Abstract
Reconstruction of neural circuits from volume electron microscopy data requires the tracing of cells in their entirety, including all their neurites. Automated approaches have been developed for tracing, but their error rates are too high to generate reliable circuit diagrams without extensive human proofreading. We present flood-filling networks, a method for automated segmentation that, similar to most previous efforts, uses convolutional neural networks, but contains in addition a recurrent pathway that allows the iterative optimization and extension of individual neuronal processes. We used flood-filling networks to trace neurons in a dataset obtained by serial block-face electron microscopy of a zebra finch brain. Using our method, we achieved a mean error-free neurite path length of 1.1 mm, and we observed only four mergers in a test set with a path length of 97 mm. The performance of flood-filling networks was an order of magnitude better than that of previous approaches applied to this dataset, although with substantially increased computational costs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Macagno, E. R., Levinthal, C. & Sobel, I. Three-dimensional computer reconstruction of neurons and neuronal assemblies. Annu. Rev. Biophys. Bioeng. 8, 323–351 (1979).
Harris, K. M., Jensen, F. E. & Tsao, B. Three-dimensional structure of dendritic spines and synapses in rat hippocampus (CA1) at postnatal day 15 and adult ages: implications for the maturation of synaptic physiology and long-term potentiation. J. Neurosci. 12, 2685–2705 (1992).
Ventura, R. & Harris, K. M. Three-dimensional relationships between hippocampal synapses and astrocytes. J. Neurosci. 19, 6897–6906 (1999).
Fiala, J. C. Reconstruct: a free editor for serial section microscopy. J. Microsc. 218, 52–61 (2005).
Briggman, K. L. & Bock, D. D. Volume electron microscopy for neuronal circuit reconstruction. Curr. Opin. Neurobiol. 22, 154–161 (2012).
Jain, V., Seung, H. S. & Turaga, S. C. Machines that learn to segment images: a crucial technology for connectomics. Curr. Opin. Neurobiol. 20, 653–666 (2010).
Kim, J. S. et al. Space-time wiring specificity supports direction selectivity in the retina. Nature 509, 331–336 (2014).
Takemura, S.-Y. et al. Synaptic circuits and their variations within different columns in the visual system of Drosophila. Proc. Natl. Acad. Sci. USA 112, 13711–13716 (2015).
Helmstaedter, M., Briggman, K. L. & Denk, W. High-accuracy neurite reconstruction for high-throughput neuroanatomy. Nat. Neurosci. 14, 1081–1088 (2011).
Helmstaedter, M. et al. Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500, 168–174 (2013).
Cardona, A. TrakEM2: an ImageJ-based program for morphological data mining and 3D modeling. in Proc. ImageJ User and Developer Conference 18–19 (Centre de Recherche Public Henri Tudor: Luxembourg, 2006).
Berning, M., Boergens, K. M. & Helmstaedter, M. SegEM: efficient image analysis for high-resolution connectomics. Neuron 87, 1193–1206 (2015).
Plaza, S. M. Focused proofreading to reconstruct neural connectomes from EM images at scale. in Deep Learning and Data Labeling for Medical Applications (eds. Carneiro, G. et al.) 249–258 (Springer, Cham, 2016).
Jain, V. et al. Supervised learning of image restoration with convolutional networks. in Proc. IEEE 11th International Conference on Computer Vision 636–643 (IEEE, New York, 2007).
Ciresan, D., Giusti, A., Gambardella, L. M. & Schmidhuber, J. Deep neural networks segment neuronal membranes in electron microscopy images. in Advances in Neural Information Processing Systems 25 (eds. Pereira, F. et al.) 2852–2860 (Neural Information Processing Systems Foundation, La Jolla, CA, 2012).
Turaga, S. C. et al. Convolutional networks can learn to generate affinity graphs for image segmentation. Neural Comput. 22, 511–538 (2010).
Funke, J. et al. A deep structured learning approach towards automating connectome reconstruction from 3D electron micrographs. arXiv Preprint at https://arxiv.org/abs/1709.02974 (2017).
Lee, K., Zung, J., Li, P., Jain, V. & Seung, H. S. Superhuman accuracy on the SNEMI3D Connectomics Challenge. arXiv Preprint at https://arxiv.org/abs/1706.00120 (2017).
Andres, B., Koethe, U., Helmstaedter, M., Denk, W. & Hamprecht, F. A. Segmentation of SBFSEM volume data of neural tissue by hierarchical classification. in Pattern Recognition: Proceedings of the 30th DAGM Symposium (ed. Rigoll, G.)142–152 (Springer, Berlin, 2008).
Kaynig, V. et al. Large-scale automatic reconstruction of neuronal processes from electron microscopy images. Med. Image Anal. 22, 77–88 (2015).
Knowles-Barley, S. et al. RhoanaNet pipeline: dense automatic neural annotation. arXiv Preprint at https://arxiv.org/abs/1611.06973 (2016).
Beier, T. et al. Multicut brings automated neurite segmentation closer to human performance. Nat. Methods 14, 101–102 (2017).
Turaga, S. C., Briggman, K. L., Helmstaedter, M., Denk, W. & Seung, H. S. Maximin affinity learning of image segmentation. in Advances in Neural Information Processing Systems 22 (eds. Bengio, Y. et al.) 1865–1873 (Neural Information Processing Systems, La Jolla, CA, 2009).
Jain, V. et al. Boundary learning by optimization with topological constraints. in Proc. IEEE Computer Society Conference on Computer Vision and Pattern Recognition 2488–2495 (IEEE, New York, 2010).
Denk, W. & Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2, e329 (2004).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, Cambridge, MA, 2016).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional networks for biomedical image segmentation. in Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015 (eds. Navab, N. et al.) 234–241 (Springer, Cham, 2015).
Nunez-Iglesias, J., Kennedy, R., Plaza, S. M., Chakraborty, A. & Katz, W. T. Graph-based active learning of agglomeration (GALA): a Python library to segment 2D and 3D neuroimages. Front. Neuroinform. 8, 34 (2014).
Maitin-Shepard, J., Jain, V., Januszewski, M., Li, P. & Abbeel, P. Combinatorial energy learning for image segmentation. arXiv Preprint at https://arxiv.org/abs/1506.04304 (2015).
Saalfeld, S., Fetter, R., Cardona, A. & Tomancak, P. Elastic volume reconstruction from series of ultra-thin microscopy sections. Nat. Methods 9, 717–720 (2012).
Dorkenwald, S. et al. Automated synaptic connectivity inference for volume electron microscopy. Nat. Methods 14, 435–442 (2017).
Pallotto, M., Watkins, P. V., Fubara, B., Singer, J. H. & Briggman, K. L. Extracellular space preservation aids the connectomic analysis of neural circuits. eLife 4, e08206 (2015).
Zlateski, A. & Seung, H. S. Image segmentation by size-dependent single linkage clustering of a watershed basin graph. arXiv Preprint at https://arxiv.org/abs/1505.00249 (2015).
Kasthuri, N. et al. Saturated reconstruction of a volume of neocortex. Cell 162, 648–661 (2015).
Martin, D. R., Fowlkes, C. C. & Malik, J. Learning to detect natural image boundaries using local brightness, color, and texture cues. IEEE Trans. Pattern Anal. Mach. Intell. 26, 530–549 (2004).
Funke, J., Andres, B., Hamprecht, F. A., Cardona, A. & Cook, M. Efficient automatic 3D-reconstruction of branching neurons from EM data. in Proc. IEEE Conference on Computer Vision and Pattern Recognition 1004–1011 (IEEE, New York, 2012).
Wolf, S., Schott, L., Kothe, U. & Hamprecht, F. Learned watershed: end-to-end learning of seeded segmentation. in Proc. IEEE International Conference on Computer Vision (ICCV) 2030–2038 (IEEE, New York, 2017).
Bai, M. & Urtasun, R. Deep watershed transform for instance segmentation. in Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2858–2866 (IEEE, New York, 2017).
Meirovitch, Y. et al. A multi-pass approach to large-scale connectomics. arXiv Preprint at https://arxiv.org/abs/1612.02120 (2016).
Romera-Paredes, B. & Torr, P. H. S. Recurrent instance segmentation. in Computer Vision—ECCV 2016 (eds. Leibe, B. et al.) 312–329 (Springer, Cham, 2016).
Pinheiro, P. O , Lin, T.-Y., Collobert, R., & Dollár, P. Learning to refine object segments. in Computer Vision–ECCV 2016 (eds. Leibe, B. et al.) 75–91 (Springer, Cham, 2016).
Ren, M. & Zemel, R. S. End-to-end instance segmentation with recurrent attention. in Proc. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 293–301 (IEEE, New York, 2017).
Abadi, M. et al. TensorFlow: large-scale machine learning on heterogeneous distributed systems. arXiv Preprint at https://arxiv.org/abs/1603.04467 (2016).
LeCun, Y. et al. Backpropagation applied to handwritten zip code recognition. Neural Comput. 1, 541–551 (1989).
LeCun, Y. A , Bottou, L , Orr, G. B., & Müller, K.-R . Efficient BackProp. in Neural Networks: Tricks of the Trade (eds. Montavon, G., Orr, G. & Müller, K.-R.) 9–48 (Springer, Berlin, 2012).
Tschopp, F. Efficient convolutional neural networks for pixelwise classification on heterogeneous hardware systems. arXiv Preprint at https://arxiv.org/abs/1509.03371 (2015).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. arXiv Preprint at https://arxiv.org/abs/1603.05027 (2016).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. arXiv Preprint at https://arxiv.org/abs/1512.03385 (2015).
Kornfeld, J. et al. EM connectomics reveals axonal target variation in a sequence-generating network. eLife 6, e24364 (2017).
Acknowledgements
We thank T.L. Dean and B. Agüera y Arcas for useful discussions and support. We also thank M.S. Fee and J. Shlens for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
M.J. and V.J. conceived FFNs. M.J. developed pipelines and performed experiments. J.K. and W.D. acquired EM data and ground truth annotations. A.P. aligned the dataset. M.J. and J.K. analyzed results. V.J., M.J., W.D., P.H.L., J.M.-S., and J.K. developed skeleton metrics. M.J. and V.J. developed tissue-classification models. T.B., L.L., M.T., and J.M. developed software for annotation and visualization. V.J., M.J., W.D., and J.K. wrote the manuscript. V.J. supervised the project.
Corresponding author
Ethics declarations
Competing interests
J.K. holds shares of Ariadne service GmbH. M.J., P.H.L., A.P., T.B., L.L., J.M.-S., M.T., and V.J. are employees of Google LLC, which sells cloud computing services.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
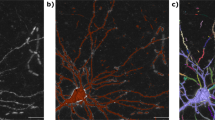
Supplementary Figure 1 Tissue classification results.
Manual annotations (left) and convolutional network inference (right) of a subset of the labeled voxel classes: blood vessel (red), myelin (blue), and cell body (green). False positive identifications of cell body voxels are visible in the automated inference (inside the myelinated area). Scale bar is 2 microns.
Supplementary Figure 2 Architecture of the FFN.
Overall computational architecture of the Flood-filling Network. Each of the eight convolutional modules are identical and implement the operations shown in the top inset box. The predicted object map (POM) is shown as the yellow mask channel, and provides recurrent feedback to the FFN.
Supplementary Information
Supplementary Text and Figures
Supplementary Figs. 1 and 2 and Supplementary Note
Supplementary Video 1
FFN reconstruction of a single neurite (i.e., seeded from a single voxel) in J0126 volume
Supplementary Software
Flood-filling network software for training and inference
Rights and permissions
About this article
Cite this article
Januszewski, M., Kornfeld, J., Li, P.H. et al. High-precision automated reconstruction of neurons with flood-filling networks. Nat Methods 15, 605–610 (2018). https://doi.org/10.1038/s41592-018-0049-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0049-4
This article is cited by
-
RoboEM: automated 3D flight tracing for synaptic-resolution connectomics
Nature Methods (2024)
-
Building an automated three-dimensional flight agent for neural network reconstruction
Nature Methods (2024)
-
Using high-resolution microscopy data to generate realistic structures for electromagnetic FDTD simulations from complex biological models
Nature Protocols (2024)
-
Attention based morphological guided deep learning network for neuron segmentation in electron microscopy
The Journal of Supercomputing (2024)
-
Seg2Link: an efficient and versatile solution for semi-automatic cell segmentation in 3D image stacks
Scientific Reports (2023)