGECO1a.
Rights and permissions
About this article
Cite this article
Liu, R., Li, Z., Marvin, J.S. et al. Direct wavefront sensing enables functional imaging of infragranular axons and spines. Nat Methods 16, 615–618 (2019). https://doi.org/10.1038/s41592-019-0434-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0434-7
This article is cited by
-
Robust and adjustable dynamic scattering compensation for high-precision deep tissue optogenetics
Communications Biology (2023)
-
Construction and use of an adaptive optics two-photon microscope with direct wavefront sensing
Nature Protocols (2023)
-
Deep learning-driven adaptive optics for single-molecule localization microscopy
Nature Methods (2023)
-
Glutamate indicators with improved activation kinetics and localization for imaging synaptic transmission
Nature Methods (2023)
-
Recent advances in optical imaging through deep tissue: imaging probes and techniques
Biomaterials Research (2022)