Abstract
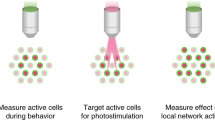
Understanding the causal relationship between neural activity and behavior requires the ability to perform rapid and targeted interventions in ongoing activity. Here we describe a closed-loop all-optical strategy for dynamically controlling neuronal activity patterns in awake mice. We rapidly tailored and delivered two-photon optogenetic stimulation based on online readout of activity using simultaneous two-photon imaging, thus enabling the manipulation of neural circuit activity ‘on the fly’ during behavior.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The original code, together with detailed instructions, is available at the following Github link: https://github.com/alloptical/ClosedLoop. The authors will keep the repository operating and freely accessible. The data sets generated and/or analyzed in this study are available from the corresponding authors upon reasonable request.
References
Emiliani, V., Cohen, A. E., Deisseroth, K. & Häusser, M. J. Neurosci. 35, 13917–13926 (2015).
Rickgauer, J. P., Deisseroth, K. & Tank, D. W. Nat. Neurosci. 17, 1816–1824 (2014).
Szabo, V., Ventalon, C., De Sars, V., Bradley, J. & Emiliani, V. Neuron 84, 1157–1169 (2014).
Packer, A. M., Russell, L. E., Dalgleish, H. W. & Häusser, M. Nat. Methods 12, 140–146 (2015).
Forli, A. et al. Cell Rep. 22, 3087–3098 (2018).
Mardinly, A. R. et al. Nat. Neurosci. 21, 881–893 (2018).
Yang, W., Carrillo-Reid, L., Bando, Y., Peterka, D. S. & Yuste, R. eLife 7, e32671 (2018).
Shadlen, M. N. & Newsome, W. T. Curr. Opin. Neurobiol. 4, 569–579 (1994).
Rieke, F. Spikes: Exploring the Neural Code. (MIT Press, Cambridge, MA, 1999).
Pouget, A., Dayan, P. & Zemel, R. Nat. Rev. Neurosci. 1, 125–132 (2000).
Averbeck, B. B., Latham, P. E. & Pouget, A. Nat. Rev. Neurosci. 7, 358–366 (2006).
Grosenick, L., Marshel, J. H. & Deisseroth, K. Neuron 86, 106–139 (2015).
deCharms, R. C. & Zador, A. Annu. Rev. Neurosci. 23, 613–647 (2000).
Helmchen, F., Imoto, K. & Sakmann, B. Biophys. J. 70, 1069–1081 (1996).
Simons, D. J. J. Neurophysiol. 41, 798–820 (1978).
Bruno, R. M. Curr. Opin. Neurobiol. 21, 701–708 (2011).
Hebb, D. O. The Organization of Behavior. (Wiley, New York, 1949).
Martin, S. J., Grimwood, P. D. & Morris, R. G. Annu. Rev. Neurosci. 23, 649–711 (2000).
Carrillo-Reid, L., Yang, W., Bando, Y., Peterka, D. S. & Yuste, R. Science 353, 691–694 (2016).
Zhang, W. & Linden, D. J. Nat. Rev. Neurosci. 4, 885–900 (2003).
Paz, J. T. et al. Nat. Neurosci. 16, 64–70 (2013).
Krook-Magnuson, E., Armstrong, C., Oijala, M. & Soltesz, I. Nat. Commun. 4, 1376 (2013).
Newman, J. P. et al. eLife 4, e07192 (2015).
Prsa, M., Galiñanes, G. L. & Huber, D. Neuron 93, 929–939 (2017).
Iaccarino, H. F. et al. Nature 540, 230–235 (2016).
Chen, T. W. et al. Nature 499, 295–300 (2013).
Nikolenko, V., Poskanzer, K. E. & Yuste, R. Nat. Methods 4, 943–950 (2007).
Nikolenko, V. et al. Front. Neural Circuits 2, 5 (2008).
Guo, Z. V., Hart, A. C. & Ramanathan, S. Nat. Methods 6, 891–896 (2009).
Dal Maschio, M. et al. Opt. Express 18, 18720–18731 (2010).
Anselmi, F., Ventalon, C., Bègue, A., Ogden, D. & Emiliani, V. Proc. Natl Acad. Sci. USA 108, 19504–19509 (2011).
Prakash, R. et al. Nat. Methods 9, 1171–1179 (2012).
Packer, A. M. et al. Nat. Methods 9, 1202–1205 (2012).
Akerboom, J. et al. Front. Mol. Neurosci. 6, 2 (2013).
Hochbaum, D. R. et al. Nat. Methods 11, 825–833 (2014).
Theis, L. et al. Neuron 90, 471–482 (2016).
Ronzitti, E. et al. J. Neurosci. 37, 10679–10689 (2017).
Rickgauer, J. P. & Tank, D. W. Proc. Natl Acad. Sci. USA 106, 15025–15030 (2009).
Gerchberg, R. W. & Saxton, W. O. Optik (Stuttg.) 35, 237–246 (1972).
Oron, D., Tal, E. & Silberberg, Y. Opt. Express 13, 1468–1476 (2005).
Zhu, G., van Howe, J., Durst, M., Zipfel, W. & Xu, C. Opt. Express 13, 2153–2159 (2005).
Papagiakoumou, E., de Sars, V., Oron, D. & Emiliani, V. Opt. Express 16, 22039–22047 (2008).
Papagiakoumou, E., de Sars, V., Emiliani, V. & Oron, D. Opt. Express 17, 5391–5401 (2009).
Papagiakoumou, E. et al. Nat. Methods 7, 848–854 (2010).
Andrasfalvy, B. K., Zemelman, B. V., Tang, J. & Vaziri, A. Proc. Natl Acad. Sci. USA 107, 11981–11986 (2010).
Hernandez, O. et al. Nat. Commun. 7, 11928 (2016).
Pégard, N. C. et al. Nat. Commun. 8, 1228 (2017).
Baker, C. A., Elyada, Y. M., Parra, A. & Bolton, M. M. eLife 5, e14193 (2016).
Shemesh, O. A. et al. Nat. Neurosci. 20, 1796–1806 (2017).
Guizar-Sicairos, M., Thurman, S. T. & Fienup, J. R. Opt. Lett. 33, 156–158 (2008).
Kitamura, K., Judkewitz, B., Kano, M., Denk, W. & Häusser, M. Nat. Methods 5, 61–67 (2008).
Margrie, T. W. et al. Neuron 39, 911–918 (2003).
Watson, B.O., Yuste, R. & Packer, A.M. bioRxiv Preprint at https://www.biorxiv.org/content/early/2016/05/18/054080 (2016).
Pachitariu, M. et al. bioRxiv Preprint at https://www.biorxiv.org/content/early/2017/07/20/061507 (2016).
Clements, J. D. & Bekkers, J. M. Biophys. J. 73, 220–229 (1997).
Okun, M. et al. Nature 521, 511–515 (2015).
Acknowledgements
We thank I. Bianco, H.W. Dalgleish, M. Fisek, B. Judkewitz, P. Latham, D. Selviah and A. Roth for helpful discussions and comments on the manuscript; K. Deisseroth (Stanford University) for access to AAVdj virus; and Bruker Corporation for software support. We thank D. Selviah for support and training of Z.Z. This work was supported by grants from the Wellcome Trust, EMBO, ERC (AdG 695709) and the BBSRC (BB/N009835) to M.H. Z.Z. was supported by the Faculty of Engineering at University College London. L.E.R. was supported by the Medical Research Council (MRC). O.M.G. was supported by the Wellcome Trust and MRC. A.M.P. was supported by the European Commission (Marie Curie International Incoming Fellowship grant no. 328048).
Author information
Authors and Affiliations
Contributions
Z.Z. and L.E.R. developed the closed-loop software. Z.Z. performed all-optical experiments and analyzed data. O.M.G., L.E.R. and A.M.P. performed animal surgeries. A.M.P. performed cell-attached recordings. O.M.G. prepared the sensory stimulation setup. L.E.R., A.M.P. and M.H. conceived the project. Z.Z., L.E.R., A.M.P. and O.M.G. designed detailed experimental protocols. Z.Z., L.E.R., A.M.P. and M.H. wrote the paper. All authors discussed the results and contributed to revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Architecture of the key hardware and software tools in the closed-loop all-optical system.
(a) Schematic diagram of the system architecture. The two-photon microscope is controlled using Prairie View (Bruker Corporation). The directions and power of imaging and photostimulation laser beams are independently driven by two sets of scanning modules and two Pockels cells. A custom written closed-loop interface links the online microscope readout with photostimulation control and sensory stimulation control. PackIO (Watson et al.53) synchronizes and records voltage signals across devices. Commands are sent from the closed-loop interface to PrairieView, the custom SLM control and sensory stimulation control software, via TCP. PMTs, photomultiplier tubes. Note that the SLM control and sensory stimulation control software could also be installed in computer 1. (b) Software architecture of the closed-loop interface: the intensities of the user-selected ROIs (regions-of-interest) are extracted from the raw data stream from Prairie View using PrairieLink (Bruker Corporation). The interface enables users to select regions of interests (ROIs), monitor calcium signal traces online, specify experiment protocols, and save recordings. If the closed-loop intervention is enabled, the intensity values are passed to different processes according to the experiment type (more in Supplementary Note 1). The sockets between the software and the closed-loop interface are read continuously. (c) Software architecture of the sensory stimulation control software. Voltage signals are sent to the piezo controller upon receiving a command from the closed-loop interface (see Supplementary Note 1 for further details). (d) Software architecture of the SLM control software. The phase mask specified by the index in the socket is sent to the SLM controller (Meadowlark Optics). An echo is sent to the client when this process is complete.
Supplementary Figure 2 Electrophysiological validation of single-cell activity clamp.
(a) Zoomed-in somatic calcium signal, action potentials and photostimulation time from Fig. 1d. (b) Calcium signal recorded during activity clamp (red bar, 30 s) from 7 cells in 5 animals (animals were under anesthesia; center value is mean; shaded-area is s.d.). (c) Average calcium signal level during steady state (from the time when the target calcium level was reached to the end of the clamp) versus the target signal level (P = 0.00081, Kruskal-Wallis test; 7 cells in 5 animals). Gray points are individual cells. (d) Spike rate extracted from cell-attached recording during activity clamp of the same cells in (b). The instantaneous firing rate exhibits an initial peak followed by a steady state (center value is mean; shaded-area is s.d.). (e) Average spike rate during steady state (as in (c)) versus target calcium signal levels (P = 0.35, Kruskal-Wallis test). Gray points are individual cells. (f) Photostimulation (10 ms spiral, 30 mW on sample, laser pulse repetition rate 70 ± 2 MHz, Fidelity laser; or 20 ms spiral, 10 mW on sample, laser pulse repetition rate 2 MHz, Satsuma laser) rate during activity clamp from the same cells in (c) (P = 0.21, Kruskal-Wallis test; 7 cells in 5 animals; center value is mean; shaded-area is s.d.). (g) Average photostimulation rate during steady state (as in (e)) versus target calcium signal levels. (h) Photostimulation success decreased throughout the clamp period (coefficients of linear fit R1 = -1.01, R2 = 71.6). % Successful photostimuli, defined as the percentage of photostimuli that activated one or more spikes (counted in 60 ms time window after the onset of each stimulation) out of the total number of photostimuli triggered in the Xth second from the start of clamp (n = 5897 spiral photostimuli). (i) Most (~63 %) photostimuli activated one or more spikes (counted in a 60 ms time window after the onset of each stimulation). (j) In most cells ( > 80%), the calcium level could be clamped to within 30% of the ΔF/F of the target levels. (k) There is a weak correlation of clamp error with baseline spike rate (P = 0.06, r = 0.44, n = 19 trials in 7 cells), which is not correlated with the baseline fluorescence level (P = 0.55, r = –0.07, n = 68 trials in 25 cells).
Supplementary Figure 3 Clamping the activity level of multiple cells simultaneously.
(a) A field of view in layer 2/3 of mouse somatosensory cortex coexpressing GCaMP6s (green) and C1V1-2A-mCherry (pink; scale bar, 50 μm, 30 mW on sample, Chameleon laser). The activity of four cells (white circles) were clamped around 50% ΔF/F for 30 s (red bar shows the clamping time, during which the photostimulation rate and pattern were adjusted according to the online ΔF/F readout, the instantaneous photostimulation rate per cell ranges from 0 to 6 Hz, 6 mW per cell, Satsuma laser). Stimulating all four cells at a fixed rate (3 Hz, 30 s) without closed-loop feedback evokes different ΔF/F profiles in different targets (traces show the averages in three trials, imaged at 30 Hz, downsampled by 5 for display). This is an example representative of 8 independent experiments with similar results. (b) The average activity level during clamping and during fixed-rate photostimulation is similar (n.s. P = 0.57, 103 trials in 8 experiments, 4 animals; 4–6 target cells were clamped simultaneously in each experiment; Wilcoxon signed-rank test (two-sided)), however, the variation between the target cells during clamping (measured as the standard deviation between the calcium traces during 30-s conditioning) is significantly reduced compared to when they were photostimulated at the same time at a fixed rate (***P = 0.0002, Wilcoxon signed-rank test (two-sided); center of value is mean; error bars show s.d.). The animals were awake in these experiments.
Supplementary Figure 4 Closed-loop photostimulation-boosted calcium transients are larger than the linear sum of the calcium transients evoked by the weak sensory stimulation and the replayed photostimuli.
(a) Comparing the calcium transients in the closed-loop trials (same as in Fig. 1k, panel 3) with the linear sum of the calcium transients evoked by the weak sensory stimuli (panel 2 in Fig. 1k) and the replayed photostimuli (panel 4 in Fig. 1k, 90 trials, 6 neurons in 4 animals. Center values are mean; shaded-areas are s.d.). (b) The peak of the stimulus-triggered calcium transients in the closed-loop trials is significantly larger than in the linear sum of the weak sensory stimulation trials and the replayed photostimulation trials (same data as in (a), **P = 0.001, Wilcoxon signed-rank test (two-sided); center of value is mean; error bars show s.d.).
Supplementary Figure 5 Validation of online GCaMP6s event detection.
(a) Online event detection results (red bars) compared with offline event detection results for the example trace (closed-loop range) in Fig. 2c. Online event detection used the average calcium signal within a circular ROI (20 pixels in diameter). Offline event detection (blue bars) was performed on the neuropil-subtracted (Pachitariu et al.54) GCaMP6s calcium trace (smoothed by low-pass Butterworth filter) extracted from motion-corrected (Guizar-Sicairos et al.50) movies using template matching (Clements and Bekkers55). Events indicated by red arrows are false alarms (i.e., events detected by the online algorithm and not by offline analysis); blue arrows indicate missed events (i.e., events detected by the offline algorithm and not by online analysis). (b) Accuracy of the online event detection results compared with the offline results. Sensitivity (the true positive rate), which measures how many ‘true events’ were successfully detected, is defined as the number of events that were detected both online and offline over the total number of events detected offline; precision, which measures how many of the ‘detected events’ were true events, is defined as the number of events that were detected both online and offline over the total number of events detected online. Data are presented as box-and-whisker plots displaying median and interquartile ranges (n = 6 trigger neurons in 3 animals; each trigger neuron was associated with a different group of 5 target neurons).
Supplementary Figure 6 Online feedback speed.
(a) Time from the decision-making frame acquired to the start of photostimulation in the singletrigger-multi-targets experiments (motion correction not enabled, top, 6.96 ± 0.92 ms (mean ± s.d.), n = 139 trials in 6 single-cell readout experiments) and multi-trigger multi-targets experiments (bottom, n = 3 multi (three)-cell readout experiments). The distribution of the feedback time in the multicell readout experiments showed two peaks corresponding to the trials that did not require a phase-mask update (7.01 ± 0.76 ms (mean ± s.d.), n = 90 trials) and the trials in which a different phase mask was loaded onto the SLM (21.46 ± 1.88 ms (mean ± s.d.), n = 426 trials). Data are presented as box-and-whisker plots displaying median and interquartile ranges. (b) Time delay introduced by each element of the closed-loop system. (c) The computation time (fifth row in panel b) increases with the number of monitored regions of interest (ROIs), but remains < 3 ms even when 30 ROIs are read out simultaneously (n = 1,000 frames in each condition; measure of center is mean; error bars show s.d.).
Supplementary Figure 7 Correlation of trigger and target activity during the spontaneous activity and closed-loop control period.
(a) Left: Pearson’s correlation coefficients between the calcium transients of the trigger cells and their associated target cells (–0.04 ± 0.27 (mean ± s.d.) during spontaneous activity (gray) and 0.39 ± 0.21 with closed-loop intervention (red); ****P = 2.9 × 10–7, Wilcoxon rank sum test (two-sided); n = 30 trigger-target pairs of 6 trigger-target groups in 3 animals; same neurons as in Fig. 2d). Right: Pearson’s correlation coefficients between the calcium transients of the target cells during spontaneous activity (gray) and closed-loop intervention (red). ****P = 2.5 × 10–12, Wilcoxon rank sum test (two-sided); n = 60 target-target pairs; same neurons as in Fig. 2d, measure of center is mean). (b) Left: Pearson’s correlation coefficients between the calcium transients of the trigger cells and their associated targets (–0.17 ± 0.30 (mean ± s.d.) during spontaneous recording and 0.32 ± 0.22 during closed-loop intervention; ****P = 1.2 × 10–9, Wilcoxon rank sum test (two-sided); n = 45 trigger-target pairs of 9 trigger-target groups in 3 animals; same neurons as in Fig. 2h). Right: Pearson’s correlation coefficients between the calcium transients of the target cells that were associated with the same trigger cell during spontaneous activity and closed-loop intervention (0.08 ± 0.25 (mean ± s.d.) during spontaneous recording and 0.44 ± 0.27 (mean ± s.d.) during closed-loop intervention; ****P = 1.0 × 10–10, Wilcoxon rank sum test (two-sided); n = 90 target-target pairs; same neurons as in Fig. 2h; measure of center is mean).
Supplementary Figure 8 GCaMP6f event time detection.
(a) An example showing detected events (red vertical lines) and inferred spikes (black) during spontaneous calcium signaling (low-pass filtered, GCaMP6f, gray line). (b) Iterative adjustment of event times. Top: event times (black vertical lines) were initialized as the spike times where the inferred spike amplitude is above mean plus s.d. Bottom: in each iteration, the events that are detected within 150 ms after the preceding event, and those that have no fluorescence intensity increase (ΔF/F < 0) were discarded, and the onsets of the calcium transients (gray line) were adjusted to be the event times (red vertical lines). (c) Normalized calcium transients around detected event times. Left: calcium transients (gray) aligned to the event times after initialization; middle: excluded events that have ΔF/F amplitude < 0; right: calcium transients aligned to the detected event onset times (red vertical line).
Supplementary Figure 9 The rate and amplitude of events in the target cells increased significantly after closed-loop conditioning.
(a) Left: the rate of the detected events in the target cells increased significantly after closed-loop conditioning (****P = 6.4 × 10–11, Wilcoxon signed-rank test (two-sided), n = 57 target cells in 6 animals). Right: the detected event amplitude increased significantly after conditioning (**P = 0.0026, Wilcoxon signed-rank test (two-sided)). (b) Target cells showed only a small increase in the event rate (*P = 0.03, Wilcoxon signed-rank test (two-sided); n = 56 target cells in 6 animals) and no significant change in the event amplitude after open-loop conditioning (n.s. P = 0.29, Wilcoxon signed-rank test (two-sided); measure of center is mean; error bars show s.d.).
Supplementary Figure 10 Photostimulation-triggered activity and spatial distribution of cells in closed-loop and open-loop experiments.
(a) Average calcium transient (mean ± s.d.; frames with photostimulation were interpolated by a spline curve) evoked by photostimulation (stimulation duration 12–25 × 10 ms spirals at 100 Hz; 30 mW per cell, Fidelity laser or 4 mW per cell, Satsuma laser) in closed-loop (n = 57 target cells in 6 animals) and open-loop experiments (n = 56 target cells in 6 animals). (b) The amplitude of the calcium transient (maximum average response in a 500-ms window after the onset of photostimulation, excluding the photostimulation period) in target cells evoked by photostimulation in open-loop (gray, n = 56 target cells in 6 animals) and closed-loop (red, n = 57 target cells in 6 animals) experiments is similar (n.s. P = 0.58, Wilcoxon rank sum test (two-sided)). (c) Average photostimulation rate during 30-min conditioning: 0.139 ± 0.049 Hz in the open-loop (gray) and 0.136 ± 0.042 Hz in the closed-loop condition (red; P = 0.59, Wilcoxon rank sum test (two-sided), n = 6 animals in each condition; measure of center is mean; error bars show s.d.). (d) The average lateral distance between the target cells (left, n.s. P = 0.48) and that between the trigger cell and the target cells are similar in closed-loop and open-loop experiments in Fig. 2 (n.s. P = 0.24, Wilcoxon rank sum test (two-sided). n = 6 animals in each condition). (e) The change in the inferred spike rate after conditioning was not significantly correlated with the level of photostimulation-evoked activity during conditioning. The coefficient of determination (R-squared) of linear regression (outliers that are above/below mean ± s.d. were excluded before fitting) is 0.02 and 0.00 for the open-loop values (n = 55 cells in 6 animals) and the closed-loop values (n = 55 cells in 6 animals), respectively.
Supplementary Figure 11 The increase in spike rate in target cells is larger than that in background cells after closed-loop photostimulation.
Bootstrap test on the change in the inferred spike rate in the target cells and background cells: increase in activity of the target cells was greater than that of 97% ± 2.7% (mean ± s.e., n = 6 animals) of the bootstrap samples of the background cells. Gray area shows the distribution of the change in inferred spike rate of 3,000 randomly selected samples (groups of all detected cells in the field of view (excluding the trigger cell) with the same number of cells as in the targets group); red lines are the upper limit of the 95% bootstrap confidence interval; black arrows locate the average change in the target cells in the distributions.
Supplementary Figure 12 The activity of the trigger and targets became more correlated after closed-loop conditioning.
(a) The inferred spike rate of the target cells and background cells in closed-loop and open-loop conditions aligned to the onsets of the detected events in the trigger cells: after closed-loop conditioning, the inferred spike rate of the target cells but not the background cells became significantly higher during periods when the trigger cell is active (dark gray bar, 170-ms window following an event in the trigger) than that during periods when the trigger cell is less active (n = 57, P = 0.004 for target cells and n = 451, P = 0.61 for background cells, Wilcoxon signed rank test (two-sided)); neither the target cells nor the background cells after open-loop conditioning showed an increase in spike rate that is locked to the events of the trigger cell (n = 56, P = 0.61 for target cells; n = 424, P = 0.17 for background cells. Center values are mean; shaded areas are s.e.). (b) Population coupling within the trigger-targets increased significantly after closed-loop photostimulation (* P = 0.040, Wilcoxon signed-rank test (one-sided), n = 63 pairs of coupling within 57 target cells and 6 trigger cells in 6 animals; center values are mean; error bars are s.d.). The increase in the coupling of one cell to the other cells in the trigger-targets group is negatively correlated with the distance between the target cell to the trigger cell (Pearson’s correlation coefficient r = –0.39, P = 0.0028) and the average distance between the target cell and the other target cells (r = –0.38, P = 0.0033). (c) Pearson’s correlation coefficients between the calcium transients of the trigger cells and their associated targets before and after conditioning (*P = 0.029, Wilcoxon signed-rank test (one-sided); n = 57 pairs of 57 target cells and 6 trigger cells in 6 animals). (d) Pearson’s correlation coefficients between the calcium transients of the target cells that were photostimulated at the same time did not increase (n.s. P = 0.46, Wilcoxon signed-rank test (one-sided); n = 244 pairs of 57 target cells in 6 animals; center values are mean; error bars are s.d.).
Supplementary Figure 13 Opsin and calcium indicator expression analysis and controls.
(a) The photostimulation-evoked change in the fluorescent signal of GCaMP6f-expressing cells is significantly lower than the GCaMP6s-expressing cells (tested by photostimulating groups of 10 cells that have detectable C1V1-mCherry signal with 25 10-ms spirals at 100 Hz, 30 mW per cell, Fidelity laser); P = 7.3 × 10–33, Kolmogorov–Smirnov test (two-sided); n = 1,101 GCaMP6s cells in 6 animals; n = 1,065 GCaMP6f cells in 5 animals). More than 65.8% of all tested GCaMP6s-expressing cells had fluorescence intensity increases (ΔF/F) larger than 20%. (b) The response to photostimulation in GCaMP6s-C1V1 cells in (a) does not correlate with the C1V1 expression level (as indicated by the mean intensity of mCherry fluorescence signal. The coefficient of determination (R-squared) of linear regression is 0.002). (c) Strong photostimulation (100 Hz, 500 ms duration, 30 mW, Fidelity laser; n = 50 trials, averaged responses from 5 cells) produces Ca signals in an animal coexpressing GCaMP6f and C1V1 (right), but not in a GCaMP-only control mouse (left). (d) The increase in spike rate (normalized to baseline spike rate) following closed-loop plasticity induction does not correlate with the C1V1 expression level (Pearson’s correlation coefficient r = 0.018, P = 0.89) or the baseline GCaMP fluorescence signal level (Pearson’s correlation coefficient r = –0.019, P = 0.89, n = 57 cells in 6 animals). Each point is a single target neuron from the experiments shown in Fig. 2m. The coefficient of determination (R-squared) of linear regression (outliers that are above/below mean ± s.d. were excluded before fitting) is 0.00 and 0.02 for the C1V1 expression level and GCaMP expression level (n = 55 cells in 6 animals), respectively.
Supplementary Figure 14 Assessing long-term effects of photostimulation.
(a) The percentage of cells exhibiting nuclear fluorescence is 4.2 ± 2.6% on day 1 (before running the conditioning protocol in Fig. 2), and 4.0 ± 2.4% on day 2 (24–48 h after running the plasticity protocol; 49 cells in 5 FOVs in 5 animals, n.s. P = 1.0, Wilcoxon signed-rank test (two-sided); measure of center is mean; error bars show s.d.). (b) The photostim-triggered ΔF/F of cells in (a) (average amplitude of 10 trials, 4 mW per cell, 15 10-ms spirals at 100 Hz, Satsuma laser) on day 2 is similar to the levels on day 1 (n.s. P = 0.69, Wilcoxon signed-rank test (two-sided); measure of center is mean; error bars show s.d.). (c) The GCaMP6f image, photostimulation-triggered ΔF and the C1V1-mCherry images of the tested cells on day 1 and day 2. The cells marked in red squares are identified as ‘filled’ according to their GCaMP6f intensity (left panel). Note that many of them still showed visible GCaMP responses when activated by photostimulation (middle panel).
Supplementary Figure 15 Quantification of motion-induced artifacts.
(a) During the 30-min closed-loop conditioning in Fig. 2 (animals were head-fixed and were freely running on a running wheel), the lateral shifts of the fields of view are within 5 μm in 99% of all frames (324,000 frames in 6 movies). (b) The average ΔF/F recorded from the trigger cell in the raw movies (without motion correction) is close to that extracted from the registered movie (81% of the differences in ΔF/F values are below 5%).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Note 1 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Zhang, Z., Russell, L.E., Packer, A.M. et al. Closed-loop all-optical interrogation of neural circuits in vivo. Nat Methods 15, 1037–1040 (2018). https://doi.org/10.1038/s41592-018-0183-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0183-z
This article is cited by
-
Real-time analysis of large-scale neuronal imaging enables closed-loop investigation of neural dynamics
Nature Neuroscience (2024)
-
FIOLA: an accelerated pipeline for fluorescence imaging online analysis
Nature Methods (2023)
-
A distributed and efficient population code of mixed selectivity neurons for flexible navigation decisions
Nature Communications (2023)
-
Transparent neural implantable devices: a comprehensive review of challenges and progress
npj Flexible Electronics (2022)
-
All-optical interrogation of neural circuits in behaving mice
Nature Protocols (2022)