Abstract
Image scanning microscopy (ISM) can improve the effective spatial resolution of confocal microscopy to its theoretical limit. However, current implementations are not robust or versatile, and are incompatible with fluorescence lifetime imaging (FLIM). We describe an implementation of ISM based on a single-photon detector array that enables super-resolution FLIM and improves multicolor, live-cell and in-depth imaging, thereby paving the way for a massive transition from confocal microscopy to ISM.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
The Matlab software for PR, multi-image deconvolution, phasor plot calculation and FRC analysis is freely available for academic use and is provided online with this paper as Supplementary Software.
References
Pawley, J.B. (ed.) Handbook of Biological Confocal Microscopy (Springer, 1995).
Ebrecht, R., Don Paul, C. & Wouters, F. S. Protoplasma 251, 293–305 (2014).
Sheppard, C. J. R. Optik (Stuttg.) 80, 53–54 (1988).
Sheppard, C. J. R., Mehta, S. B. & Heintzmann, R. Opt. Lett. 38, 2889–2892 (2013).
Müller, C. B. & Enderlein, J. Phys. Rev. Lett. 104, 198101 (2010).
Castello, M., Sheppard, C. J. R., Diaspro, A. & Vicidomini, G. Opt. Lett. 40, 5355–5358 (2015).
York, A. G. et al. Nat. Methods 9, 749–754 (2012).
Schulz, O. et al. Proc. Natl Acad. Sci. USA 110, 21000–21005 (2013).
De Luca, G. M. et al. Biomed. Opt. Express 4, 2644–2656 (2013).
York, A. G. et al. Nat. Methods 10, 1122–1126 (2013).
Roth, S., Sheppard, C. J. R., Wicker, K. & Heintzmann, R. Opt. Nanoscopy 2, 5 (2013).
Winter, P. W. et al. Optica 1, 181–191 (2014).
Gregor, I. et al. Nat. Methods 14, 1087–1089 (2017).
Sheppard, C. J. R., Castello, M., Tortarolo, G., Vicidomini, G. & Diaspro, A. J. Opt. Soc. Am. A Opt. Image Sci. Vis. 34, 1339–1350 (2017).
Huff, J. Nat. Methods 12, i–ii (2015).
Zappa, F., Tisa, S., Tosi, A. & Cova, S. Sens. Actuators A Phys. 140, 103–112 (2007).
Becker, W. (ed.) Advanced Time-Correlated Single Photon Counting Applications (Springer, 2015).
Bertero, M., Boccacci, P., Desider’a, G. & Vicidomini, G. Inverse Probl. 25, 123006 (2009).
Tortarolo, G., Castello, M., Diaspro, A., Koho, S. & Vicidomini, G. Optica 5, 32–35 (2018).
Owen, D. M. et al. Biophys. J. 90, L80–L82 (2006).
Booth, M. J. Light Sci. Appl. 3, e165 (2014).
Digman, M. A. & Gratton, E. Annu. Rev. Phys. Chem. 62, 645–668 (2011).
Israel, Y., Tenne, R., Oron, D. & Silberberg, Y. Nat. Commun. 8, 14786 (2017).
Hoebe, R. A. et al. Nat. Biotechnol. 25, 249–253 (2007).
Yu, Z., Liu, S., Zhu, D., Kuang, C. & Liu, X. Opt. Commun. 404, 139–146 (2017).
Vicidomini, G., Bianchini, P. & Diaspro, A. Nat. Methods 15, 173–182 (2018).
Vicidomini, G., Schmidt, R., Egner, A., Hell, S. & Schönle, A. Opt. Express 18, 10154–10167 (2010).
Ingaramo, M. et al. ChemPhysChem 15, 794–800 (2014).
Castello, M., Diaspro, A. & Vicidomini, G. Appl. Phys. Lett. 105, 234106 (2014).
Koho, S., Deguchi, T. & Hänninen, P. E. J. Microsc. 260, 208–218 (2015).
Ströhl, F. & Kaminski, C. F. Methods Appl. Fluoresc. 3, 014002 (2015).
Digman, M. A., Caiolfa, V. R., Zamai, M. & Gratton, E. Biophys. J. 94, L14–L16 (2008).
Chung, K. et al. Nature 497, 332–337 (2013).
Acknowledgements
This work was partially supported by the Compagnia di San Paolo (ROL 20641 to G.T. and G.V.). We thank R. Nakamura (Nikon Corporation, Japan) for support with confocal A1R measurements, and Nikon Corporation for sharing useful technical information about confocal A1R and for dual-color (mitochondria and tubulin) sample preparation. We thank R. Kawakami, K. Otomo and T. Nemoto (Research Institute for Electronic Science, Hokkaido University) for advice on whole-brain imaging. We thank F. Cella Zanacchi (Istituto Italiano di Tecnologia) for support in cell preparation, and S. Piazza (Istituto Italiano di Tecnologia), E. Slenders (Hasselt University) and E. Tcarenkova (Turku University) for useful discussions.
Author information
Authors and Affiliations
Contributions
M.C. and G.V. conceived the idea. A.T. and G.V. supervised the project, with support from C.J.R.S., P.B. and A.D. M.C., M.B., F.V., A.T. and G.V. designed the SPAD array. M.B., F.V. and A.T. realized and characterized the SPAD array. M.C., G.T. and G.V. designed and implemented the custom ISM system. T.D., P.B. and G.V. integrated the ISM method on the commercial microscope. M.C., G.T. and G.V. designed and developed the controlling architecture. M.C., G.T., S.K. and G.V. developed the image processing and analysis software. M.C., G.T., T.D., M.O., S.P., P.B. and G.V. performed the experiments and analyzed the data with support from all other authors. L.P., M.O., S.P. and L.L. assisted in live-cell imaging and fluorescence lifetime experiments. M.C., G.T. and G.V. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
M.C., G.T., M.B., F.V., P.B., C.J.R.S., A.D., A.T. and G.V. have filed a patent application (application number IT 102018000001891) on the method presented.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The SPAD array detection system developed for image scanning microscopy.
Photograph of the SPAD array detection system. The photograph shows the two stacked printed circuit boards (PCBs). Inset shows the 5 × 5 SPAD array. (b) Photon detection efficiency (PDE) of the central element of the 5 × 5 SPAD array, at different excess bias voltages. (c) Temporal response (or impulse-response function (IRF)) of the central element (Vex = 6 V excess-bias) of the SPAD array to a pulsed laser source at 850 nm (20 ps FWHM) when all the other pixels are turned ON. The long tail on the right side of the IRF is due to the optical cross-talk with the other SPAD elements.
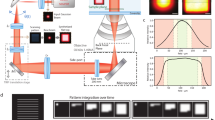
Supplementary Figure 2 Custom image scanning microscopy schematic setup.
LPD, laser-pulsed diode; HWP, half-wave plate; QWP, quarter-wave plate; AOM, acoustic optical modulator; PMF, polarized maintaining fiber; PC, personal computer; TDCs, time-to-digital converters; TTL, transistor–transistor logic; 3A-S, three-axis stage; 3A-PS, three-axis piezo stage; DM, dichroic mirror; GMs, galvanometer mirrors; SL, scan lens; OL, objective lens.
Supplementary Figure 3 Image scanning microscopy with adaptive (A) pixel reassignment (PR).
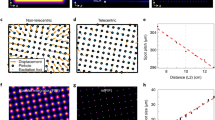
(a) Side-by-side comparison of the effective PSFs for ‘ideal’ confocal (0.2 AU), ‘open’ confocal (1.4 AU), uncorrected ISM (the pixel- reassignment method uses theoretical shift vectors), and ISM (the pixel-reassignment method uses estimated shift vectors). Scale bar, 500 nm. (b) Fingerprint maps superimposed with the shift vectors for the uncorrected ISM (left) and the true ISM (right). Scale bars, 50 nm. (c) Radial PSFs obtained from the intensity profiles of (a). The top panel shows the un-normalized PSFs, and the middle and bottom panels show the normalized PSFs, together with the Gaussian fitted data. Pixel-dwell time, 50 μs. Pixel-size, 5 nm. Image format, 400 × 400 pixels. Scale bars, 500 nm.
Supplementary Figure 4 Image scanning microscopy on nuclear pore complexes (NPCs).
(a) Comparison of large-field-of-view images of the nuclear pore scaffold structures stained with Alexa Fluor 488: ‘ideal’ confocal (top) and APR-ISM (bottom). Image format, 1,500 × 1,500 pixels; scale bars, 10 μm. (b) Side-by-side comparison of single-cell images, ideal confocal (0.25 AU), open confocal (1.7 AU), APR-ISM and deconvolved ISM++ (five iterations), respectively. Notably, to highlight the property of the proposed ISM implementation to generate high-resolution large-field-of-view images, all images are a digital zoom of the white boxes in (a). Image format, 400 × 400 pixels; scale bars, 1 μm. (c) Magnified views of the white boxes in (b). Image format, 100 × 100 pixels; scale bars, 1 μm. (d) Line intensity profiles across two closely packed NPCs at the position of the arrowheads for the different imaging modalities. For all images: pixel dwell time, 100 μs; pixel size, 66.6 nm; excitation power Pexc, 840 nW. The FRC-based resolution values for the 1,500 × 1,500 pixel images are 213 nm, 253 nm and 207 nm for ideal confocal, open confocal and APR-ISM, respectively. Data are representative of n = 10 experiments.
Supplementary Figure 5 Image scanning microscopy of nuclear lamin.
(a) Side-by-side comparison of HEK cell nuclear lamin stained with Alexa Fluor 488: ‘ideal’ confocal (0.25 AU), open confocal (1.7 AU), APR-ISM and deconvolved ISM++ (five iterations). Image format, 600 × 600 pixels. (b) Magnified views of the regions outlined by white boxes in (a). (c) Line intensity profiles across a nuclear invagination (nucleoplasmic reticulum) at the position of the arrowheads in (b) for the different imaging modalities. For all images: pixel dwell time, 100 μs; pixel size, 50 nm; excitation power Pexc, 840 nW. Scale bars, 1 μm. The FRC-based resolution values for the images are 238 nm, 263 nm, and 206 nm for ideal confocal, open confocal and APR-ISM, respectively. Data are representative of n = 10 experiments.
Supplementary Figure 6 Image scanning microscopy of fluorescent beads.
(a) Side-by-side comparison of ‘ideal’ confocal, ‘open’ confocal, APR-ISM and deconvolved ISM++ (ten iterations) images (500 × 500 pixels, 40-nm pixel size) of 20-nm red fluorescent beads. Pixel dwell time, 50 μs. Pixel size, 40 nm. Image format, 500 × 500 pixels. Excitation power Pexc, 56 nW. Scale bars, 1 μm. (b) Magnified views of the regions outlined by white boxes in (a). Scale bars, 1 μm. (c) Line intensity profiles across two close fluorescent beads at the position of the arrowheads in (b) for the different imaging modalities. Data are representative of n = 5 experiments.
Supplementary Figure 7 FRC-based resolution scaling on ISM with increasing excitation power.
(a) Series of tubulin (labeled with Abberior STAR red) images of the same area for ‘ideal’ confocal (top), ‘open’ confocal (middle) and APR-ISM (bottom) as a function of the excitation beam power. Bleaching was negligible across the whole imaging experiment. Pixel dwell time, 100 μs. Pixel size, 37.5 nm. Image format, 400 × 400 pixels. Excitation power Pexc: 50, 55, 62, 70, 90, 110, 170, 220, 250, 350, 520, 700, 890, 1,000, 1,300 and 2,000 nW. Scale bars, 1 μm. (b) Fourier-ring-correlation curves for different excitation beam powers and different imaging modalities: ideal confocal (top), open confocal (middle) and APR-ISM (bottom). The fixed 1/7 threshold value is also reported with the curves. Resolution (based on the FRC analysis and the 1/7 threshold value) as a function of excitation power for the three different imaging modalities is reported in Fig. 1f. Data are representative of n = 2 experiments.
Supplementary Figure 8 Image scanning microscopy for live-cell mitochondria imaging.
(a) Side-by-side comparison of ‘ideal’ confocal (top), ‘open’ confocal (middle-top), APR-ISM (middle-bottom) and deconvolved ISM++ time-lapse (3 min, 24 frames) of mitochondria labeled with MitoTracker Deep Red in a live cell. For each imaging modality, three representative frames are shown (0 s, 90 s and 180 s). Pixel dwell time, 30 μs. Pixel size, 40 nm. Image format, 500 × 500 pixels. Excitation power Pexc, 140 nW. Scale bars, 1 μm. (b) Maximum intensity projections (color-coded by time) of the time-lapses for the different imaging modalities. These projections allow one to identify the fraction of mitochondria with minimal mobility (white) from the mobile one (colored). Data are representative of n = 10 experiments.
Supplementary Figure 9 Image scanning microscopy for live-cell tubulin imaging.
(a) Side-by-side comparison of ‘ideal’ confocal (top), ‘open’ confocal (middle), and APR-ISM (bottom) time-lapses (4.5 min, 19 frames) of tubulin labeled with SiR-tubulin in a live HeLa cell. For each imaging modality, three representative frames are shown (0 s, 142 s and 264 s). All frames are normalized in intensity to the maximum and minimum of the first frame of the respective time-lapse. This allows one to appreciate the negligible photobleaching. Pixel dwell time, 50 μs. Pixel size, 57.2 nm. Image format, 700 × 700 pixels. Excitation power Pexc, 2 μW. Scale bars, 1 μm. (b) Magnified view of the regions outlined by white dashed boxes in (a). Data are representative of n = 3 experiments.
Supplementary Figure 10 Photon-arrival-time histograms for ISM.
Comparison between the histograms of the photon-arrival times for the different imaging modalities: ISM, ‘open’ CLSM (1.4 AU) and ‘ideal’ CLSM (0.2 AU). We obtained the histograms by integrating all the pixels from the 3D datasets of the tubulin experiment (Supplementary Fig. 11). Clearly, because the pixel reassignment simply shifts photons from one pixel to another, the ISM and open CLSM histograms are identical. This is also an indication that our extension of the pixel-reassignment method to fluorescence lifetime imaging does not introduce artifacts. The reduced SNR of the ‘close’ CLSM appears clear in the comparison between the histograms. We report also the impulse response function (IRF) of the scanning microscope obtained by measuring the reflection (488 nm) from a gold bead.
Supplementary Figure 11 Fluorescence lifetime image scanning microscopy on tubulin labeled with Abberior STAR red.
(a) Side-by-side comparison of the fitting-based ‘ideal’ confocal FLIM (left; 1.4 AU), FLISM (center) and ‘open’ confocal FLIM (right) images of tubulin labeled with Abberior STAR red. Purple pixels correspond to out-of-range fluorescence lifetime values, which are probably the result of artifacts during the time-domain (fitting) analysis. All pixels were analyzed. (b) The respective intensity images are also reported. Pixel dwell time, 100 μs. Pixel size, 30 nm. Image format, 500 × 500 pixels. Excitation power Pexc, 500 nW. Scale bars, 1 μm. Data are representative of n = 4 experiments.
Supplementary Figure 12 Fluorescence lifetime image scanning microscopy of membrane labeled with ANEP.
Side-by-side comparison of the fitting-based FLIM (left; 0.25 AU) and FLISM (right) images of membrane labeled with ANEP. Purple pixels correspond to out-of-range fluorescence lifetime values, which probably resulted from artifacts during the time-domain (fitting) analysis. All pixels were analyzed. Pixel dwell time, 300 μs. Pixel size, 62.6 nm. Image format, 800 × 800 pixels. Excitation power Pexc, 3 μW. Scale bars, 1 μm. Data are representative of n = 4 experiments.
Supplementary Figure 13 Commercial-system-based image scanning microscopy setup.
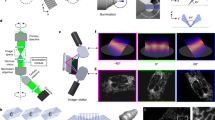
(a) Schematic describing the connections between the different hardware components of the image scanning microscope based on a commercial Nikon A1R system. Carma microscope control registers the photons collected by the SPAD array and is in charge of the detector initialization. The synchronization with the scanning system and with the other actuators of the Nikon A1R, which allows generation of the scanned images, is obtained through communication with the Nikon microscope controller. In the case of imaging with galvanometer mirrors, the carma controller provides the analog scanning signal to the Nikon controller, which successively communicates with the galvanometer mirrors located in the confocal scan head (i.e., carma is the master). In the case of imaging with a resonant mirror (for the fast axis), the carma microscope unit receives the synchronization signals (pixel, line and frame clocks) from the Nikon microscope (i.e., carma is the slave). Both the Nikon and the carma microscope controls communicate with the personal computer (PC). In particular, the PC hosts the carma software, which visualizes, analyzes and processes (deconvolution, pixel reassignment and Fourier-ring correlation) the data. (b) Simplified scheme of the Nikon scan head. Only the important elements for the ISM implementation are reported. The excitation beam (blue) is sent to the galvanometer mirrors (GMs) or resonant mirror thanks to a first dichroic mirror (DM1). The beam is scanned on the specimen/object plane thanks to the scanning lens (SL), the tube lens and the objective lens; tube and objective lenses are not shown in the scheme. The fluorescence (green) is collected by the objective lens and descanned by the GMs. The SL and the TL generate a second conjugate image plane, the pinhole plane, with magnification M2 = 3.9 × M1. M1 is the magnification on the first image plane (not shown in the scheme), which corresponds to the nominal magnification of the objective lens (10 × , 20 × or 60 × in our experiments). A second dichoric mirror (DM2), which usually deflects the fluorescence to the conventional single-point detector (light green), is removed, and the pinhole is completely opened during ISM. The zoom lens (i) is positioned on a five-axis stage (5A-S) to align the fluorescence beam with respect to the SPAD array; (ii) conjugates the pinhole plane on the SPAD array and adds an extra magnification (M3) on the detector plane; and (iii) allows a projected size of the SPAD array detector on the object plane equal to 1 Airy unit.
Supplementary Figure 15 Image scanning microscopy combined with fast resonant scanning.
Side-by-side comparison between ‘ideal’ confocal, ‘open’ confocal and APR-ISM images of tubulin stained with Alexa Fluor 546 (format, 256 × 256 pixels; resonant frequency, 7.9 kHz; zoom factor, 8 × , which results in a pixel size of 103 nm and a minimum pixel dwell time of about 70 ns; 64 line integrations). Insets show magnified views of the regions outlined by white boxes. Scale bars, 1 μm. Data are representative of n = 10 experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 Supplementary Notes 1–4 and Supplementary Table 1
Supplementary Video 1
‘Ideal’ confocal time-lapse imaging of mitochondria. Ideal CLSM (0.2 AU) time-lapse images (3 min, 24 frames) of mitochondria labeled with MitoTracker Deep Red in a live HEK cell. The video is representative of n = 10 experiments
Supplementary Video 2
‘Open’ confocal time-lapse imaging of mitochondria. Open CLSM (1.4 AU) time-lapse images (3 min, 24 frames) of mitochondria labeled with MitoTracker Deep Red in a live HEK cell. The video is representative of n = 10 experiments
Supplementary Video 3
APR-ISM time-lapse imaging of mitochondria. APR-ISM time-lapse images (3 min, 24 frames) of mitochondria labeled with MitoTracker Deep Red in a live HEK cell. The video is representative of n = 10 experiments
Supplementary Video 4
Deconvolved ISM++ time-lapse imaging of mitochondria. Deconvolved ISM++ time-lapse images (3 min, 24 frames) of mitochondria labeled with MitoTracker Deep Red in a live HEK cell. The video is representative of n = 10 experiments
Supplementary Video 5
‘Ideal’ confocal large-field-of-view time-lapse imaging of tubulin. Ideal CLSM (0.2 AU) large-field-of-view time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. Scale bars, 1 μm. The video is representative of n = 3 experiments
Supplementary Video 6
‘Open’ confocal large-field-of-view time-lapse imaging of tubulin. Open CLSM (1.4 AU) large-field-of-view time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. Scale bars, 1 μm. The video is representative of n = 3 experiments
Supplementary Video 7
APR-ISM large-field-of-view time-lapse imaging of tubulin. APR-ISM large-field-of-view time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. Scale bars, 1 μm. The video is representative of n = 3 experiments
Supplementary Video 8
‘Ideal’ confocal time-lapse imaging of tubulin. Ideal CLSM (0.2 AU) time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. The power of the 635-nm excitation laser was 0.5 μW for the first 10 frames and was increased to 5 μW thereafter. The intensities of all frames are normalized to the maximum and minimum values of the first frame. The video is representative of n = 4 experiments
Supplementary Video 9
‘Open’ confocal time-lapse imaging of tubulin. Open CLSM (1.4 AU) time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. The power of the 635-nm excitation laser was 0.5 μW for the first ten frames and was increased to 5 μW thereafter. The intensities of all frames are normalized to the maximum and minimum values of the first frame. The video is representative of n = 4 experiments
Supplementary Video 10
APR-ISM time-lapse imaging of tubulin. APR-ISM time-lapse images of tubulin labeled with SiR-tubulin in a live HeLa cell. The power of the 635-nm excitation laser was 0.5 μW for the first ten frames and was increased to 5 μW thereafter. The intensities of all frames are normalized to the maximum and minimum values of the first frame
Supplementary Video 11
Comparison of APR-ISM and ‘ideal’ confocal time-lapses of tubulin: Comparison of APR-ISM and ideal CLSM time-lapses of tubulin in a live HeLa cell with excitation (635 nm) powers able to provide similar resolution between the two imaging modalities (0.5 μW and 5 μW, respectively). The video shows the time-lapse images we obtained by concatenating the first ten frames of the APR-ISM video (Supplementary Video 7) and the successive frame of the ideal CLSM video (Supplementary Video 5). The intensities of all frames are normalized to the maximum and minimum values of the first frame. This video clearly shows that photobleaching is negligible during APR-ISM imaging, but that it substantially degrades when the excitation power is increased to obtain CLSM imaging at a resolution comparable to that of ISM. The video is representative of n = 4 experiments
Supplementary Software
FRC and APR-ISM analysis Matlab software. Scripts for Fourier-ring correlation analysis, as well as stand-alone executable files (from Matlab) for pixel-reassignment, multi-image deconvolution, and phasor-plot analysis. A user introduction and test images are also provided.
Rights and permissions
About this article
Cite this article
Castello, M., Tortarolo, G., Buttafava, M. et al. A robust and versatile platform for image scanning microscopy enabling super-resolution FLIM. Nat Methods 16, 175–178 (2019). https://doi.org/10.1038/s41592-018-0291-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0291-9
This article is cited by
-
Enhancing image resolution of confocal fluorescence microscopy with deep learning
PhotoniX (2023)
-
Open-source tools enable accessible and advanced image scanning microscopy data analysis
Nature Photonics (2023)
-
SPLIT-PIN software enabling confocal and super-resolution imaging with a virtually closed pinhole
Scientific Reports (2023)
-
High-resolution single-photon imaging with physics-informed deep learning
Nature Communications (2023)
-
Single-photon microscopy to study biomolecular condensates
Nature Communications (2023)