Abstract
Optical assays of synaptic strength could facilitate studies of neuronal transmission and its dysregulation in disease. Here we introduce a genetic toolbox for all-optical interrogation of synaptic electrophysiology (synOptopatch) via mutually exclusive expression of a channelrhodopsin actuator and an archaerhodopsin-derived voltage indicator. Optically induced activity in the channelrhodopsin-expressing neurons generated excitatory and inhibitory postsynaptic potentials that we optically resolved in reporter-expressing neurons. We further developed a yellow spine-targeted Ca2+ indicator to localize optogenetically triggered synaptic inputs. We demonstrated synOptopatch recordings in cultured rodent neurons and in acute rodent brain slice. In synOptopatch measurements of primary rodent cultures, acute ketamine administration suppressed disynaptic inhibitory feedbacks, mimicking the effect of this drug on network function in both rodents and humans. We localized this action of ketamine to excitatory synapses onto interneurons. These results establish an in vitro all-optical model of disynaptic disinhibition, a synaptic defect hypothesized in schizophrenia-associated psychosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request. Add Gene numbers for constructs developed in this study are listed in Supplementary Table 1.
References
van den Pol, A. N. Neuropeptide transmission in brain circuits. Neuron 76, 98–115 (2012).
Tritsch, N. X. & Sabatini, B. L. Dopaminergic modulation of synaptic transmission in cortex and striatum. Neuron 76, 33–50 (2012).
Kuner, T. & Augustine, G. J. A genetically encoded ratiometric indicator for chloride: capturing chloride transients in cultured hippocampal neurons. Neuron 27, 447–459 (2000).
Emiliani, V., Cohen, A. E., Deisseroth, K. & Häusser, M. All-optical interrogation of neural circuits. J. Neurosci. 35, 13917–13926 (2015).
Hochbaum, D. R. et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat. Methods 11, 825–833 (2014).
Werley, C. A. et al. All-optical electrophysiology for disease modeling and pharmacological characterization of neurons. Curr. Protoc. Pharmacol. 78, 11.20.1–11.20.24 (2017).
Lou, S. et al. Genetically targeted all-optical electrophysiology with a transgenic Cre-dependent optopatch mouse. J. Neurosci. 36, 11059–11073 (2016).
Shemesh, O. A. et al. Temporally precise single-cell-resolution optogenetics. Nat. Neurosci. 20, 1796–1806 (2017).
Krystal, J. H. et al. Subanesthetic effects of the NMDA antagonist, ketamine, in humans: psychotomimetic, perceptual, cognitive, and neuroendocrine effects. Arch. Gen. Psychiatry 51, 199–214 (1994).
Kraguljac, N. et al. Ketamine modulates hippocampal neurochemistry and functional connectivity: a combined magnetic resonance spectroscopy and resting-state fMRI study in healthy volunteers. Mol. Psychiatry 22, 562–569 (2017).
Jackson, M. E., Homayoun, H. & Moghaddam, B. NMDA receptor hypofunction produces concomitant firing rate potentiation and burst activity reduction in the prefrontal cortex. Proc. Natl. Acad. Sci. USA 101, 8467–8472 (2004).
Di Lazzaro, V. et al. Ketamine increases human motor cortex excitability to transcranial magnetic stimulation. J. Physiol. 547, 485–496 (2003).
Hamm, J. P., Peterka, D. S., Gogos, J. A. & Yuste, R. Altered cortical ensembles in mouse models of schizophrenia. Neuron 94, 153–167 (2017).
Izumi, Y. & Zorumski, C. F. Metaplastic effects of subanesthetic ketamine on CA1 hippocampal function. Neuropharmacology 86, 273–281 (2014).
Grunze, H. C. R. et al. NMDA-dependent modulation of CA1 local circuit inhibition. J. Neurosci. 76, 2034–2043 (1996).
Lisman, J. E. et al. Circuit-based framework for understanding neurotransmitter and risk gene interactions in schizophrenia. Trends Neurosci. 31, 234–242 (2008).
Adam, Y. et al. All-optical electrophysiology reveals brain-state dependent changes in hippocampal subthreshold dynamics and excitability. bioRxiv Preprint at https://doi.org/10.1101/281618 (2018).
Gradinaru, V. et al. Molecular and cellular approaches for diversifying and extending optogenetics. Cell 141, 154–165 (2010).
Branda, C. S. & Dymecki, S. M. Talking about a revolution: the impact of site-specific recombinases on genetic analyses in mice. Dev. Cell. 6, 7–28 (2004).
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. A FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Saunders, A., Johnson, C. A. & Sabatini, B. L. Novel recombinant adeno-associated viruses for Cre activated and inactivated transgene expression in neurons. Front. Neural Circuits 6, 47 (2012).
Petreanu, L., Mao, T., Sternson, S. M. & Svoboda, K. The subcellular organization of neocortical excitatory connections. Nature 457, 1142–1145 (2009).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. eL ife 5, e12727 (2016).
Fischer, M., Kaech, S., Knutti, D. & Matus, A. Rapid actin-based plasticity in dendritic spines. Neuron 20, 847–854 (1998).
Burkel, B. M., Von Dassow, G. & Bement, W. M. Versatile fluorescent probes for actin filaments based on the actin-binding domain of utrophin. Cell Motil. Cytoskeleton 64, 822–832 (2007).
Gross, G. et al. Recombinant probes for visualizing endogenous synaptic proteins in living neurons. Neuron 78, 971–985 (2013).
Baker, C. A., Elyada, Y. M., Parra-Martin, A. & Bolton, M. Cellular resolution circuit mapping in mouse brain with temporal-focused excitation of soma-targeted channelrhodopsin. eLife 5, 1–15 (2016).
Gerfen, C. R., Paletzki, R. & Heintz, N. GENSAT BAC cre-recombinase driver lines to study the functional organization of cerebral cortical and basal ganglia circuits. Neuron 80, 1368–1383 (2013).
Edwards, F. A., Konnerth, A. & Sakmann, B. Quantal analysis of inhibitory synaptic transmission in the dentate gyrus of rat hippocampal slices: a patch-clamp study. J. Physiol. 430, 213–249 (1990).
Dingledine, R. Brain Slices (Springer Science & Business Media, New York, 2013).
Potter, G. B. et al. Generation of Cre-transgenic mice using Dlx1/Dlx2 enhancers and their characterization in GABAergic interneurons. Mol. Cell. Neurosci. 40, 167–186 (2009).
Poitras, L., Ghanem, N., Hatch, G. & Ekker, M. The proneural determinant MASH1 regulates forebrain Dlx1/2 expression through the I12b intergenic enhancer. Development 134, 1755–1765 (2007).
Colasante, G. et al. Rapid conversion of fibroblasts into functional forebrain GABAergic interneurons by direct genetic reprogramming. Cell Stem Cell 17, 719–734 (2015).
McCormick, D. A., Connors, B. W., Lighthall, J. W. & Prince, D. A. Comparative electrophysiology of pyramidal and sparsely spiny stellate neurons of the neocortex. J. Neurophysiol. 54, 782–806 (1985).
Kwan, A. C. & Dan, Y. Dissection of cortical microcircuits by single-neuron stimulation in vivo. Curr. Biol. 22, 1459–1467 (2012).
McNally, J. M., McCarley, R. W., McKenna, J. T., Yanagawa, Y. & Brown, R. E. Complex receptor mediation of acute ketamine application on in vitro gamma oscillations in mouse prefrontal cortex: modeling gamma band oscillation abnormalities in schizophrenia. Neuroscience 199, 51–63 (2011).
Kinney, J. W. A specific role for NR2A-containing NMDA receptors in the maintenance of parvalbumin and GAD67 immunoreactivity in cultured interneurons. J. Neurosci. 26, 1604–1615 (2006).
Xi, D., Keeler, B., Zhang, W., Houle, J. D. & Gao, W. J. NMDA receptor subunit expression in GABAergic interneurons in the prefrontal cortex: application of laser microdissection technique. J. Neurosci. Methods 176, 172–181 (2009).
Paoletti, P. & Neyton, J. NMDA receptor subunits: function and pharmacology. Curr. Opin. Pharmacol. 7, 39–47 (2007).
Jones, R. S. G. & Bühl, E. H. Basket-like interneurones in layer II of the entorhinal cortex exhibit a powerful NMDA-mediated synaptic excitation. Neurosci. Lett. 149, 35–39 (1993).
Klein, J. S., Jiang, S., Galimidi, R. P., Keefe, J. R. & Bjorkman, P. J. Design and characterization of structured protein linkers with differing flexibilities. Protein Eng. Des. Sel. 27, 325–330 (2014).
Geraerts, M., Micheils, M., Baekelandt, V., Debyser, Z. & Gijsbers, R. Upscaling of lentiviral vector production by tangential flow filtration. J. Gene Med. 7, 1299–1310 (2005).
Nehme, A. R. et al. Combining NGN2 programming with developmental patterning generates human excitatory neurons with NMDAR-mediated synaptic transmission. Cell Rep. 23, 2509–2523 (2018).
Kralj, J. M., Douglass, A. D., Hochbaum, D. R., Maclaurin, D. & Cohen, A. E. Optical recording of action potentials in mammalian neurons using a microbial rhodopsin. Nat. Methods 9, 90–95 (2012).
Acknowledgements
We thank V. Joshi, K. Williams, M. Lee, and S. Begum for technical assistance. We thank V. Parot for help with analysis of spine Ca2+ data, S. Turney and the Harvard Center Brain Science (CBS) for loan of optical equipment, B. Sabatini (Harvard Medical School, Boston, MA, USA) for Rbp4-Cre mice, and C. Werley and L. Williams (Q-State Biosciences, Cambridge, MA, USA) for the mI12b plasmid. This work was supported by the Howard Hughes Medical Institute and a grant from the Gordon and Betty Moore Foundation. Y.A. was supported by a long-term fellowship from the Human Frontiers Science Program and by a postdoctoral fellowship from the Edmund and Lili Safra center for Brain Sciences. D.B.A. and E.S.J. were supported by NIH grant NS-089491. K.E. and R.N. were supported by the Stanley Center and NIMH grants (grant nos. 5U01MH105669-04 and 1U01MH115727-01).
Author information
Authors and Affiliations
Contributions
L.Z.F. and A.E.C. conceived synOptopatch. L.Z.F. designed the synOptopatch and system, built the dual-view system, and acquired the optical electrophysiology data. L.Z.F. and A.E.C. analyzed the data and wrote the paper. L.Z.F., R.N., K.E., and A.E.C. designed the approach for implementation in human neurons. D.B.A., E.S.J., and L.Z.F. engineered spine-jRGECO1a. Y.A. created QuasAr2-Citrine, designed soma-localized QuasAr2 and CheRiff, and developed the brain slice imaging system. H.W. set up the patterned illumination system. All authors participated in revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.E.C. and K.E. are founders of Q-State Biosciences. L.Z.F. and A.E.C. have filed a patent related to synOptopatch.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Co-expression of CheRiff- and QuasAr2-introduced optical cross-talk in measurements of synaptic transmission.
a, In neurons co-expressing CheRiff-CFP and QuasAr2-Citrine, the QuasAr2 citrine fluorescence reported action potentials with high signal-to-noise ratio. Left: images of QuasAr2 fluorescence. Scale bar, 20 μm. Right: fluorescence of QuasAr2 during optogenetic stimulation, recorded at 500 Hz. Blue: optogenetic stimulation waveform. b, Top: red: QuasAr2 fluorescence, blue: DMD mask for patterned blue light stimulation. Scale bar, 10 μm. Bottom: fluorescence signal of the circled cells before (red) and after (black) addition of excitatory blockers NBQX (20 μM) and AP5 (50 μM). The persistence of the signal indicates that it arose from direct optogenetic stimulation of a postsynaptic neurite rather than from synaptic transmission.
Supplementary Figure 2 Redundant use of Cre recombination sites caused spurious cross-reactivity.
a, Schematic of DIO Cre-on CheRiff-CFP and DO Cre-off QuasAr2-mOrange. b, Neurons co-infected with DIO Cre-on CheRiff-CFP and Cre-GFP showed Cre-activated expression. c, Neurons co-infected with DO Cre-off QuasAr2-mOrange and Cre-GFP showed Cre-inactivated expression. d, Neurons co-infected with DIO CheRiff-CFP, DO QuasAr2-mOrange and Cre did not show mutually exclusive expression of the actuator and reporter. Scale bars in b–d, 20 μm (n in b–d = 3 culture dishes; representative data are shown). e, Quantification of the data in (d) showing lack of mutually exclusive expression (n = 102 neurons). f, Top: schematic of DIO Cre-on CheRiff-CFP and FAS Cre-off QuasAr2-Citrine. Bottom: mean expression levels of FAS Cre-off QuasAr2-Citrine depended on the titers of the FAS Cre-off QuasAr2-Citrine lentiviruses (n = 391 neurons). g, Mean expression levels in expressing cells of (left, n = 112 neurons) FAS Cre-off QuasAr2-Citrine and (right, n = 149 neurons) DIO Cre-on CheRiff-CFP did not depend on the titer of the Cre virus. h, SynOptopatch constructs mediated mutually exclusive expression of actuator and reporter. No co-expression of DIO Cre-on CheRiff-CFP and FAS Cre-off QuasAr2-Citrine was observed across a range of titers of Cre virus. All statistics are mean ± s.e.m. All error bars, s.e.m.
Supplementary Figure 3 Calibration of the synOptopatch constructs.
a, Simultaneous fluorescence and manual patch-clamp recordings of PSPs and action potentials evoked by presynaptic optogenetic stimulation. Blue, 10 ms blue light stimulation; red, whole-cell single-trial unfiltered fluorescence; black, patch-clamp recordings. b, Overlay of mean optically and electrically recorded PSP waveforms from a single cell (n = 9 repeats). c, Quantification of fluorescence versus postsynaptic potential for synOptopatch recordings (n = 10 neurons, R2 = 0.9). d, Quantification of photocurrent of QuasAr2 under red and blue illumination (red: 400 W cm–2; blue: 10 ms, 20 – 120 mW cm–2). e,f, Quantification of optical crosstalk of blue illumination into QuasAr2 fluorescence. e, Neurons expressing QuasAr2 were exposed to continuous excitation at 640 nm and pulses of illumination at 488 nm (10 ms, 60 mW cm–2) (n = 17 neurons). f, Quantification of crosstalk amplitude as a function of blue light intensity. The shading represents the range of blue light intensity used for optogenetic stimulation of primary neurons. Error bars represent s.e.m. g, SynOptopatch fluorescence recordings of EPSPs were stable for at least 2 h. h, Photobleaching of QuasAr2 in the synOptopatch assay. A neuron was illuminated for 30 minutes continuously at 640 nm, 400 W cm–2 and probed at 60 s intervals with blue light to induce PSPs (2 pulses of 10 ms, 2 Hz, 60 mW cm–2). i, Stability of PSP waveform recorded as a function of duration of continuous red illumination. (In h,i, n = 3 neurons; representative data are shown.) All shaded error bars, s.e.m.
Supplementary Figure 4 SynOptopatch detects synaptic plasticity.
a, Schematic showing protocol for inducing homeostatic plasticity. b, A typical ten-trial average trace of a same neuron measured before gabazine incubation (left) and when wash-out after 48 h incubation with gabazine (right). c, Neurons incubated for 48 h with gabazine had increased ratio of IPSP amplitude over EPSP amplitude (n = 9 neurons, **P = 0.04, two-sided paired-sample t-test), while in d control cells did not show significant increase in the ratio of IPSP amplitude over EPSP amplitude (n = 5 neurons, P = 0.76, two-sided paired-sample t-test). All error bars, s.e.m. All statistics are mean ± s.e.m.
Supplementary Figure 5 SynOptopatch in hiPSC-derived neurons.
a, Schematic of synOptopatch in hiPSC-derived neurons. b, Schematic of excitatory cortical neuron differentiation, gene transduction, maturation and measurement. c, Blue light stimulation induced single-trial PSPs and action potentials (red: before; black: after NBQX addition). Due to lower CheRiff expression than in primary neurons, stimulation required higher blue light intensities (400 mW cm–2 versus 20–120 mW cm–2 for primary neurons) to evoke presynaptic action potentials (n = 12 neurons; representative data are shown). d, PSPs were stable for 1 h (red: before; black: 1 h without perturbation). Right: quantification of AUC before and after 1 h (AUC, A.U., 0.58 ± 0.16 versus 0.50 ± 0.11, n = 7 neurons, P = 0.4, two-sided paired-sample t-test). e, A synaptic blocker (25 μM NBQX) largely eliminated the PSPs. The residual transient in the presence of NBQX occurred concurrent with the blue light stimulation, indicating that this signal arose from blue light crosstalk, a consequence of the higher blue stimulation power needed (also visible in d and f). Right: quantification of AUC before and after addition of NBQX (AUC, A.U., 0.97 ± 0.1 versus 0.53 ± 0.07, n = 12 neurons, ***P = 2 × 10−5, two-sided paired-sample t-test). f, A positive allosteric modulator of AMPARs and kainate receptors (CTZ, 50 μM) increased the PSP amplitude. Right: quantification of AUC before and after addition of CTZ (AUC, A.U., 0.63 ± 0.26 versus 1.4 ± 0.48, n = 6 neurons, *P = 0.055, two-sided paired-sample t-test). g, Exclusive expression of QuasAr2 and CheRiff in hiPSC-derived neurons. Green: presynaptic cells expressing CheRiff-CFP. Red: postsynaptic cells expressing QuasAr2-Citrine, showing Citrine fluorescence. All scale bars, 10 μm (n = 3 culture dishes; representative data are shown) All shaded error bars and error bars, s.e.m. All statistics are mean ± s.e.m.
Supplementary Figure 6 NMDAR component of postsynaptic potential.
To isolate the NMDAR component of the PSP, we added 10 μM NBQX and 10 μM picrotoxin to block AMPA and GABAa receptors, respectively, and imaged in a medium containing 0 Mg2+ to remove the voltage-dependent Mg2+ NMDAR block. Ten-trial average (n = 10 neurons; representative data are shown).
Supplementary Figure 7 Targeting GCaMP6s and jRGECO1a to dendritic spines with and without transcriptional regulatory system.
a, Image of a neuron expressing GCaMP6s fused to the CH domain of Utrophin (GCaMP6s-Utr) with zinc finger binding sequence. GCaMP6s was stained with anti-GFP antibody. GCaMP6s co-localized with endogenous PSD-95, a marker for dendritic spines. PSD95 puncta not associated with GCaMP6s puncta were due to spines from neighboring neurons not expressing GCaMP6s. Scale bar, 10 μm. b, GCaMP6s-Utr without ZFBS exhibited more diffuse localization compared to regulated GCaMP6s-Utr. c,d, Examples from additional neurons as in a and b, respectively (a–d, n = 5 neurons; representative data are shown). e,f, Example fluorescence line sections transecting two spines and a parent dendrite along the lines shown in a and b, respectively. g, Quantification of ratio of fluorescence in spines to adjacent parent dendrites in cells expressing GCaMP6-Utr with or without the ZFBS (n = 5 neurons of each type, ~200 spines per neuron, **P = 0.027, two-sided t-test). h, Quantification of spine fluorescence as a function of distance from the soma (n = 5 neurons of each type, ~200 spines per neuron). i, Spine-jRGECO1a and cytosolic jRGECO1a expressed in HEK293 cells gave the same fluorescence response to a Ca2+ transient induced by 10 μM ionomycin (ΔF/F 1.6 ± 0.1 versus 1.5 ± 0.07, n = 26 cells for spine-jRGECo1a, n = 29 for cytosolic jRGECO1a, P = 0.4, two-sided student’s t-test). All error bars, s.e.m. All statistics are mean ± s.e.m.
Supplementary Figure 8 Simultaneous imaging of spine-jRGECO1a and QuasAr2.
a, Co-expression of spine-jRGECO1a and QuasAr2. Schematic showing three color imaging with blue light excitable soma-localized CheRiff in presynaptic cells, yellow light excitable spine-jRGECO1a and red light excitable QuasAr2 co-expressed in postsynaptic cells. b, Dual-view microscope for simultaneous imaging of spine-jRGECO1a and QuasAr2. Fluorescence from spine-jRGECO1a and QuasAr2 were passed to electron-multiplying charge-coupled-device camera and sCMOS camera, respectively, by a 640 nm dichroic beamsplitter.
Supplementary Figure 9 Correlation of synaptic and bAP-induced Ca2+ activity in dendritic spines.
a, Red: mean action potential and PSP reported by QuasAr2; orange: mean spine-jRGECO1a fluorescence induced by bAP and sub-threshold synaptic inputs, respectively. b, Distribution of Ca2+ transient amplitudes in spines, induced by bAPs (red) or subthreshold synaptic inputs (brown). Synaptic inputs only drove Ca2+ transients in a subset of synaptic spines, while bAPs drove Ca2+ signals synchronously in almost all dendritic spines. c, Correlation of somatic PSPs and sum of Ca2+ transient amplitude across all spines (n = 14, R2 = 0.42, P = 0.01). d, bAP failure in spines. Blue: 10 ms blue light stimulation of soma-localized CheRiff in a presynaptic cell. Red: postsynaptic QuasAr2 fluorescence; dark yellow: average over all the spines of spine-jRGECO1a fluorescence (n = 100 spines). Right: single-spine fluorescence traces within a single dendritic branch, corresponding to the regions circled in the figure. The orange trace shows an example of a bAP-induced spine Ca2+ transient distal to a spine that did not respond to the bAP. Scale bar, 15 μm. All shaded error bars, s.e.m.
Supplementary Figure 10 IPSPs in acute slices under high K+ concentration.
IPSP detected under the condition of 5 mM K+, twice higher than usual ACSF, and 10 μM NBQX, 10 μM AP5. Blue: 10 ms blue light stimulation. Light and dark red: IPSPs. Scale bar, 10 μm. Five-trial average (n = 8 cells; representative data are shown).
Supplementary Figure 11 Development and characterization of an inhibitory neuron-specific enhancer.
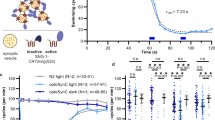
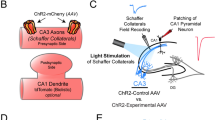
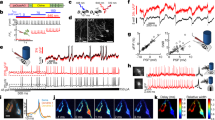
a, The inhibitory marker Gad67 co-localized with eGFP expression. Left: mI12b-eGFP; Right: anti-GAD67 immunostaining (Methods). Scale bars, 30 μm (n = 39 cells; representative data are shown). b, Correspondence of mI12b-eGFP fluorescence and Gad67 immunostaining in neurons that were positive for at least one of these reporters. Of n = 41 Gad67+ neurons, 39 were mI12b-eGFP+. Of n = 41 mI12b-eGFP+ neurons, 39 were also Gad67+. c, Top: Schematic showing patterned light stimulation onto an mI12b-eGFP+ neuron and voltage imaging from a nearby cell expressing QuasAr2. Middle: dark green: mI12b-eGFP, red: QuasAr2-dark Citrine, blue: DMD mask for patterned blue light stimulation. Right: IPSP was evoked by stimulation of the presynaptic cell expressing mI12b-EGFP. Scale bar, 10 μm. Two-trial average (n = 3 cells; representative data are shown). d, Optopatch measurements of spiking patterns and action potential waveforms revealed differences between mI12b-eGFP+ and mI12b-eGFP− neurons consistent with differences between inhibitory versus excitatory neurons reported by patch-clamp measurements (Methods).35,36 Blue: blue light stimulation of cells expressing both CheRiff and QuasAr2. Example traces show QuasAr2 fluorescence from a nominal excitatory neuron (orange, mI12b-EGFP−) and a nominal inhibitory neuron (green, mI12b-EGFP+). Middle: raster plots of spiking. Bottom: spike rate of nominal excitatory neurons (orange, mI12b-EGFP−) and inhibitory neurons (green, mI12b-EGFP+). e, Spiking rate and f, spiking waveform of cells not expressing mI12b (orange) and expressing mI12b (dark green). All shaded error bars and error bars, s.e.m. mI12b-eGFP+ neurons (putative interneurons) showed higher average evoked firing rates (13.6 ± 1.7 versus 6.7 ± 1 Hz, P = 0.001, two-sided two-sample t-test), lower probability of depolarization block under strong stimulus, narrower action potentials (6.9 ± 0.2 ms versus 9.4 ± 0.4 ms, P = 1 × 10−6, two-sided two-sample t-test) and larger after-hyperpolarization compared to simultaneously measured mI12b-eGFP− cells (n = 36 mI12b-eGFP+ and 32 mI12b-eGFP− neurons).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Table 1
Supplementary Video 1
Spine-jRGECO1a in a cultured rat hippocampal neuron. The neuron receives uncorrelated synaptic inputs that activate a subset of spines and also produces action potentials that activate most spines synchronously. The movie was acquired at 20 frames s−1 and is shown at 10 frames s−1
Supplementary Video 2
Spine-GCaMP6s in a cultured rat hippocampal neuron. The neuron receives uncorrelated synaptic inputs that activate a subset of spines and also produces action potentials that activate most spines synchronously. The movie was acquired at 20 frames s−1 and is shown at 10 frames s−1
Rights and permissions
About this article
Cite this article
Fan, L.Z., Nehme, R., Adam, Y. et al. All-optical synaptic electrophysiology probes mechanism of ketamine-induced disinhibition. Nat Methods 15, 823–831 (2018). https://doi.org/10.1038/s41592-018-0142-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0142-8
This article is cited by
-
A positively tuned voltage indicator for extended electrical recordings in the brain
Nature Methods (2023)
-
Multiregion neuronal activity: the forest and the trees
Nature Reviews Neuroscience (2022)
-
A far-red hybrid voltage indicator enabled by bioorthogonal engineering of rhodopsin on live neurons
Nature Chemistry (2021)
-
Ketamine disinhibits dendrites and enhances calcium signals in prefrontal dendritic spines
Nature Communications (2020)
-
Neurobiology of rapid-acting antidepressants: convergent effects on GluA1-synaptic function
Molecular Psychiatry (2019)