Abstract
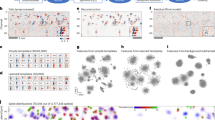
Thus far, optical recording of neuronal activity in freely behaving animals has been limited to a thin axial range. We present a head-mounted miniaturized light-field microscope (MiniLFM) capable of capturing neuronal network activity within a volume of 700 × 600 × 360 µm3 at 16 Hz in the hippocampus of freely moving mice. We demonstrate that neurons separated by as little as ~15 µm and at depths up to 360 µm can be discriminated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
21 May 2018
In the version of this Brief Communication originally published online, ref. 21 included details for a conference paper (Pegard, N. C. et al. Paper presented at Novel Techniques in Microscopy: Optics in the Life Sciences, Vancouver, BC, Canada, 12–15 April 2015). The correct reference is the following: Pégard, N. C. et al. Optica 3, 517–524 (2016). This error has been corrected in the print, HTML and PDF versions of the paper.
References
Yang, W. & Yuste, R. Nat. Methods 14, 349–359 (2017).
Prevedel, R. et al. Nat. Methods 13, 1021–1028 (2016).
Nöbauer, T. et al. Nat. Methods 14, 811–818 (2017).
Duemani Reddy, G., Kelleher, K., Fink, R. & Saggau, P. Nat. Neurosci. 11, 713–720 (2008).
Katona, G. et al. Nat. Methods 9, 201–208 (2012).
Botcherby, E. J. et al. Proc. Natl Acad. Sci. USA 109, 2919–2924 (2012).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Nat. Methods 10, 413–420 (2013).
Bouchard, M. B. et al. Nat. Photonics 9, 113–119 (2015).
Yang, W. et al. Neuron 89, 269–284 (2016).
Lu, R. et al. Nat. Neurosci. 20, 620–628 (2017).
Song, A. et al. Nat. Methods 14, 420–426 (2017).
Helmchen, F., Fee, M. S., Tank, D. W. & Denk, W. Neuron 31, 903–912 (2001).
Zong, W. et al. Nat. Methods 14, 713–719 (2017).
Flusberg, B. A. et al. Nat. Methods 5, 935–938 (2008).
Ghosh, K. K. et al. Nat. Methods 8, 871–878 (2011).
Ziv, Y. et al. Nat. Neurosci. 16, 264–266 (2013).
Cai, D. J. et al. Nature 534, 115–118 (2016).
Barbera, G. et al. Neuron 92, 202–213 (2016).
Levoy, M., Ng, R., Adams, A., Footer, M. & Horowitz, M. ACM Trans. Graph. 25, 924 (2006).
Prevedel, R. et al. Nat. Methods 11, 727–730 (2014).
Pégard, N. C. et al. Optica 3, 517–524 (2016).
Broxton, M. et al. Opt. Express 21, 25418–25439 (2013).
Pnevmatikakis, E. A. et al. Neuron 89, 285–299 (2016).
Acknowledgements
We thank M. Colombini (IMP Vienna) and J.M. Petrillo (Rockefeller University) for manufacturing of mechanical components and advice on mechanical design issues. We are grateful to M. Chen and A. Pernía-Andrade for advice on surgeries and viral injections. T.N. acknowledges support from the Leon Levy Foundation (Leon Levy Fellowship in Neuroscience). This work was supported in part by the Human Frontiers Science Program Project RGP0041/2012 (to A.V.), the Institute of Molecular Pathology (to M.I.M. and A.V.), the Kavli Foundation (to A.V.), the Smith Family Foundation (to D.D.C.), the Harvard Mind Brain Behavior Interfaculty Initiative (to D.D.C.), the Intelligence Advanced Research Projects Activity (IARPA; Department of Interior/Interior Business Center (DoI/IBC) contract number D16PC00002 to A.V. and D.D.C.) and the National Science Foundation (grant no. DBI-1707408 to A.V. and P.G.).
Author information
Authors and Affiliations
Contributions
O.S. contributed to design and implementation of signal extraction and motion detection, data analysis and writing of the manuscript. T.N. contributed to design and implementation of imaging and signal extraction, performed experiments, analyzed data and wrote the manuscript. L.W. implemented an early version of the imaging system, performed surgeries and experiments, analyzed data and contributed to writing of the manuscript. F.M.T. performed injections, surgeries, imaging and behavioral experiments and contributed to writing of the manuscript. C.N.X. contributed to injections and surgeries. M.I.M. performed simulations. A.G. and M.Y. developed nucleus-confined GCaMP under the guidance of D.D.C. D.A. developed the original Miniscope and helped with implementation of the MiniLFM under the supervision of P.G. A.V. conceived and led the project, conceptualized imaging and signal extraction, designed in vivo experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Alignment of MLA to sensor in MiniLFM.
(a) Alignment jig. For aligning the MLA to the sensor chip, a pair of custom 4-finger holders (silver cylindrical slotted parts, center left) was designed that can be tightened using hose clamps (not shown). One clamp holds the MLA (not visible, occluded by clamp) and is mounted statically on a post/post holder (leftmost part). The other clamp holds the sensor (turquoise rectangle) and is itself held by a 6-axis kinematic mount (Thorlabs K6XS) for adjusting tip, tilt and rotation, and lateral position. The kinematic mount is attached to a 3-axis linear stage assembly (Thorlabs PTA3A/M) for adjusting MLA-to-sensor distance as well as for convenient coarse adjustment of lateral position. (b) Zooms into sensor images of spots formed by passing collimated light through the microlens array (MLA), before (left) and after (right) optimizing microlens-to-sensor distance (focusing). Brightness is shown oversaturated to emphasize dim pixels. Spot spacing is approx. 20 px. Insert: Zoom into one of the focused spots, not oversaturated. (c) Mean focal spot area (blue trace) and mean focal spot peak intensity, evaluated over all focal spots on the sensor, versus MLA-sensor distance. Vertical axis is normalized to maximal values, horizontal axis is relative to optimal focus distance. At the optimal focus distance, the relative standard deviation of the spot size was as low as 12%, indicating highly homogeneous alignment.
Supplementary Figure 2 Comparison of agility of animals wearing no device, Miniscope, or MiniLFM.
Quantification of animal agility from recordings of behavior on a linear track, after completion of training (see Online Methods). Three mice; one trial under each condition per animal and day, for three consecutive days, resulting in a total n=27 trials. Trial duration: 10 minutes. Inter-trial break: 1 hour. Wide horizontal bars indicated mean, error bars are s.e.m. Data point color indicates animal. (a) Average walking speed. ns, not significant by one-way ANOVA (F = 1.776, p = 0.089, 8 DOF). (b) Distance travelled per trial. ns, not significant by one-way ANOVA (F = 2.112, p = 0.465, 8 DOF). (c) Number of stops made during trial. ns, not significant; *, significant at p < 0.05 by one-way ANOVA (F = 4.11, p = 0.011, 8 DOF).
Supplementary Figure 3 Sketch of experimental setup used for simultaneous 2PM + MiniLFM–SID recordings.
See Methods for description of setup and procedure.
Supplementary Figure 4 Characterization of cross-talk on the level of individual Ca2+ transients as a function of neuronal distance.
Putative crosstalk event strength between pairs of neuronal traces on the level of individual Ca2+ transients versus neuron pair distance. Color scale gives column-wise, top-to-bottom cumulative density of events (normalized to one for each column). Solid magenta lines are contour lines of the resulting set of cumulative distribution functions for the 0.1, 0.25 and 0.5 quantiles, as indicated by the black labels. The majority of events are concentrated in the lowest row (low event strength, i.e. predominantly false positives due to noise). Identification and quantification of the transient pairs that resulted in crosstalked events was performed by first using the CaImAn package for inferring the maximum likelihood firing rate from both the ground truth (2PM) and the SID-extracted MiniLFM traces (recorded simultaneously from a 2P-excited plane as described in the Online Methods). Continuously connected regions of nonzero firing rate were detected in the resulting time series and classified as a single event. For each matched pair of SID-extracted and ground-truth activity traces, those events were identified that appeared in the latter but not in the former. The origin of the resulting false positive events in the SID traces was determined by finding matching events in ground-truth traces. The putative crosstalk events were weighted by the relative amplitude of the correctly assigned and crosstalked peaks, respectively, and binned according to distance between the source and crosstalk target neuron. The resulting distributions of events (shown in this figure by the cumulative density) were tested for equality using the two-sample Kolmogorov-Smirnov test, resulting in a significant difference only between the lowest distance bin (5–15 µm) compared to the others (p = 0.041). This can be interpreted as the signature of crosstalk events that have not been successfully demixed and assigned by the SID algorithm and hence establishes a criterion for the neuron discriminability threshold of SID under the given conditions (see main text). Note that no signature of crosstalk was detected in the excess mutual information analysis presented in Fig. 2h of the main text. Only when cutting the data to the level of individual calcium transients, as is done here, and thereby artificially boosting the sensitivity, a minimal but significant increase in the value of the crosstalk metric could be detected for neuronal pairs separated by less than ~15 µm. Data from n = 5 recordings from two mice. Total number of data points (neuron pairs): 726.
Supplementary Figure 5 MiniLFM–SID volumetric Ca2+ imaging up to 260 µm deep in hippocampus CA1 of a freely moving mouse expressing nucleus-confined Ca2+ indicator.
(a) Neuron positions obtained by SID-analysis of an 8-minute MiniLFM recording at 16 Hz frame rate in mouse hippocampus expressing nucleus-localized indicator AAV9.Syn.H2B.GCaMP6f.WPRE.Pzac2.1 (top and side views). (b) Heatmap of motion-corrected and de-noised temporal signals corresponding to the 127 neuron positions shown in (a). (c) Stacked neuronal activity traces, same data as in (b). (d) Histogram of the neuron positions shown in (a) versus depth. (e) Histogram of nearest-neighbor distances between the neurons shown in (a).
Supplementary Figure 6 Comparison of the motion detected by the motion-detection metric derived from MiniLFM raw data, and the transverse acceleration as measured by an accelerometer.
Top trace: Acceleration metric derived from a miniature accelerometer attached to the MiniLFM sensor circuit board (Online Methods) during a 5-minute trial of free behavior on a linear track. Raw signal was processed by hi-pass filtering, re-binning and taking the modulus of the transverse acceleration component (i.e., in the sensor plane). Bottom trace: Motion metric derived from MiniLFM raw data of the same trial. Metric is based on range of difference frame values (Online Methods).
Supplementary Figure 7 Motion correction of MiniLFM data.
(a) Lateral motion-induced image shift versus time during a 10-minute, regular (non-LFM) Miniscope recording from hippocampal CA1 of a mouse moving back and forth on a linear track. One pixel corresponds to ~1 µm in the sample. Quasi-periodic peaks in traces occur when animal initiates track traversal after reward administration at track ends. Right panels: Histograms of shifts in the lateral-medial (L-M) and anterior-posterior (A-P) directions. (b) Illustration of motion correction pipeline (see Supplementary Note 3 and Online Methods): A motion metric (magenta trace) is calculated on the raw data and thresholded at a high value (black horizontal line) to detect strong motion bursts. The time series of raw data frames (illustrated as a film strip) is split into low-motion segments (black rectangles) that lie in between the motion bursts. The low-motion segments are SID-processed individually, which yields a set of neuron footprints (blue images) and corresponding activity traces (green) for each segment. The neurons from each segment are then pooled, merging strongly overlapping footprints and extracting the corresponding activity traces across the full dataset. To further reduce motion artefacts and interpolate motion-affected frames, the same motion metric is now thresholded at a lower value, and the time steps above threshold are masked out (grey areas) from the activity traces. For each neuron, a model is trained on the motion-masked activity traces that estimates the underlying firing rate (black trace) and GECI response time constants. The model is then used to interpolate the masked-out time steps, yielding a motion-corrected estimate of the physiological neural activity (blue trace). (c) Examples of motion-corrected activity traces (see also Supplementary Fig. 8). Top: Motion metric calculated from the raw data (magenta trace), with black horizontal line indicating the threshold above which frames are considered motion-affected. Bottom: Three examples of motion-corrected Ca2+ activity signals (blue) together with maximum-likelihood-estimate of underlying firing rate (black), each corresponding to a line of the heatmap shown in Fig. 2c.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Notes 1–3
Supplementary Software
Custom code for MiniLFM alignment and data analysis pipeline. Updated versions of this code will be made available at https://github.com/vazirilab
Supplementary Table 1
Source data for SID-extracted neuron centroids and activity time series
Supplementary Video 1
Mouse with head-mounted MiniLFM moving freely in a square arena Video of adult mouse behaving and moving spontaneously for 50 s in a square arena. MiniLFM is screw-clamped into a baseplate that had been glued to skull, centered on an implanted GRIN objective lens (not visible; see Online Methods and Fig. 1). Data cable (white) is suspended from an arm above the center of the arena
Supplementary Video 2
Mouse with head-mounted MiniLFM moving freely on a linear track Video of adult mouse behaving and moving spontaneously for 43 s on a linear track. MiniLFM is screw-clamped into a baseplate that had been glued to skull, centered on an implanted GRIN objective lens (not visible; see Online Methods and Fig. 1). Data cable (white) is suspended from a ring that is gliding along a steel suspension wire above the track
Supplementary Video 3
Perspective rendering of neuron positions and Ca2+ activity during free movement, recorded with a MiniLFM from mouse hippocampus CA1 and extracted with the SID algorithm Recording frame rate: 16 Hz. Real-time recording duration: 30 min. Playback speed: 10.7×. White dashed lines indicate field-of-view of MiniLFM with dimensions 800 × 800 × 400 µm. Neurons are rendered as spheres placed at their SID-extracted locations. Color indicates neuron brightness
Supplementary Video 4
Wide-field imaging of Ca2+ activity with a head-mounted, unmodified (i.e., non-LFM) Miniscope in mouse hippocampal CA1 during free movement in an arena Unprocessed widefield recording of Ca2+ activity during free behavior in arena, using an unmodified, head-mounted Miniscope system. Frame rate: 16 Hz. Playback speed: 4×. Field-of-view is cropped by approx. 75% to approx. 370 × 240 µm for clarity
Supplementary Video 5
3D rendering of conventionally (non-SID) frame-by-frame-reconstructed MiniLFM data. Conventional frame-by-frame, deconvolution-based LFM reconstruction of an excerpt (duration: 80 s) of the same recording that was also analysed using SID and is shown in Supplementary Video 3 and Fig. 2a-c. Recording frame rate: 16 Hz. Playback includes every third recorded frame, played at a third of the recording frame rate, resulting in a playback duration equal to the recording duration. Field-of-view: approx. 620 × 500 × 320 µm
Supplementary Video 6
MiniLFM raw data Unprocessed MiniLFM sensor data as a multipage TIFF file, excerpt (recording duration 30 s = 480 frames at 16 Hz frame rate) from the same recording shown in Supplementary Video 2, Supplementary Video 4, and Fig. 2a-c
Rights and permissions
About this article
Cite this article
Skocek, O., Nöbauer, T., Weilguny, L. et al. High-speed volumetric imaging of neuronal activity in freely moving rodents. Nat Methods 15, 429–432 (2018). https://doi.org/10.1038/s41592-018-0008-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-018-0008-0
This article is cited by
-
Mesoscopic calcium imaging in a head-unrestrained male non-human primate using a lensless microscope
Nature Communications (2024)
-
Correlated-photon imaging at 10 volumetric images per second
Scientific Reports (2023)
-
Imagining the future of optical microscopy: everything, everywhere, all at once
Communications Biology (2023)
-
Large depth-of-field ultra-compact microscope by progressive optimization and deep learning
Nature Communications (2023)
-
Large-scale recording of neuronal activity in freely-moving mice at cellular resolution
Nature Communications (2023)