Abstract
To capture highly dynamic biological processes at cellular resolution is a recurring challenge in biology. Here we show that combining selective-volume illumination with simultaneous acquisition of orthogonal light fields yields three-dimensional images with high, isotropic spatial resolution and a significant reduction of reconstruction artefacts, thereby overcoming current limitations of light-field microscopy implementations. We demonstrate medaka heart and blood flow imaging at single-cell resolution and free of motion artefacts at volume rates of up to 200 Hz.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Code availability
All custom code used during the current study are either available as Supplementary Software or from the corresponding authors on reasonable request.
References
Winter, P. W. & Shroff, H. Curr. Opin. Chem. Biol. 20, 46–53 (2014).
Katona, G. et al. Nat. Methods 9, 201–208 (2012).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. K. Science 305, 1007–1009 (2004).
Abrahamsson, S. et al. Nat. Methods 10, 60–63 (2013).
Tomer, R. et al. Cell 163, 1796–1806 (2015).
Lu, R. et al. Nat. Neurosci. 20, 620–628 (2017).
Levoy, M., Ng, R., Adams, A., Footer, M. & Horowitz, M. ACM Trans. Graph. 25, 924 (2006).
Broxton, M. et al. Opt. Express 21, 25418 (2013).
Prevedel, R. et al. Nat. Methods 11, 727–730 (2014).
Nöbauer, T. et al. Nat. Methods 14, 811–818 (2017).
Pégard, N. C. et al. Optica 3, 517 (2016).
Cohen, N. et al. Opt. Express 22, 24817–24839 (2014).
Truong, T. V. et al. Preprint at bioRxiv https://doi.org/10.1101/403303 (2018).
Wu, Y. et al. Nat. Biotechnol. 31, 1032–1038 (2013).
Preibisch, S. et al. Nat. Methods 11, 645–648 (2014).
Fahrbach, F. O., Voigt, F. F., Schmid, B., Helmchen, F. & Huisken, J. Opt. Express 21, 21010 (2013).
Mickoleit, M. et al. Nat. Methods 11, 919 (2014).
Taylor, J. M. Front. Physiol. 5, 481 (2014).
Gong, Y. et al. Science 350, 1361–1366 (2015).
de Medeiros, G. et al. Nat. Commun. 6, 8881 (2015).
Botcherby, E. J., Juškaitis, R., Booth, M. J. & Wilson, T. Opt. Commun. 281, 880–887 (2008).
Manton, J. D. & Rees, E. J. Opt. Lett. 41, 4170–4173 (2016).
Koster, R., Stick, R., Loosli, F. & Wittbrodt, J. Development 124, 3147–3156 (1997).
Consortium, T. E. P. et al. Nature 489, 57 (2012).
Kirchmaier, S. et al. Development 140, 4287–4295 (2013).
Kirchmaier, S., Lust, K. & Wittbrodt, J. PLoS ONE 8, e76117 (2013).
Stemmer, M., Thumberger, T., del Sol Keyer, M., Wittbrodt, J. & Mateo, J. L. PLoS ONE 10, e0124633 (2015).
Meijering, E., Dzyubachyk, O. & Smal, I. Methods Enzymol. 504, 183–200 (2012).
Acknowledgements
We would like to thank the European Molecular Biology Laboratory (EMBL) Heidelberg mechanical and electronic workshop, the EMBL IT department as well as Luxendo GmbH (www.luxendo.eu) for technical support, as well as C. Tischer (EMBL ALMF) for help. We further thank T. Thumberger for advice on CRISPR−Cas donor design and M. Majewski, E. Leist and A. Saraceno for fish husbandry. J.G. is a fellow of the MD/PhD program of the Medical Faculty Heidelberg and of the Heidelberg Biosciences International Graduate School (HBIGS), and is grateful to M. Gorenflo for supervision and guidance. J.G. was supported by a Heidelberg Research Center for Molecular Medicine (HRCMM) Career Development Fellowship (CDF) and by the Deutsche Herzstiftung e.V. (S/02/17). N.N. acknowledges support from VINNOVA and the Åke Wiberg foundation. This work was supported by the EMBL (N.W., N.N., B.B., G.M., L.H and R.P.).
Author information
Authors and Affiliations
Contributions
N.N., L.H. and R.P. designed the imaging system. N.W. and N.N. built the imaging system and performed experiments together with J.G. and G.M. R.P. designed microlenses, J.G. generated transgenic animals under guidance of J.W. N.W. wrote analysis software with help from N.N. B.B. contributed to hardware control and data processing, and N.N. wrote computational cluster pipeline with assistance from N.W. R.P. wrote the manuscript with input from all authors. R.P. and L.H. conceived and led the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare competing financial interests. L.H. is scientific co-founder and advisor of Luxendo GmbH (part of Bruker), which makes light sheet-based microscopes commercially available.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
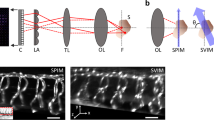
Supplementary Fig. 1 Iso-LFM setup.
(a) Schematic 2D drawing of the setup showing the main opto-mechanical components. Note that the two imaging objectives do not face each other co-axially but are arranged in an orthogonal fashion as further indicated in (c). Depending on the excitation beam size (adjustable beam expander) and scan mode of the galvanometric mirrors either a conventional light-sheet (yellow plane), or a selective volume can be illuminated (b). For very fast imaging modes (50 Hz or more), we chose to overfill a quadratic 2D slit which was relayed into the focal volume of the detection objectives, in order to instantaneously excite the entire volume of interest. The fluorescence is detected by two orthogonally oriented detection paths. A narrow bandpass-filter (BP filter - Semrock, 525/50 nm BrightLine) is placed in front of the tube lens, which focuses either on the camera sensor (light sheet mode) or on the microlens array (Iso-LFM mode). In the latter case, the camera sensor is displaced by one focal length of the microlenses further away from the tube lens. A 1-1 relay lens system is used to relay the image plane of the camera sensor (see Methods for details). (c) Experimental implementation of the microscope. Optical paths are colored as in (a). Zoom-in shows a close-up of the three mutually orthogonal objectives and the sample area.
Supplementary Fig. 2 Sample stage and mounting.
(a) Close-up of the composite stage (left panel). Fine positioning of the sample is achieved with a combination of coarse manual XYZ positioning, fast piezo stack positioning for light-sheet acquisition or changing the volumetric field-of-view in Iso-LFM, as well as a miniature rotation stage. (b) Medaka fish hatchlings are first mounted in an agarose filled capillary (inner diameter 1.6 mm) and thereafter the agarose tube is attached to a custom sample mount which is positioned into the focal volume of excitation and both detection objectives. (c) Close-up of the sample stage. Note that only one detection objective is shown here for clarity (i.e. View 2 has been removed). Illumination (detection) objective is labelled by a cyan (green) arrow.
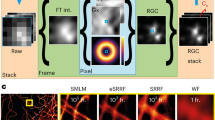
Supplementary Fig. 3 Light-field image rectification.
(a) Every acquired light-field image has to be rectified in order to allow for proper light-field reconstruction. This involves resizing and interpolating the images to ensure that the number of pixels behind each microlens exactly correspond to the number of pixels defined in the PSF computation. (b) An automated image rectification procedure is implemented by detecting the lenslet center position and periodicity based on a calibration or background image. This was verified in 10 independent images.
Supplementary Fig. 4 Iso-LFM reconstruction pipeline.
The workflow for reconstructing the acquired 2D light-field images to multiview deconvolved 3D datasets. The pipeline is based on MATLAB (orange) as well as FIJI (blue). The upper left numbers correspond to the protocol sections in Supplementary Note 1. An automated rectification script as well as a custom job preparation script for automated distribution of multi-view deconvolution jobs facilitate this workflow and are highlighted by orange boxes (see Supplementary Software).
Supplementary Fig. 5 Overlap of light-sheet and Iso-LFM volumes.
Lateral (top) as well as axial (bottom) maximum intensity projections of fluorescent beads distributed in 3D in agarose, imaged by either light sheet (cyan) or Iso-LFM (red) and a 40x 0.8NA objective. The mean distance in 3D space between light-sheet and Iso-LFM reconstructions of the same beads is ~1.5 ± 0.7 µm (mean ± SD, n = 24 beads) and therefore smaller than the resolution of Iso-LFM. Scale bar is 10 µm.
Supplementary Fig. 6 Characterization of performance for 22.5x 0.5NA objectives.
(a,b) Maximum intensity projections of reconstructed fluorescent beads and their respective resolution (FWHM) for single-view and dual-view, respectively. The lower panels depict the measured PSF at −80 µm off the focal plane (averaged over n = 12 beads) with lateral and axial FWHM indicated. Scalebars upper panel 10 µm and lower left panel 5 µm. (c) Mean values of lateral and axial PSF FWHM for the single-view (n = 2192 beads total) and the dual-view (n = 2816 beads total) case across imaging volume (steps of 30 µm). Multiview deconvolution improves axial resolution across the entire overlapping volume and achieves near-isotropic resolution of better than ~5 µm (5.2 ± 1.3 µm, ~2.7 ± 0.5 µm and ~4.8 ± 1.2 µm in x/y and z, respectively). Centre lines represent mean and shadings denote SD.
Supplementary Fig. 7 Blood flow quantifications from single-cell tracking using Iso-LFM.
Single blood cells were manually tracked in 3D using the Fiji plugin MtrackJ1. Mean velocity (solid line) of all tracked blood cells (n = 28) over time. Shaded region denotes SD. Green and red dots denote velocities of tracked cells in the center and close to the vessel boundary, respectively, as indicated in the inset (scale bar 20 µm). On average, cells in the center of the vessel show higher velocities compared to those close to the vessel’s wall.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Notes 1–4
Supplementary Software
Scripts for automated rectification and distribution of dual-view deconvolution jobs.
Supplementary Video 1
Imaging of the beating heart at 143Hz volume rate in a Medaka fish hatchling. Heart imaging of cardiomyocyte nuclei (myl7::H2B-eGFP) labeled Medaka hatchlings. Maximum intensity projections of single-view (top panels) and multi-view. deconvolved dual-view Iso-LFM (bottom panels). Dashed white box indicates zoom-in region displayed on right most panels. Representative video of 10 recording data sets in n=3 independent fish. Scale bars 20 µm.
Supplementary Video 2
Imaging of blood flow dynamics at single cell resolution and with 200Hz volume rate in a Medaka fish hatchling. Maximum intensity projections of single-view and multiview deconvolved dual-view Iso-LFM of mature blood cells (ENH-HMMA-4s-hsp70::eGFP) in Medaka hatchlings. For dual-view (bottom panels), single blood cells are resolved in all three orientations. Representative video of 10 recording data sets in n=2 independent fish. Scale bars 20 µm.
Supplementary Video 3
Imaging of blood flow through the heart at single cell resolution and with 166Hz volume rate in a Medaka fish hatchling. Maximum intensity projections of single-view and multiview deconvolved dual-view Iso-LFM of sparsely labelled blood cells (ENH-HMMA-4s-hsp70::eGFP) in Medaka hatchlings. For better visualization, A, V and the dashed white line denote the atrium, ventricle and rough outline of the heart, respectively. Representative video of 10 recording data sets in n=2 independent fish. Scale bars 20 µm.
Rights and permissions
About this article
Cite this article
Wagner, N., Norlin, N., Gierten, J. et al. Instantaneous isotropic volumetric imaging of fast biological processes. Nat Methods 16, 497–500 (2019). https://doi.org/10.1038/s41592-019-0393-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0393-z
This article is cited by
-
Light-field flow cytometry for high-resolution, volumetric and multiparametric 3D single-cell analysis
Nature Communications (2024)
-
Multi-focus light-field microscopy for high-speed large-volume imaging
PhotoniX (2022)
-
A practical guide to scanning light-field microscopy with digital adaptive optics
Nature Protocols (2022)
-
3D Reconstruction of cellular images from microfabricated imagers using fully-adaptive deep neural networks
Scientific Reports (2022)
-
Computational Analysis of Cardiac Contractile Function
Current Cardiology Reports (2022)