Abstract
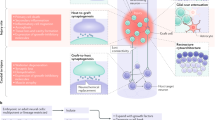
Spinal cord neural stem cells (NSCs) have great potential to reconstitute damaged spinal neural circuitry, but they have yet to be generated in vitro. We now report the derivation of spinal cord NSCs from human pluripotent stem cells (hPSCs). Our observations show that these spinal cord NSCs differentiate into a diverse population of spinal cord neurons occupying multiple positions along the dorso-ventral axis, and can be maintained for prolonged time periods. Grafts into injured spinal cords were rich with excitatory neurons, extended large numbers of axons over long distances, innervated their target structures, and enabled robust corticospinal regeneration. The grafts synaptically integrated into multiple host intraspinal and supraspinal systems, including the corticospinal projection, and improved functional outcomes after injury. hPSC-derived spinal cord NSCs could enable a broad range of biomedical applications for in vitro disease modeling and constitute an improved clinically translatable cell source for ‘replacement’ strategies in several spinal cord disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kadoya, K. et al. Spinal cord reconstitution with homologous neural grafts enables robust corticospinal regeneration. Nat. Med. 22, 479–487 (2016).
Ma, L. et al. Human embryonic stem cell-derived GABA neurons correct locomotion deficits in quinolinic acid-lesioned mice. Cell Stem Cell 10, 455–464 (2012).
Goldman, S. A. Stem and progenitor cell-based therapy of the central nervous system: hopes, hype, and wishful thinking. Cell Stem Cell 18, 174–188 (2016).
Lindvall, O. & Kokaia, Z. Stem cells in human neurodegenerative disorders—time for clinical translation? J. Clin. Invest. 120, 29–40 (2010).
Lemon, R. N. Descending pathways in motor control. Annu. Rev. Neurosci. 31, 195–218 (2008).
Tuszynski, M. H. & Steward, O. Concepts and methods for the study of axonal regeneration in the CNS. Neuron 74, 777–791 (2012).
Lu, P. et al. Long-distance growth and connectivity of neural stem cells after severe spinal cord injury. Cell 150, 1264–1273 (2012).
Gouti, M. et al. In vitro generation of neuromesodermal progenitors reveals distinct roles for Wnt signalling in the specification of spinal cord and paraxial mesoderm identity. PLoS Biol. 12, e1001937 (2014).
Henrique, D., Abranches, E., Verrier, L. & Storey, K. G. Neuromesodermal progenitors and the making of the spinal cord. Development 142, 2864–2875 (2015).
Gouti, M., Metzis, V. & Briscoe, J. The route to spinal cord cell types: a tale of signals and switches. Trends Genet. 31, 282–289 (2015).
Mazzoni, E. O. et al. Saltatory remodeling of Hox chromatin in response to rostrocaudal patterning signals. Nat. Neurosci. 16, 1191–1198 (2013).
Lippmann, E. S. et al. Deterministic HOX patterning in human pluripotent stem cell-derived neuroectoderm. Stem Cell Rep. 4, 632–644 (2015).
Liu, J. P., Laufer, E. & Jessell, T. M. Assigning the positional identity of spinal motor neurons: rostrocaudal patterning of Hox-c expression by FGFs, Gdf11, and retinoids. Neuron 32, 997–1012 (2001).
Philippidou, P. & Dasen, J. S. Hox genes: choreographers in neural development, architects of circuit organization. Neuron 80, 12–34 (2013).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Alaynick, W. A., Jessell, T. M. & Pfaff, S. L. SnapShot: spinal cord development. Cell 146, 178 (2011).
Callaway, E. M. & Luo, L. Monosynaptic circuit tracing with glycoprotein-deleted rabies viruses. J. Neurosci. 35, 8979–8985 (2015).
Grskovic, M., Javaherian, A., Strulovici, B. & Daley, G. Q. Induced pluripotent stem cells—opportunities for disease modelling and drug discovery. Nat. Rev. Drug Discov. 10, 915–929 (2011).
Avior, Y., Sagi, I. & Benvenisty, N. Pluripotent stem cells in disease modelling and drug discovery. Nat. Rev. Mol. Cell Biol. 17, 170–182 (2016).
Sterneckert, J. L., Reinhardt, P. & Schöler, H. R. Investigating human disease using stem cell models. Nat. Rev. Genet. 15, 625–639 (2014).
Termsarasab, P., Thammongkolchai, T. & Frucht, S. J. Spinal-generated movement disorders: a clinical review. J. Clin. Mov. Disord. 2, 18 (2015).
Tao, Y. & Zhang, S. C. Neural subtype specification from human pluripotent stem cells. Cell Stem Cell 19, 573–586 (2016).
Zhang, X. et al. Pax6 is a human neuroectoderm cell fate determinant. Cell Stem Cell 7, 90–100 (2010).
Deschamps, J. & van Nes, J. Developmental regulation of the Hox genes during axial morphogenesis in the mouse. Development 132, 2931–2942 (2005).
Kim, W. Y. et al. GSK-3 is a master regulator of neural progenitor homeostasis. Nat. Neurosci. 12, 1390–1397 (2009).
Fuccillo, M., Joyner, A. L. & Fishell, G. Morphogen to mitogen: the multiple roles of Hedgehog signalling in vertebrate neural development. Nat. Rev. Neurosci. 7, 772–783 (2006).
Dunn, N. R., Vincent, S. D., Oxburgh, L., Robertson, E. J. & Bikoff, E. K. Combinatorial activities of Smad2 and Smad3 regulate mesoderm formation and patterning in the mouse embryo. Development 131, 1717–1728 (2004).
Takemoto, T. et al. Tbx6-dependent Sox2 regulation determines neural or mesodermal fate in axial stem cells. Nature 470, 394–398 (2011).
Gómez-Skarmeta, J. L., Campuzano, S. & Modolell, J. Half a century of neural prepatterning: the story of a few bristles and many genes. Nat. Rev. Neurosci. 4, 587–598 (2003).
Davis-Dusenbery, B. N., Williams, L. A., Klim, J. R. & Eggan, K. How to make spinal motor neurons. Development 141, 491–501 (2014).
Kelly, T. K., Karsten, S. L., Geschwind, D. H. & Kornblum, H. I. Cell lineage and regional identity of cultured spinal cord neural stem cells and comparison to brain-derived neural stem cells. PLoS One 4, e4213 (2009).
Goulding, M. Circuits controlling vertebrate locomotion: moving in a new direction. Nat. Rev. Neurosci. 10, 507–518 (2009).
Soshnikova, N. & Duboule, D. Epigenetic temporal control of mouse Hox genes in vivo. Science 324, 1320–1323 (2009).
Ni, Y. et al. Characterization of long descending premotor propriospinal neurons in the spinal cord. J. Neurosci. 34, 9404–9417 (2014).
Lu, P. et al. Prolonged human neural stem cell maturation supports recovery in injured rodent CNS. J. Clin. Invest. 127, 3287–3299 (2017).
Grealish, S. et al. Monosynaptic tracing using modified rabies virus reveals early and extensive circuit integration of human embryonic stem cell-derived neurons. Stem Cell Rep. 4, 975–983 (2015).
Adler, A. F., Lee-Kubli, C., Kumamaru, H., Kadoya, K. & Tuszynski, M. H. Comprehensive monosynaptic rabies virus mapping of host connectivity with neural progenitor grafts after spinal cord Injury. Stem Cell Rep. 8, 1525–1533 (2017).
Del Barrio, M. G. et al. A transcription factor code defines nine sensory interneuron subtypes in the mechanosensory area of the spinal cord. PLoS One 8, e77928 (2013).
Flynn, J. R., Graham, B. A., Galea, M. P. & Callister, R. J. The role of propriospinal interneurons in recovery from spinal cord injury. Neuropharmacology 60, 809–822 (2011).
Esposito, M. S., Capelli, P. & Arber, S. Brainstem nucleus MdV mediates skilled forelimb motor tasks. Nature 508, 351–356 (2014).
Kirkeby, A. et al. Generation of regionally specified neural progenitors and functional neurons from human embryonic stem cells under defined conditions. Cell Rep. 1, 703–714 (2012).
Basso, D. M., Beattie, M. S. & Bresnahan, J. C. A sensitive and reliable locomotor rating scale for open field testing in rats. J. Neurotrauma 12, 1–21 (1995).
Kloos, A. D., Fisher, L. C., Detloff, M. R., Hassenzahl, D. L. & Basso, D. M. Stepwise motor and all-or-none sensory recovery is associated with nonlinear sparing after incremental spinal cord injury in rats. Exp. Neurol. 191, 251–265 (2005).
Rosenzweig, E. S. et al. Restorative effects of human neural stem cell grafts on the primate spinal cord. Nat. Med. 24, 484–490 (2018).
Krtolica, A. et al. GROα regulates human embryonic stem cell self-renewal or adoption of a neuronal fate. Differentiation 81, 222–232 (2011).
Elkabetz, Y. et al. Human ES cell-derived neural rosettes reveal a functionally distinct early neural stem cell stage. Genes Dev. 22, 152–165 (2008).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Acknowledgements
We thank P. Lu, J. Dulin, C. Lee-Kubli, and E. Staufenberg for experimental assistance; R. Kawaguchi and F. Gao for analysis of RNA-seq data; R.C. Addis (University of Pennsylvania, Philadelphia, PA, USA) for providing lentivirus expressing GCaMP5; S. Fisher (UCSF, San Francisco, CA, USA) for providing UCSF4 hESCs; T. Müller and C. Birchmeier (Max Delbrück Center for Molecular Medicine, Berlin, Germany) for providing TLX3 and LBX1 antibodies; S. Ross (University of Pittsburgh, Pittsburgh, PA, USA) for providing BHLHB5 antibody; and R. Darnell (the Rockefeller University, New York, NY, USA) for providing Hu antibody. This work was supported by the Veterans Administration Gordon Mansfield Spinal Cord Injury Consortium (to M.H.T.), the NIH (grants NS042291 and EB014986 to M.H.T.), the Craig H. Neilsen Foundation (to H.K. and K.K.), the Japan Society for the Promotion of Science (to H.K.), the Bernard and Anne Spitzer Charitable Trust (to M.H.T.), and the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation (to M.H.T.).
Author information
Authors and Affiliations
Contributions
H.K. designed and carried out experiments, interpreted results, and wrote the manuscript. K.K. contributed to the conception of the project and interpretation of results. A.F.A. contributed to rabies tracing experiments. Y.T. performed electrophysiological analysis. L.G. contributed to the behavior analysis. G.C. contributed to RNA-seq analysis. M.H.T. contributed to the conception of the project, data review, and interpretation of results, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
H.K. and M.H.T are submitting a patent application related to this work.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Induction of spinal cord NSCs from H9 ESCs.
(a,b) Immunolabeling for SOX2, CDX2, and PAX6 (a; without CHIR + FGF2/8, b; with CHIR + FGF2/8), ten days after neural induction. Scale bars, 100 um. LSBD; LDN + SB + DAPT. (c,d) qPCR for (c) forebrain markers FOXG1, OTX2, IRX3, and SIX3 and hindbrain markers EGR2 (KROX20) and MAFB, and (d) mesodermal markers TBX6, FOXC1, and MEOX1, endodermal marker SOX17, and neural crest cell markers FOXD3, SNAI1 (SNAIL), and SOX10, ten days after neural induction (n = 3 for each gene, three independent experiments). (e,f) Immunolabeling for SOX2, FOXG1, and Nestin (Nes) (e; without CHIR + FGF2/8, f; with CHIR + FGF2/8), ten days after neural induction, indicating CHIR and FGF2/8 treated cells do not express FOXG1 and do not form neural rosettes. Scale bars, 100 μm. (g-j) Immunolabeling for Brachyury (Bra) and DAPI (g; LDN + SB + DAPT (LSBD), h; LSBD + FGF2/8, i; LSBD + CHIR, j; LSBD + CHIR + FGF2/8) four days after neural induction (P0). Scale bars, 100 um. (k-n) Immunolabeling for brain markers OTX2, PAX6, and DAPI, eight days after neural induction (k; LSBD, l; LSBD + FGF2/8, m; LSBD + CHIR, n; LSBD + CHIR + FGF2/8). Scale bars, 100 um. (o-r) Immunolabeling for CDX2, PAX6, and SOX2 (o; LSBD, p; LSBD + FGF2/8, q; LSBD + CHIR, r; LSBD + CHIR + FGF2/8), eight days after neural induction. Scale bars, 100 um. (s) HOX C gene expression nine days after neural induction (n = 3 for each gene, three independent experiments). Spinal cord-specific HOXC 6-10 expression was dramatically up-regulated in the LSBD + CHIR + FGF2/8 condition. Each gene expression was normalized to the highest gene expressing sample (which was set at a value of 1.0). Data are presented as mean ± s.e.m. Immunolabeling was independently repeated at least three times with similar results.
Supplementary Figure 2 Purity of H9-cell-derived spinal cord NSCs.
(a-h) Immunolabeling for the neural stem cell marker SOX2, mesodermal stem cell marker TBX6, and DAPI (a,e; CHIR + FGF2/8, b,f; CHIR + FGF2/8 + DAPT, c,g; CHIR + FGF2/8 + LDN + SB (LSB), d,h; CHIR + FGF2/8 + LSB + DAPT), three days (P0, a-d) and four days (P1, e-h) after neural induction. Scale bars, 100 um. (i-l) Immunolabeling for CDX2, SOX2, and DAPI (i; CHIR + FGF2/8, j; CHIR + FGF2/8 + DAPT, k; CHIR + FGF2/8 + LSB, l; CHIR + FGF2/8 + LSB + DAPT), eight days after neural induction. Scale bars, 100 um. (m-o) Quantification in e-h (m, n = 4 for each group, four independent experiments) and i-l (n,o, n = 4 for each group, four independent experiments). Two-way ANOVA ((m) F(3,12) = 19.6, P < 0.0001. (n) F(3,12) = 5.5, P = 0.0129. (o) F(3,12) = 6.2, P = 0.0089), followed by Tukey’s multiple comparisons. *P < 0.05, **P < 0.01, ***P < 0.001. (p-s) Immunolabeling for the definitive endodermal marker CXCR4 and DAPI, four days after neural induction (P0, p; CHIR + FGF2/8, q; CHIR + FGF2/8 + DAPT, r; CHIR + FGF2/8 + LSB, s; CHIR + FGF2/8 + LSB + DAPT). Scale bars, 100 um. LSB blocked mesodermal and endodermal induction15. Data are presented as mean ± s.e.m. Immunolabeling was independently repeated four times (Suppl. Figure 2a-l) and twice (Suppl. Figure 2p-s) with similar results.
Supplementary Figure 3 Rostro-caudal and dorso-ventral axis of H9-cell-derived spinal cord NSCs.
(a) Immunolabeling for the post-mitotic neuronal marker DCX, neural intermediate filament Nestin (Nes), and mitotic NSC marker SOX2 at P18. Scale bar, 100 um. (b,c) Temporal analysis of proportion of Nestin-expressing cells (b) and SOX2- or DCX- expressing cells (c). Almost all cells expressed Nestin, DCX, and/or SOX2 at each passage. Data were normalized to the number of DAPI + cells (n = 3, three independent experiments). (d-f) Immunolabeling for dorsal (PAX6) and ventral (NKX6.1) markers (d), p3 marker NKX2.2 (e), or pMN marker OLIG2 (f) with NSC marker SOX2, 20 days after neural induction. Insets show higher magnifications. Scale bars, 100 μm. (g,h) Immunolabeling for NKX6.1, PAX6, and SOX2, 18 days after neural induction. Insets show the boxed regions. Hh-Ag1.5 (SHH) disrupted dorso-ventral polarity of induced cells. Scale bars, 100 μm. (i) Temporal analysis of dorso-ventral axis of H9-derived spinal cord NSCs. The number of PAX6- or NKX6.1- expressing cells were normalized to the number of SOX2-expressing cells (n = 3, three independent experiments). (j-l) qPCR for HOXA (j), HOXB (k), and HOXC (l) genes. Gene expression was normalized to the time point of maximum expression in each gene (n = 3 for each gene, three independent experiments). HOX genes were sequentially activated after neural induction12. Data are presented as mean ± s.e.m. Immunolabeling was independently repeated at least three times (Suppl. Figure 3a, d-f) and twice (Suppl. Figure 3g,h) with similar results.
Supplementary Figure 4 Differentiation of H9-cell-derived spinal cord NSCs.
(a) Representative traces of spontaneous excitatory postsynaptic currents (sEPSCs; left panels) and corresponding action potential traces elicited by depolarizing current injections (right panels) from the same neuron. Both 250 nM and 1 uM tetrodotoxin (TTX; sodium channel blocker) completely blocked sEPSCs and action potentials. (b) Representative traces of sEPSCs (superimposed 10 sweeps) in the presence or absence of the glutamate receptor antagonists, DNQX (AMPA antagonist) and APV (NMDAR antagonist), and the GABA-A receptor antagonist picrotoxin (PTX). The sEPSCs were blocked by DNQX/APV and PTX further blocked spontaneous activities, suggesting that spontaneous activities observed were result from trans-synaptic transmissions onto the neuron recorded. (c,d) Ten weeks after neuronal differentiation, cells differentiated into NeuN, Hu, or TUJ1-expressing neurons (c) and GFAP or S100β -expressing astrocytes (d). Scale bars, 100 μm. (e-g) Immunolabeling for neuronal markers (MAP2, DCX, or TUJ1) and neurotransmitter phenotype markers, eight weeks after neuronal differentiation, showing that these cells have the potential to differentiate into glutamatergic (glutamate (Glu); e), GABAergic (GABA; f), and glycinergic (GlyT2; g) neurons. Insets show higher magnifications. Scale bars, 50 μm. (h-p) Immunolabeling for neuronal marker DCX and transcription factors that specify neuronal subtypes, including HB9 (h), ISL1/2 (i), LIM1 + 2 (j), BRN3A (k), LBX1 (l), TLX3 (m), CHX10 (n), LHX3 (o), and PAX2 (p)16 two weeks after neuronal differentiation. Scale bars, 40 μm. (q-z) Clonal analysis of H9-derived spinal cord NSCs. Single GFP-expressing H9-derived spinal cord NSC was expanded on GFP negative H9-derived spinal cord NSCs and then purified by a flow cytometry. These subclones differentiated into GFAP-expressing astrocytes (q) and DCX (r) or TUJ1 (s) -expressing neurons. These neurons included HB9 (t) or ISL1/2 (u) -expressing motor neurons and LIM1 + 2-expressing spinal interneurons (v). These interneurons are further characterized and included CHX10 (w) or LHX3 (x) -expressing excitatory interneurons and PAX2 (y) or FOXP2 (z) -expressing inhibitory interneurons. Scale bars, 40 um (q, t-z) and 60 um (r,s). Immunolabeling was independently repeated at least twice with similar results.
Supplementary Figure 5 UCSF4 ESC-derived spinal cord NSCs.
(a) mRNA expression of HOXC and HOXD clusters in each sample by RNA-Sequencing. Expression levels were normalized to the cell sample with the highest level ( = 100). (b) Transcriptional activities of HOXD cluster genes in default H9-NSCs, H9-derived spinal cord NSCs, and fetal spinal cord-derived NPCs, clearly showing that H9-derived spinal cord NSCs express spinal cord-specific HOXD genes. (c-e) Immunolabeling of UCSF4-derived spinal cord NSCs for CDX2, OCT4, and SOX2 (c), CDX2, PAX6, and SOX2 (d), and SOX2, FOXG1, and Nestin (Nes, e), ten days after neural induction. Scale bars, 250 μm (c) and 100 um (d,e). (f,g) qPCR for pluripotent cell markers NANOG and OCT4, neural markers SOX1 and SOX2, brain markers FOXG1 and OTX2, and neural crest marker FOXD3 (f), and CDX2 and HOX genes (g) showing collinear HOX gene activation (n = 3 for each gene, three independent experiments). Gene expression levels were normalized to expression in UCSF4 ESCs for each gene. (h) Gene expression of fetal central nervous system (CNS) tissues and UCSF4-derived spinal cord NSCs at passage 13 (n = 3 for each gene, three independent experiments). Each gene expression was normalized to the sample with the highest expression level ( = 1). (i) Immunolabeling for neural marker SOX2, SOX1, and Nestin (Nes) at passage 12, indicating that long-term cultured UCSF4-derived cells maintain their neural identity. Scale bar, 100 μm. (j) Immunolabeling for the neuronal marker MAP2 and astrocyte marker GFAP, five weeks after differentiation. Scale bar, 100 μm. (k) Transcriptional activities of HOXA-D clusters in UCSF4-derived spinal cord NSCs (passage 8). Data are presented as mean ± s.e.m. Immunolabeling was independently repeated at least twice with similar results.
Supplementary Figure 6 Induction of spinal cord NSCs from human iPSCs.
(a-d) Immunolabeling of IPS11-derived spinal cord NSCs for Brachyury (Bra, a), PAX6 (b), and SOX2 (c), three days after neural induction. Scale bars, 20 μm. (e,f) qPCR of IPS11-derived cells for pluripotent cell markers NANOG and OCT4, neural markers SOX1 and SOX2, mesodermal marker TBXT, and ectodermal marker PAX6 (e, n = 3 for each gene, three independent experiments), and spinal cord marker CDX2 and HOXC genes (f, n = 3 for each gene, three independent experiments). Gene expression levels were normalized to expression in IPS11 for each gene. (g) Triple immunolabeling for CDX2, PAX6, and SOX2 nine days after induction. Scale bar, 50 μm. (h) Triple immunolabeling for SOX2, SOX1, and Nestin (Nes) in passage 12 IPS11-derived spinal cord NSCs. Scale bar, 50 μm. (i,j) Triple immunolabeling for NKX6.1, PAX6, and SOX2 (i) and NKX2.2, OLIG2, and SOX2 (j), 18 days after neural induction. Scale bars, 100 μm (i) and 50 um (j). (k) Immunolabeling for the neuronal markers DCX and MAP in passage 15 IPS11-derived NSCs after ten days of neuronal differentiation, suggesting that they maintain their neuronal identity over several passages. Scale bar, 50 um. (l-s) Immunolabeling for DCX and transcription factors that specify neuronal subtypes, including HB9 (l), ISL1/2 (m), LIM1 + 2 (n), BRN3A (o), TLX3 (p), LBX1 (q), PAX2 (r), and LHX3 (s)16, two weeks after neuronal differentiation. Scale bars, 20 um. Data are presented as mean ± s.e.m. Immunolabeling was independently repeated at least twice with similar results.
Supplementary Figure 7 Neuronal subtypes generated from H9-cell-derived spinal cord NSCs.
(a) Confocal images at graft-host border (dotted line: left; graft, right; host), six weeks post-grafting reveals that grafted cells express the neuronal markers DCX and NeuN. Scale bar, 200 μm. (b) Neuronal and glial phenotype quantification. 80% of grafted cells express the neuronal marker Hu when assessed three months post-grafting, whereas 11% of grafted cells express the mature astrocyte marker GFAP, and 1% of cells express the oligodendrocyte precursor cell marker NG2 three months post-grafting (n = 4). (c-j) Confocal images of co-localization of the human-specific nuclear marker, HuNu with spinal cord neuronal subtype-specific transcription factors: sensory interneurons (LBX1 (c) and TLX3 (d)), inhibitory interneurons (PAX2 (e)), or pre-motor interneurons (BHLHB5 (f), FOXP1 (g), FOXP2 (h), LHX3 (i), and CHX10 (j)). Scale bars, 10 μm. (k-t) Confocal images from center of graft reveal co-localization of the neuronal markers DCX or NeuN with neuronal subtype-specific transcription factors of dorsal interneurons (BRN3A (k), LBX1 (l), and TLX3 (m)), inhibitory interneurons (PAX2 (n)), or intermediate-ventral interneurons (LIM 1 + 2 (o), BHLHB5 (p), FOXP1 (q), FOXP2 (r), LHX3 (s), and CHX10 (t)), three months post-grafting. Insets show triple labeling of GFP, neuronal markers, and subtype-specific transcription factors. Scale bars, 25 μm. Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary Figure 8 Synaptic connectivity of host and H9-cell-derived spinal cord NSCs.
(a) Confocal image of co-localization of the graft (GFP) and human-specific axonal TAU (hTAU) at C8 level. Scale bar, 20 um. (b,c) Triple labeling for GFP, human SYN (hSYN), and CaMKII (b) or ChAT (c), indicating co-association of graft-derived human axon terminals with a synaptic marker in direct association with host neurons. Scale bars, 5 μm. (d-f) Sagittal sections triple-labeled for GFP, rat VGLUT1/2 (rVGLUT1/2), and VGLUT1/2, three months post-grafting. Scale bars, 500 μm (d,f) and 100 um (e). High-magnification views of boxed area in d is shown in e. Inset shows higher magnification in boxed area in f. Scale bar, 5 μm. Rat VGLUT1/2 (rVGLUT1/2)-expressing rat excitatory pre-synaptic elements were present at (d,e) host-graft border (dotted line: left; host, right; graft) and (f) inside the graft. VGLUT1/2 is immunoreactive for both human and rat antigens; rVGLUT1/2 labels only rat antigen. (g) Close apposition of rat VGLUT1/2 (rVGLUT1/2) axons terminals in graft onto GFP-expressing cell bodies or dendrites, suggesting host-graft excitatory connection. Scale bar, 5 μm. Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary Figure 9 Growth of H9-cell-derived spinal cord NSC grafts.
(a-c) Immunolabeling for the human nuclear marker HuNu, proliferation marker Ki67, and DAPI, six months post-grafting. Dotted lines indicate graft-host border. Scale bars, 1 mm (a,b) and 50 um (c). Inset shows higher magnification of boxed area in c. Scale bar, 5 um. The graft occupies the lesion cavity without extending beyond it. Only 0.61 ± 0.29 % of HuNu-expressing human cells expressed Ki67 (n = 3 grafts sampled). (d-h) Immunolabeling for HuNu, the pluripotent marker OCT4, and DAPI, six months post-grafting. OCT4-expressing cells were not detected. f-h show higher magnification of boxed areas in d and e. Scale bars, 500 um (d,e) and 100 um (f-h). Data are presented as mean ± s.e.m. Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary Figure 10 Connectivity of host intraspinal neurons with H9-cell-derived spinal cord NSC grafts.
(a,b) Double immunostaining of mCherry and HuNu for control (w/o TVA + G-protein (GP)) and experimental (w/ TVA + GP) rats. No mCherry-expressing cells were founded in control animals. Inset shows mCherry +HuNu + cell in b. Scale bars, 500 μm. (c) Confocal image shows co-localization of rodent-specific alpha-internexin (rNF66) and mCherry. Scale bar, 10 um. (d) Sagittal section showing retrogradely, trans-synaptically traced host mCherry-expressing cells in the cervical spinal cord rostral to the C4 dorsal column SCI site. Connected host neurons are scattered throughout gray matter. Dotted lines indicate rostral host-graft border. Scale bar, 1 mm; inset shows the boxed region. Scale bar, 200 um. (e) Trans-synaptically labeled cells at the L2 level mainly located in lamina III-IV. Inner lamina II is indicated by PKCγ staining. Scale bar, 250 μm. (f-h) Characterization of mCherry + host cells in the cervical spinal cord. Synaptically connected mCherry + host neurons include sensory interneurons labeled for TLX3 (f), LBX1 (g), and BRN3A (h). Scale bars, 20 μm. (i,j) Sensory interneurons at the L2 host spinal cord level are synaptically connected with the spinal cord NSC graft, indicated by double labeling for mCherry and sensory interneuronal marker c-Maf. Dotted line indicates boundary between host dorsal column (DC) and gray matter. Scale bars, 100 μm (i) and 20 μm (j). Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary Figure 11 Connectivity of host supraspinal neurons with H9-cell-derived spinal cord NSC grafts.
(a) Neurons of the red nucleus (RN) in the pons were connected to the graft, indicated by mCherry labeling. Inset is high magnification of arrowhead. CP; cerebral peduncle. SN; substantia nigra. Scale bar, 500 μm. (b) In the reticular formation, host neurons in the gigantocellular reticular nucleus ventral part (GiV) are synaptically connected to the human graft. IRt; intermediate reticular nucleus, PCRt; parvocellular reticular nucleus, PY; pyramidal tract, Sp5i; spinal trigeminal nucleus, interpolar part, 4 V; fourth ventricle. Scale bar, 500 μm. (c-e) Host raphespinal axons (labeled for 5-HT) regenerate into GFP+ H9-derived spinal cord NSC grafts in the lesion site (c); the boxed region is shown in d. These regenerating axons express the presynaptic marker synaptophysin (SYN) in close association with graft cell somata (e). Dotted line indicates rostral host-graft border. Scale bars, 50 μm (c), 10 μm (d), and 5 um (e). Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary Figure 12 Connectivity of host DRG neurons with H9-cell-derived spinal cord NSC grafts.
(a-c) Neurons in multiple dorsal root ganglia (DRG) in proximity to the graft at C4 are labeled for mCherry, indicating connectivity with the human NSC graft. (a) Cervical, (b) thoracic, and (c) lumbar DRGs. Scale bars, 1 mm. (d,e) High magnification images of mCherry-expressing cells in C7 (d) and L3 (e) DRGs. Scale bars, 500 um. (f) Double immunolabeling for mCherry and NF200 in C5 DRGs showing co-localization of large diameter DRG neurons with mCherry. Scale bar, 100 μm. A similar pattern was observed in all experimental animals (n = 4).
Supplementary Figure 13 Corticospinal regeneration into H9-cell-derived spinal cord NSC grafts.
To demonstrate that rat corticospinal axons clearly regenerate into the human neural stem cell graft within the lesion site, we performed double labeling for rat Thy1 (rThy1, blue) and host corticospinal axons (CST, red). (a) Rat Thy1 is expressed in host gray matter and is absent from the lesion site containing the human neural stem cell graft. (b) In the same section, corticospinal axons regenerate into the lesion site containing the human neural stem cell graft, which lacks Thy1 labeling. (c) Panels a and b are shown in a single image. Boxed area in c is shown in Fig. 5j. Dotted lines indicate host-graft border. Scale bars, 500 um (a-c). (d-f) Another example of host corticospinal axons regenerating in a human neural stem cell graft occupying the lesion site; the graft does not label for rThy1. Boxed area in d is shown in e and f. Dotted lines indicate host-graft border. Scale bars, 250 um (d) and 100 um (e,f). (g,h) The graft in the lesion site is labeled for the human-specific nuclear marker HuNu. Host corticospinal axons (CST) regenerate into the graft. Boxed area (the lesion center) is shown in h. Scale bars, 500 um (g) and 100 um (h). Corticospinal tracing experiments were independently repeated twice with similar results.
Supplementary Figure 14 Corticospinal regeneration into H9-cell-derived spinal cord NSCs.
(a) Double labeling for corticospinal axons (CST) and rat-specific VGLUT1 (rVGLUT1) within spinal cord NSCs. Scale bar, 20 μm. (b) Triple labeling for GFP, corticospinal axons (CST), and synaptophysin (SYN) reveals co-localization of regenerating corticospinal axon terminals with SYN, suggesting synaptic connectivity. Scale bars, 5 μm. (c-f) Quantification of graft-derived neurons (Hu+HuNu+ cells) among H9-brain (c, n = 4), brainstem (d, n = 5), and spinal cord NSCs (e, n = 6). Hu labels all neurons (human or rodent; images are from within graft which contains only human neurons), whereas HuNu exclusively labels the nuclei of human cells. There were no statistical differences among groups (One-way ANOVA, F(2, 12) = 0.28, P = 0.78), indicating that differences in the proportion of neurons among graft types did not account for the support of corticospinal regeneration. Scale bars, 50 μm. Data are presented as mean ± s.e.m. Data was obtained from independent two experiments with similar results.
Supplementary Figure 15 Survival, differentiation, and axonal extension of H9-cell-derived spinal cord NSCs grafted to thoracic contusions.
(a) GFP-labeled H9-spinal cord NSCs were grafted into sites of T10 spinal cord contusion. Horizontal section labeled for GFP and NeuN, showing graft fill of the lesion cavity assessed five months after grafting. Scale bar, 1 mm. Rostral is to left, and caudal is to right. (b) Ungrafted T10 contusion cavity. Scale bar, 1 mm. (c,d) Immunolabeling for doublecortin (DCX) and human nuclei (HuNu) around the center of grafts, indicating neuronal differentiation of grafted cells. Scale bars, 50 um (c) and 5 um (d). (e) Immunolabeling for the neuronal marker NeuN and GFP. GFP-expressing grafts differentiate into neurons expressing a mature neuronal marker. Scale bar, 50 um. Separate images of boxed area are shown in right panels. Scale bars, 10 um. (f,g) Confocal images show the human-specific nuclear marker HuNu co-localizing with the neuronal marker NeuN (f) and the V2a interneuronal marker CHX10 (g). Scale bars, 5 um. (h,i) Horizontal section labeled for GFP and NeuN showing very large numbers of GFP-labeled axons that extend caudally into the host. Scale bar, 1 mm. Dotted line indicates caudal graft-host border. Boxed region is shown in i. Scale bar, 100 um. (j) Confocal image shows co-localization of GFP and human specific axonal marker TAU (hTAU). Scale bar, 5 um. (k) High magnification image of GFP-expressing axons in the host gray matter caudal to the lesion. Scale bar, 20 um. (l) Confocal image shows GFP-labeled human axon terminal expresses human specific pre-synaptic marker synaptophysin (hSYN). Scale bar, 2 um. (m) Confocal image shows human synaptophysin (hSYN)-expressing human synaptic terminals in close apposition with the host CaMKII-expressing neurons caudal to the lesion, indicating graft-host connectivity. Scale bar, 10 um. Immunohistochemistry of the same sections was repeated at least twice with similar results.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Kumamaru, H., Kadoya, K., Adler, A.F. et al. Generation and post-injury integration of human spinal cord neural stem cells. Nat Methods 15, 723–731 (2018). https://doi.org/10.1038/s41592-018-0074-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0074-3
This article is cited by
-
Grafted human-induced pluripotent stem cells-derived oligodendrocyte progenitor cells combined with human umbilical vein endothelial cells contribute to functional recovery following spinal cord injury
Stem Cell Research & Therapy (2024)
-
BAF45D-binding to HOX genes was differentially targeted in H9-derived spinal cord neural stem cells
Scientific Reports (2024)
-
Intrinsic and extrinsic actions of human neural progenitors with SUFU inhibition promote tissue repair and functional recovery from severe spinal cord injury
npj Regenerative Medicine (2024)
-
The effect of nanomaterials on embryonic stem cell neural differentiation: a systematic review
European Journal of Medical Research (2023)
-
Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells
Cell Regeneration (2023)