Abstract
We describe Strelka2 (https://github.com/Illumina/strelka), an open-source small-variant-calling method for research and clinical germline and somatic sequencing applications. Strelka2 introduces a novel mixture-model-based estimation of insertion/deletion error parameters from each sample, an efficient tiered haplotype-modeling strategy, and a normal sample contamination model to improve liquid tumor analysis. For both germline and somatic calling, Strelka2 substantially outperformed the current leading tools in terms of both variant-calling accuracy and computing cost.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
McKenna, A. et al. Genome Res. 20, 1297–1303 (2010).
DePristo, M. A. et al. Nat. Genet. 43, 491–498 (2011).
Garrison, E. & Marth, G. arXiv Preprint available at https://arxiv.org/abs/1207.3907 (2012).
Cibulskis, K. et al. Nat. Biotechnol. 31, 213–219 (2013).
Rimmer, A. et al. Nat. Genet. 46, 912–918 (2014).
Narzisi, G. et al. Nat. Methods 11, 1033–1036 (2014).
Saunders, C. T. et al. Bioinformatics 28, 1811–1817 (2012).
Durbin, R., Eddy, S. R., Krogh, A. & Mitchison, G. Biological Sequence Analysis: Probabilistic Models of Proteins and Nucleic Acids (Cambridge Univ. Press, Cambridge, UK, 1998).
Ding, J. et al. Bioinformatics 28, 167–175 (2012).
Van der Auwera, G. A. et al. Curr. Protoc. Bioinforma. 43, 11.10.1–11.10.33 (2013).
Poplin, R. et al. bioRxiv Preprint at https://www.biorxiv.org/content/early/2016/12/14/092890 (2016).
Eberle, M. A. et al. Genome Res. 27, 157–164 (2017).
Altman, R. B. et al. Sci. Transl. Med. 8, 335ps10 (2016).
Poplin, R. et al. bioRxiv Preprint at https://www.biorxiv.org/content/early/2017/11/14/201178.1 (2017).
Zook, J. et al. bioRxiv Preprint at https://www.biorxiv.org/content/early/2018/05/25/281006 (2018).
Lai, Z. et al. Nucleic Acids Res. 44, e108 (2016).
Alioto, T. S. et al. Nat. Commun. 6, 10001 (2015).
Freed, D. N., Aldana, R., Weber, J. A. & Edwards, J. S. bioRxiv Preprint at https://www.biorxiv.org/content/early/2017/05/12/115717 (2017).
Li, H. arXiv Preprint at https://arxiv.org/abs/1303.3997 (2013).
Chen, K. et al. Genome Res. 24, 310–317 (2014).
Acknowledgements
We thank S. Kruglyak, B. Moore, J. O’Connell, and E. Kanterakis for helpful discussions and comments.
Author information
Authors and Affiliations
Contributions
S.K., K.S., A.L.H., M.A.B., E.N., M.K., X.C., Y.K., D.B., P.K., and C.T.S. designed the algorithms and implemented the Strelka2 software. S.K. and C.T.S. designed and performed the analyses. S.K., K.S., and C.T.S. wrote the manuscript, with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
S.K., K.S., A.L.H., M.A.B., E.N., X.C., Y.K., P.K., and C.T.S. are employees of Illumina, Inc., a public company that develops and markets systems for genetic analysis.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
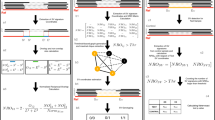
Supplementary Figure 1 Strelka2 variant-calling workflows.
Strelka2 supports detection of germline variants in small sample cohorts (up to ~10 individuals), and somatic variants from matched tumor-normal sample pairs. These two types of analyses share several high-level steps, including: (1) parameter estimation, (2) candidate variant discovery, (3) realignment and variant probability inference, and (4) empirical scoring and filtration. Here we diagram an overview of the major workflow components for both (a) germline and (b) somatic analyses.
Supplementary Figure 2 Structure of the germline indel error and variant-calling models in probabilistic graphical model plate notation (some details omitted).
(a) Indel error model. At each locus l, a preliminary estimate of the indel allele count vector C is modeled as a mixture binomial distribution governed by the two true haplotypes h1 and h2 (a function of the unobserved genotype hypothesis H), a set of indel error rates e (unobserved) and the total count X (observed). The error rates are selected from the full set of error parameters E according to the sequence context (summarized as an integer pair denoting the size s and number r of STR repeats; observed) and a binary state variable N (unobserved) categorizing the locus as clean (essentially zero error rates) or noisy (prone to indel errors). The genotype H and the noisy-clean state variable N are drawn from prior distributions that depend, respectively, on a context-specific mutation rate θ shared across samples and a context-specific noisy-state probability pn. (b) Variant calling model. The reads dj at every locus are modeled as depending on the corresponding base call quality strings qj, the unobserved haplotype hj that generated the read, and the locus-specific error rates e. The read-specific haplotype is drawn from the set of haplotypes in the locus-specific hypothesis H, of which the prior again depends on a parameter selected from θ according to context. The error rates are again selected from the global vector E of error parameters (now treated as fixed), with the difference that all loci analyzed by this model are assumed to be in the noisy state.
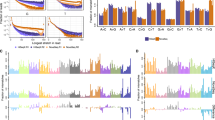
Supplementary Figure 3 Germline-indel-calling accuracy stratified by indel size and type.
The indel calling accuracy of various pipelines are plotted for the Consistency challenge Garvan dataset (left), Truth challenge HG002 dataset (center) and the GIAB HG005 dataset (right), for insertions (denoted by Ins) and deletions (denoted by Del) of different sizes (length 1-5: 88% of cases; length 6-15: 10% of cases; length 16 + : 2% of cases). For FreeBayes, the recall dropped substantially for long indels. For Strelka2 and GATK4, both of which employ local assembly, the recall drop was considerably smaller.
Supplementary Figure 4 Accuracy of germline indel and SNV calling for additional test datasets.
Results are shown for the Consistency challenge HLI dataset (left) and the Truth challenge HG001 dataset (right). Filled circles denote the pass threshold of each tool.
Supplementary Figure 5 Comparison of performance characteristics for Strelka2 versus Strelka.
(a) Comparison of somatic variant calling accuracy for the in-silico germline mixtures datasets described in Fig. 2a. Strelka2 has improved indel accuracy on impure tumor samples and is far more robust to contamination in the normal sample. (b) Comparison of runtime (wallclock time) and memory usage (peak resident set size) for the same datasets, measured on servers with two Intel Xeon E5-2680 v4 CPUs (total 28 physical cores) with 256 GB of memory.
Supplementary information
Supplementary Text and Figures
Supplementary Figs. 1–5 and Supplementary Notes 1–3
Supplementary Table 1
Germline variant calling accuracy
Supplementary Software 1
Strelka2 source code for version 2.9.0
Rights and permissions
About this article
Cite this article
Kim, S., Scheffler, K., Halpern, A.L. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods 15, 591–594 (2018). https://doi.org/10.1038/s41592-018-0051-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0051-x
This article is cited by
-
COSAP: Comparative Sequencing Analysis Platform
BMC Bioinformatics (2024)
-
A cautionary tale of low-pass sequencing and imputation with respect to haplotype accuracy
Genetics Selection Evolution (2024)
-
Race-specific coregulatory and transcriptomic profiles associated with DNA methylation and androgen receptor in prostate cancer
Genome Medicine (2024)
-
Missense mutation of NRAS is associated with malignant progression in neurocutaneous melanosis
Acta Neuropathologica Communications (2024)
-
Mutational landscape of inflammatory breast cancer
Journal of Translational Medicine (2024)