Abstract
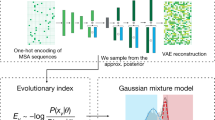
The functions of proteins and RNAs are defined by the collective interactions of many residues, and yet most statistical models of biological sequences consider sites nearly independently. Recent approaches have demonstrated benefits of including interactions to capture pairwise covariation, but leave higher-order dependencies out of reach. Here we show how it is possible to capture higher-order, context-dependent constraints in biological sequences via latent variable models with nonlinear dependencies. We found that DeepSequence (https://github.com/debbiemarkslab/DeepSequence), a probabilistic model for sequence families, predicted the effects of mutations across a variety of deep mutational scanning experiments substantially better than existing methods based on the same evolutionary data. The model, learned in an unsupervised manner solely on the basis of sequence information, is grounded with biologically motivated priors, reveals the latent organization of sequence families, and can be used to explore new parts of sequence space.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The sequence data and code supporting this work are available at https://github.com/debbiemarkslab/DeepSequence. The mutation-effects data from all analyzed experiments, as well as all model predictions, are available in Supplementary Table 2.
References
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Kosuri, S. & Church, G. M. Large-scale de novo DNA synthesis: technologies and applications. Nat. Methods 11, 499–507 (2014).
Romero, P. A., Tran, T. M. & Abate, A. R. Dissecting enzyme function with microfluidic-based deep mutational scanning. Proc. Natl Acad. Sci. USA 112, 7159–7164 (2015).
Roscoe, B. P. & Bolon, D. N. Systematic exploration of ubiquitin sequence, E1 activation efficiency, and experimental fitness in yeast. J. Mol. Biol. 426, 2854–2870 (2014).
Roscoe, B. P., Thayer, K. M., Zeldovich, K. B., Fushman, D. & Bolon, D. N. Analyses of the effects of all ubiquitin point mutants on yeast growth rate. J. Mol. Biol. 425, 1363–1377 (2013).
Melamed, D., Young, D. L., Gamble, C. E., Miller, C. R. & Fields, S. Deep mutational scanning of an RRM domain of the Saccharomyces cerevisiae poly(A)-binding protein. RNA 19, 1537–1551 (2013).
Stiffler, M. A., Hekstra, D. R. & Ranganathan, R. Evolvability as a function of purifying selection in TEM-1 β-lactamase. Cell 160, 882–892 (2015).
McLaughlin, R. N. Jr, Poelwijk, F. J., Raman, A., Gosal, W. S. & Ranganathan, R. The spatial architecture of protein function and adaptation. Nature 491, 138–142 (2012).
Kitzman, J. O., Starita, L. M., Lo, R. S., Fields, S. & Shendure, J. Massively parallel single-amino-acid mutagenesis. Nat. Methods 12, 203–206 (2015).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T. S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Araya, C. L. et al. A fundamental protein property, thermodynamic stability, revealed solely from large-scale measurements of protein function. Proc. Natl Acad. Sci. USA 109, 16858–16863 (2012).
Firnberg, E., Labonte, J. W., Gray, J. J. & Ostermeier, M. A comprehensive, high-resolution map of a gene’s fitness landscape. Mol. Biol. Evol. 31, 1581–1592 (2014).
Starita, L. M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Rockah-Shmuel, L., Tóth-Petróczy, Á. & Tawfik, D. S. Systematic mapping of protein mutational space by prolonged drift reveals the deleterious effects of seemingly neutral mutations. PLoS Comput. Biol. 11, e1004421 (2015).
Jacquier, H. et al. Capturing the mutational landscape of the beta-lactamase TEM-1. Proc. Natl Acad. Sci. USA 110, 13067–13072 (2013).
Qi, H. et al. A quantitative high-resolution genetic profile rapidly identifies sequence determinants of hepatitis C viral fitness and drug sensitivity. PLoS Pathog. 10, e1004064 (2014).
Wu, N. C. et al. Functional constraint profiling of a viral protein reveals discordance of evolutionary conservation and functionality. PLoS Genet. 11, e1005310 (2015).
Mishra, P., Flynn, J. M., Starr, T. N. & Bolon, D. N. A. Systematic mutant analyses elucidate general and client-specific aspects of Hsp90 function. Cell Rep. 15, 588–598 (2016).
Doud, M. B. & Bloom, J. D. Accurate measurement of the effects of all amino-acid mutations to influenza hemagglutinin. bioRxiv Preprint at https://www.biorxiv.org/content/early/2016/04/07/047571 (2016).
Deng, Z. et al. Deep sequencing of systematic combinatorial libraries reveals β-lactamase sequence constraints at high resolution. J. Mol. Biol. 424, 150–167 (2012).
Starita, L. M. et al. Activity-enhancing mutations in an E3 ubiquitin ligase identified by high-throughput mutagenesis. Proc. Natl Acad. Sci. USA 110, E1263–E1272 (2013).
Aakre, C. D. et al. Evolving new protein-protein interaction specificity through promiscuous intermediates. Cell 163, 594–606 (2015).
Julien, P., Miñana, B., Baeza-Centurion, P., Valcárcel, J. & Lehner, B. The complete local genotype-phenotype landscape for the alternative splicing of a human exon. Nat. Commun. 7, 11558 (2016).
Li, C., Qian, W., Maclean, C. J. & Zhang, J. The fitness landscape of a tRNA gene. Science 352, 837–840 (2016).
Mavor, D. et al. Determination of ubiquitin fitness landscapes under different chemical stresses in a classroom setting. eLife 5, e15802 (2016).
Gasperini, M., Starita, L. & Shendure, J. The power of multiplexed functional analysis of genetic variants. Nat. Protoc. 11, 1782–1787 (2016).
Starita, L. M. et al. Variant interpretation: functional assays to the rescue. Am. J. Hum. Genet. 101, 315–325 (2017).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Hecht, M., Bromberg, Y. & Rost, B. Better prediction of functional effects for sequence variants. BMC Genomics 16, S1 (2015).
Huang, Y.-F., Gulko, B. & Siepel, A. Fast, scalable prediction of deleterious noncoding variants from functional and population genomic data. Nat. Genet. 49, 618–624 (2017).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Ng, P. C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Finn, R. D. et al. HMMER web server: 2015 update. Nucleic Acids Res. 43, W30–W38 (2015).
Hopf, T. A. et al. Mutation effects predicted from sequence co-variation. Nat. Biotechnol. 35, 128–135 (2017).
Mann, J. K. et al. The fitness landscape of HIV-1 gag: advanced modeling approaches and validation of model predictions by in vitro testing. PLoS Comput. Biol. 10, e1003776 (2014).
Figliuzzi, M., Jacquier, H., Schug, A., Tenaillon, O. & Weigt, M. Coevolutionary landscape inference and the context-dependence of mutations in beta-lactamase TEM-1. Mol. Biol. Evol. 33, 268–280 (2016).
Lapedes, A., Giraud, B. & Jarzynski, C. Using sequence alignments to predict protein structure and stability with high accuracy. arXiv Preprint at https://arxiv.org/abs/1207.2484 (2012).
Weinreich, D. M., Lan, Y., Wylie, C. S. & Heckendorn, R. B. Should evolutionary geneticists worry about higher-order epistasis? Curr. Opin. Genet. Dev. 23, 700–707 (2013).
Bendixsen, D. P., Østman, B. & Hayden, E. J. Negative epistasis in experimental RNA fitness landscapes. J. Mol. Evol. 85, 159–168 (2017).
Rodrigues, J. V. et al. Biophysical principles predict fitness landscapes of drug resistance. Proc. Natl Acad. Sci. USA 113, E1470–E1478 (2016).
Echave, J. & Wilke, C. O. Biophysical models of protein evolution: understanding the patterns of evolutionary sequence divergence. Annu. Rev. Biophys. 46, 85–103 (2017).
Schmidt, M. & Hamacher, K. Three-body interactions improve contact prediction within direct-coupling analysis. Phys. Rev. E 96, 052405 (2017).
Roweis, S. & Ghahramani, Z. A unifying review of linear gaussian models. Neural Comput. 11, 305–345 (1999).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Kingma, D. P. & Welling, M. Auto-encoding variational Bayes. arXiv Preprint at https://arxiv.org/abs/1312.6114 (2013).
Rezende, D. J., Mohamed, S. & Wierstra, D. Stochastic backpropagation and approximate inference in deep generative models. arXiv Preprint at https://arxiv.org/abs/1401.4082 (2014).
Gómez-Bombarelli, R. et al. Automatic chemical design using a data-driven continuous representation of molecules. arXiv Preprint at https://arxiv.org/abs/1610.02415 (2016).
Wainwright, M. J. & Jordan, M. I. Graphical Models, Exponential Families, and Variational Inference (Now Publishers, Hanover, MA, 2008).
Ingraham, J. & Marks, D. in Proceedings of the 34th International Conference on Machine Learning Vol. 70 (eds Precup, D. & Teh, Y. W.) 1607–1616 (PMLR/Microtome Publishing, Brookline, MA, 2017).
Kingma, D. P. et al. in Advances in Neural Information Processing Systems 29 (eds Lee, D. D. et al.) 4743–4751 (Curran Associates, Red Hook, NY, 2016).
Murphy, K. P. Machine Learning: A Probabilistic Perspective (MIT Press, Cambridge, MA, 2012).
Srivastava, N., Hinton, G. E., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Hopf, T. A. et al. Three-dimensional structures of membrane proteins from genomic sequencing. Cell 149, 1607–1621 (2012).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS One 6, e28766 (2011).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Natl Acad. Sci. USA 108, E1293–E1301 (2011).
Jones, D. T., Singh, T., Kosciolek, T. & Tetchner, S. MetaPSICOV: combining coevolution methods for accurate prediction of contacts and long range hydrogen bonding in proteins. Bioinformatics 31, 999–1006 (2015).
Sim, N. L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–W457 (2012).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum. Genet. 76, 7.20.1–7.20.41 (2013).
Tubiana, J., Cocco, S. & Monasson, R. Learning protein constitutive motifs from sequence data. arXiv Preprint at https://arxiv.org/abs/1803.08718 (2018).
Sinai, S., Kelsic, E., Church, G. M. & Nowak, M. A. Variational auto-encoding of protein sequences. arXiv Preprint at https://arxiv.org/abs/1712.03346 (2017).
Rezende, D. J. & Mohamed, S. Variational inference with normalizing flows. arXiv Preprint at https://arxiv.org/abs/1505.05770 (2015).
Burda, Y., Grosse, R. & Salakhutdinov, R. Importance weighted autoencoders. arXiv Preprint at https://arxiv.org/abs/1509.00519 (2015).
Johnson, M., Duvenaud, D. K., Wiltschko, A., Adams, R. P. & Datta, S. R. in Advances in Neural Information Processing Systems 29 (eds Lee, D. D. et al.) 2946–2954 (Curran Associates, Red Hook, NY, 2016).
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015).
Weinreb, C. et al. 3D RNA and functional interactions from evolutionary couplings. Cell 165, 963–975 (2016).
Toth-Petroczy, A. et al. Structured states of disordered proteins from genomic sequences. Cell 167, 158–170 (2016).
Boucher, J. I., Bolon, D. N. & Tawfik, D. S. Quantifying and understanding the fitness effects of protein mutations: laboratory versus nature. Protein Sci. 25, 1219–1226 (2016).
Doud, M. B. & Bloom, J. D. Accurate measurement of the effects of all amino-acid mutations on influenza hemagglutinin. Viruses 8, 155 (2016).
Wrenbeck, E. E., Azouz, L. R. & Whitehead, T. A. Single-mutation fitness landscapes for an enzyme on multiple substrates reveal specificity is globally encoded. Nat. Commun. 8, 15695 (2017).
Chan, Y. H., Venev, S. V., Zeldovich, K. B. & Matthews, C. R. Correlation of fitness landscapes from three orthologous TIM barrels originates from sequence and structure constraints. Nat. Commun. 8, 14614 (2017).
Kelsic, E. D. et al. RNA structural determinants of optimal codons revealed by MAGE-Seq. Cell Syst. 3, 563–571 (2016).
Brenan, L. et al. Phenotypic characterization of a comprehensive set of MAPK1/ERK2 missense mutants. Cell Rep. 17, 1171–1183 (2016).
Bandaru, P. et al. Deconstruction of the Ras switching cycle through saturation mutagenesis. eLife 6, e27810 (2017).
Findlay, G. M. et al. Accurate functional classification of thousands of BRCA1 variants with saturation genome editing. bioRxiv Preprint at https://www.biorxiv.org/content/early/2018/04/05/294520 (2018).
Matreyek, K. A. et al. Multiplex assessment of protein variant abundance by massively parallel sequencing. bioRxiv Preprint at https://www.biorxiv.org/content/early/2018/01/16/211011 (2018).
Klesmith, J. R., Bacik, J.-P., Michalczyk, R. & Whitehead, T. A. Comprehensive sequence-flux mapping of a levoglucosan utilization pathway in E. coli. ACS Synth. Biol. 4, 1235–1243 (2015).
Haddox, H. K., Dingens, A. S., Hilton, S. K., Overbaugh, J. & Bloom, J. D. Mapping mutational effects along the evolutionary landscape of HIV envelope. eLife 7, e34420 (2018).
Pokusaeva, V. et al. Experimental assay of a fitness landscape on a macroevolutionary scale. bioRxiv Preprint at https://www.biorxiv.org/content/early/2018/04/06/222778 (2018).
Weile, J. et al. A framework for exhaustively mapping functional missense variants. Mol. Syst. Biol. 13, 957 (2017).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Suzek, B. E., Wang, Y., Huang, H., McGarvey, P. B. & Wu, C. H. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
Ekeberg, M., Lövkvist, C., Lan, Y., Weigt, M. & Aurell, E. Improved contact prediction in proteins: using pseudolikelihoods to infer Potts models. Phys. Rev. E 87, 012707 (2013).
Tipping, M. E. & Bishop, C. M. Probabilistic principal component analysis. J. R. Stat. Soc. Series B Stat. Methodol. 61, 611–622 (1999).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. arXiv Preprint at https://arxiv.org/abs/1412.6980 (2014).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982).
Acknowledgements
We thank C. Sander, F. Poelwijk, D. Duvenaud, S. Sinai, E. Kelsic, the Cold Spring Harbor Laboratory Sequence-Function Relationship Journal Club and members of the Marks lab for helpful comments and discussions. A.J.R. is supported by DOE CSGF fellowship DE-FG02-97ER25308. D.S.M. and J.B.I. were funded by NIGMS (R01GM106303).
Author information
Authors and Affiliations
Contributions
A.J.R., J.B.I., and D.S.M. designed the study. A.J.R. and J.B.I. performed the computations. All authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1
Distribution of experimental mutation effects and predictions made by DeepSequence.
Supplementary Figure 2 Mutation-effect predictions from generative models can be generalized to unseen sequences.
(above) Spearman ρ of mutation effect prediction of β-lactamase7 of each of the three generative models (N = 4788). Sequences with a normalized hamming distance greater than 0.53, 0.6, 0.8, and 0.95 with respect to the reference sequence are removed from the alignment before model fitting and inference. The distribution of hamming distances of the alignment and the cutoff of inclusion into each alignment is shown below.
Supplementary Figure 3 Predictions from all generative models for sequence families exhibited biases when compared to experimental data.
By transforming all model predictions and mutations to normalized ranks, we can compare effect predictions to experimental data across all biological datasets and models. The site-independent, pairwise, and latent variable models systematically over and under predict the effects of mutations according to amino acid identity. These biases vary in magnitude and direction depending on the amino acid identity before mutation (wildtype) or the residue identity it is mutated to (mutant).
Supplementary Figure 4 Supervised calibration of mutation-effect predictions improves predictive performance.
Amino acid bias was corrected with linear regression for all generative models, leaving one protein out for test and training a model on the rest (Methods). The bottom of the bar is Spearman ρ before correction, while the top is Spearman ρ after correction. Predictions without any evolutionary information (Supervised) performed considerably worse than other predictors.
Supplementary Figure 5 Differential improvement was strongest for deleterious effects.
Top five positions with largest reduction in rank error from independent model to DeepSequence for eight proteins are shown on the crystal structure of the protein.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5
Supplementary Table 1
Identification of sequences and datasets analyzed
Supplementary Table 2
Experimental and computed mutation effects
Supplementary Table 3
Correlation of DeepSequence and other evolutionary models to mutation effects
Supplementary Table 4
Statistical comparison to other mutation-effect prediction algorithms
Supplementary Table 5
Biologically motivated priors and Bayesian learning improve model performance
Supplementary Table 6
Dictionary parameters from all protein models
Supplementary Table 7
Group sparsity prior log enrichment statistics
Supplementary Table 8
PDB files used for scale parameter analysis
Supplementary Table 9
Residual analysis of effect predictions
Rights and permissions
About this article
Cite this article
Riesselman, A.J., Ingraham, J.B. & Marks, D.S. Deep generative models of genetic variation capture the effects of mutations. Nat Methods 15, 816–822 (2018). https://doi.org/10.1038/s41592-018-0138-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0138-4
This article is cited by
-
Prop3D: A flexible, Python-based platform for machine learning with protein structural properties and biophysical data
BMC Bioinformatics (2024)
-
Sparks of function by de novo protein design
Nature Biotechnology (2024)
-
Deep generative design of RNA family sequences
Nature Methods (2024)
-
Protein design using structure-based residue preferences
Nature Communications (2024)
-
Variational autoencoder for design of synthetic viral vector serotypes
Nature Machine Intelligence (2024)