Abstract
Erythropoietin (Epo) is the master regulator of erythropoiesis and oxygen homeostasis. Despite its physiological importance, the molecular and genomic contexts of the cells responsible for renal Epo production remain unclear, limiting more-effective therapies for anemia. Here, we performed single-cell RNA and transposase-accessible chromatin (ATAC) sequencing of an Epo reporter mouse to molecularly identify Epo-producing cells under hypoxic conditions. Our data indicate that a distinct population of kidney stroma, which we term Norn cells, is the major source of endocrine Epo production in mice. We use these datasets to identify the markers, signaling pathways and transcriptional circuits characteristic of Norn cells. Using single-cell RNA sequencing and RNA in situ hybridization in human kidney tissues, we further provide evidence that this cell population is conserved in humans. These preliminary findings open new avenues to functionally dissect EPO gene regulation in health and disease and may serve as groundwork to improve erythropoiesis-stimulating therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Mouse single-cell RNA-seq and single-cell ATAC-seq data, and human single-cell RNA-seq data that support the findings of this study, were deposited in the Gene Expression Omnibus (GEO) under accession code GSE193321. Previously published scRNA-seq data that were re-analyzed here are available under the accession codes GSE114530 (ref. 47), GSE129798 (ref. 44), EGAS00001002325GSE155794 (ref. 46) and GSE107585 (ref. 43), as well as from the European Genome-phenome Archive (EGA) under study IDs EGAS00001002325 (ref. 42). We used mouse genome mm10 and human genome Hg38 as reference genomes. ATAC-seq tracks are accessible on UCSC browser https://genome.ucsc.edu/s/bjoert/Mouse%20hypoxia%20kidney%20bulk%20scATAC%2Dseq%20from%20single%20nuclei.
Code availability
Metacell source code can be found at https://github.com/tanaylab/metacell. Source code to identify Norn cells in human single cell atlas can be found at https://github.com/AmitLab/Kidney-Norn-cells-identification. Source code used for transcription factor motif analysis can be found at https://github.com/vanheeringen-lab/gimmemotifs.
References
Orkin, S. H. Diversification of haematopoietic stem cells to specific lineages. Nat. Rev. Genet. 1, 57–64 (2000).
Jacobson, L. O., Goldwasser, E., Fried, W. & Plzak, L. Role of the kidney in erythropoiesis. Nature 179, 633–634 (1957).
Wenger, R. H. & Kurtz, A. Erythropoietin. Compr. Physiol. 1, 1759–1794 (2011).
Lacombe, C. et al. Peritubular cells are the site of erythropoietin synthesis in the murine hypoxic kidney. J. Clin. Invest. 81, 620–623 (1988).
Wu, H., Liu, X., Jaenisch, R. & Lodish, H. F. Generation of committed erythroid BFU-E and CFU-E progenitors does not require erythropoietin or the erythropoietin receptor. Cell 83, 59–67 (1995).
Haemoglobin Concentrations for the Diagnosis of Anaemia and Assessment of Severity (World Health Organization, 2011); https://apps.who.int/iris/bitstream/handle/10665/85839/WHO_NMH_NHD_MNM_11.1_eng.pdf
Prchal, J. T. Polycythemia vera and other primary polycythemias. Curr. Opin. Hematol. 12, 112–116 (2005).
Haase, V. H. Hypoxia-inducible factor–prolyl hydroxylase inhibitors in the treatment of anemia of chronic kidney disease. Kidney Int. Suppl. 11, 8–25 (2021).
Gangaraju, R. et al. Upregulation of thrombo-inflammatory pathways may contribute to increased thrombotic risk in polycythemia vera and essential thrombocythemia. Blood 128, 3143 (2016).
Gordeuk, V. R. et al. Thrombotic risk in congenital erythrocytosis due to up-regulated hypoxia sensing is not associated with elevated hematocrit. Haematologica 105, e87–e90 (2020).
Rankin, E. B. et al. Hypoxia-inducible factor-2 (HIF-2) regulates hepatic erythropoietin in vivo. J. Clin. Invest. 117, 1068–1077 (2007).
Kapitsinou, P. P. et al. Hepatic HIF-2 regulates erythropoietic responses to hypoxia in renal anemia. Blood 116, 3039–3048 (2010).
Paliege, A. et al. Hypoxia-inducible factor-2alpha-expressing interstitial fibroblasts are the only renal cells that express erythropoietin under hypoxia-inducible factor stabilization. Kidney Int. 77, 312–318 (2010).
Semenza, G. L., Nejfelt, M. K., Chi, S. M. & Antonarakis, S. E. Hypoxia-inducible nuclear factors bind to an enhancer element located 3′ to the human erythropoietin gene. Proc. Natl Acad. Sci. USA 88, 5680–5684 (1991).
Appelhoff, R. J. et al. Differential function of the prolyl hydroxylases PHD1, PHD2, and PHD3 in the regulation of hypoxia-inducible factor. J. Biol. Chem. 279, 38458–38465 (2004).
Simonson, T. S. et al. Genetic evidence for high-altitude adaptation in Tibet. Science 329, 72–75 (2010).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014).
Witt, K. E. & Huerta-Sánchez, E. Convergent evolution in human and domesticate adaptation to high-altitude environments. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180235 (2019).
Moore, L. G., Young, D., McCullough, R. E., Droma, T. & Zamudio, S. Tibetan protection from intrauterine growth restriction (IUGR) and reproductive loss at high altitude. Am. J. Hum. Biol. 13, 635–644 (2001).
Beall, C. M. et al. Natural selection on EPAS1 (HIF2α) associated with low hemoglobin concentration in Tibetan highlanders. Proc. Natl Acad. Sci. USA 107, 11459–11464 (2010).
Lorenzo, F. R. et al. A genetic mechanism for Tibetan high-altitude adaptation. Nat. Genet. 46, 951–956 (2014).
Wiesener, M. S. et al. Widespread hypoxia-inducible expression of HIF-2alpha in distinct cell populations of different organs. FASEB J. 17, 271–273 (2003).
Imeri, F. et al. Generation of renal Epo-producing cell lines by conditional gene tagging reveals rapid HIF-2 driven Epo kinetics, cell autonomous feedback regulation, and a telocyte phenotype. Kidney Int. 95, 375–387 (2019).
Kobayashi, H. et al. Distinct subpopulations of FOXD1 stroma-derived cells regulate renal erythropoietin. J. Clin. Invest. 126, 1926–1938 (2016).
Obara, N. et al. Repression via the GATA box is essential for tissue-specific erythropoietin gene expression. Blood 111, 5223–5232 (2008).
Chang, Y.-T. et al. DNA methyltransferase inhibition restores erythropoietin production in fibrotic murine kidneys. J. Clin. Invest. 126, 721–731 (2016).
Loya, F., Yang, Y., Lin, H., Goldwasser, E. & Albitar, M. Transgenic mice carrying the erythropoietin gene promoter linked to lacZ express the reporter in proximal convoluted tubule cells after hypoxia. Blood 84, 1831–1836 (1994).
Bussolati, B. et al. Renal CD133(+)/CD73(+) progenitors produce erythropoietin under hypoxia and prolyl hydroxylase inhibition. J. Am. Soc. Nephrol. 24, 1234–1241 (2013).
Gerl, K. et al. Erythropoietin production by PDGFR-β(+) cells. Pflug. Arch. 468, 1479–1487 (2016).
Broeker, K. A. E. et al. Different subpopulations of kidney interstitial cells produce erythropoietin and factors supporting tissue oxygenation in response to hypoxia in vivo. Kidney Int. 98, 918–931 (2020).
Marchetti, P., Bugliani, M., De Tata, V., Suleiman, M. & Marselli, L. Pancreatic beta cell identity in humans and the role of type 2 diabetes. Front. Cell Dev. Biol. 5, 55 (2017).
Laha, D., Grant, R., Mishra, P. & Nilubol, N. The role of tumor necrosis factor in manipulating the immunological response of tumor microenvironment. Front. Immunol. 12, 656908 (2021).
Dahl, S.L. et al. Fate‐mapping of erythropoietin‐producing cells in mouse models of hypoxaemia and renal tissue remodelling reveals repeated recruitment and persistent functionality. Acta Physiol. (Oxf.) 234, e13768 (2022).
Dahl, S. L., Bapst, A. M., Khodo, S. N., Scholz, C. C. & Wenger, R. H. Fount, fate, features, and function of renal erythropoietin-producing cells. Pflug. Arch. 474, 783–797 (2022).
Shanks, J. H., Hill, C. M., Lappin, T. R. & Maxwell, A. P. Localization of erythropoietin gene expression in proximal renal tubular cells detected by digoxigenin-labelled oligonucleotide probes. J. Pathol. 179, 283–287 (1996).
Yamazaki, S. et al. A mouse model of adult-onset anaemia due to erythropoietin deficiency. Nat. Commun. 4, 1950 (2013).
Franke, K. et al. HIF-1α is a protective factor in conditional PHD2-deficient mice suffering from severe HIF-2α-induced excessive erythropoiesis. Blood 121, 1436–1445 (2013).
Kobayashi, H., Davidoff, O., Pujari-Palmer, S., Drevin, M. & Haase, V. H. EPO synthesis induced by HIF-PHD inhibition is dependent on myofibroblast transdifferentiation and colocalizes with non-injured nephron segments in murine kidney fibrosis. Acta Physiol. 235, e13826 (2022).
Bek-Pedersen, K. Norns in Old Norse Mythology (Dunedin Academic Press, 2013).
Suresh, S., Rajvanshi, P. K. & Noguchi, C. T. The many facets of erythropoietin physiologic and metabolic response. Front. Physiol. 10, 1534 (2019).
Young, M. D. et al. Single-cell transcriptomes from human kidneys reveal the cellular identity of renal tumors. Science 361, 594–599 (2018).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Ransick, A. et al. Single-cell profiling reveals sex, lineage, and regional diversity in the mouse kidney. Dev. Cell 51, 399–413 (2019).
Combes, A. N. et al. Single cell analysis of the developing mouse kidney provides deeper insight into marker gene expression and ligand-receptor crosstalk. Development 146, dev178673 (2019).
England, A. R. et al. Identification and characterization of cellular heterogeneity within the developing renal interstitium. Development 147, dev190108 (2020).
Hochane, M. et al. Single-cell transcriptomics reveals gene expression dynamics of human fetal kidney development. PLoS Biol. 17, e3000152 (2019).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Keren-Shaul, H. et al. MARS-seq2.0: an experimental and analytical pipeline for indexed sorting combined with single-cell RNA sequencing. Nat. Protoc. 14, 1841–1862 (2019).
Baran, Y. et al. MetaCell: analysis of single-cell RNA-seq data using K-nn graph partitions. Genome Biol. 20, 206 (2019).
Nishiyama, A. & Kim-Mitsuyama, S. New approaches to blockade of the renin–angiotensin–aldosterone system: overview of regulation of the renin–angiotensin–aldosterone system. J. Pharmacol. Sci. 113, 289–291 (2010).
Pan, X. et al. Isolation and characterization of renal erythropoietin-producing cells from genetically produced anemia mice. PLoS ONE 6, e25839 (2011).
Gimm, T. et al. Hypoxia-inducible protein 2 is a novel lipid droplet protein and a specific target gene of hypoxia-inducible factor-1. FASEB J. 24, 4443–4458 (2010).
Mondéjar-Parreño, G. et al. Uncovered contribution of Kv7 channels to pulmonary vascular tone in pulmonary arterial hypertension. Hypertension 76, 1134–1146 (2020).
Schödel, J. et al. High-resolution genome-wide mapping of HIF-binding sites by ChIP-seq. Blood 117, e207–e217 (2011).
Orlando, I. M. C. et al. Distal and proximal hypoxia response elements cooperate to regulate organ-specific erythropoietin gene expression. Haematologica 105, 2774–2784 (2020).
Semenza, G. L., Koury, S. T., Nejfelt, M. K., Gearhart, J. D. & Antonarakis, S. E. Cell-type-specific and hypoxia-inducible expression of the human erythropoietin gene in transgenic mice. Proc. Natl Acad. Sci. USA 88, 8725–8729 (1991).
Hirano, I. et al. Renal anemia model mouse established by transgenic rescue with an erythropoietin gene lacking kidney-specific regulatory elements. Mol. Cell. Biol. 37, e00451-16 (2017).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Bruse, N. & van Heeringen, S. J. GimmeMotifs: an analysis framework for transcription factor motif analysis. Preprint at bioRxiv https://doi.org/10.1101/474403 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Bornstein, C. et al. Single-cell mapping of the thymic stroma identifies IL-25-producing tuft epithelial cells. Nature 559, 622–626 (2018).
Plasschaert, L. W. et al. A single-cell atlas of the airway epithelium reveals the CFTR-rich pulmonary ionocyte. Nature 560, 377–381 (2018).
Quaggin, S. E. et al. The basic-helix-loop-helix protein pod1 is critically important for kidney and lung organogenesis. Development 126, 5771–5783 (1999).
Ide, S. et al. Transcription factor 21 is required for branching morphogenesis and regulates the Gdnf-axis in kidney development. J. Am. Soc. Nephrol. 29, 2795–2808 (2018).
Helman, A. & Melton, D. A. A stem cell approach to cure type 1 diabetes. Cold Spring Harb. Perspect. Biol. 13, a035741 (2021).
Bapst, A. M., Dahl, S. L., Knöpfel, T. & Wenger, R. H. Cre-mediated, loxP independent sequential recombination of a tripartite transcriptional stop cassette allows for partial read-through transcription. Biochim. Biophys. Acta Gene Regul. Mech. 1863, 194568 (2020).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Geirsdottir, L. et al. Cross-species single-cell analysis reveals divergence of the primate microglia program. Cell 179, 1609–1622 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Acknowledgements
We thank members of the Amit laboratory for critical discussions. We thank Y. Kuperman, L. Adler and S. Viukov for assisting with hypoxia experiments. We thank K. Pozyuchenko, G.H. Siloni, C. Padberg and T. Bajanowski for assisting with human kidney samples, and P. Spielmann and T. Knöpfel for technical support. We thank T.-M. Salame, E. Hagai and E. Kopitman for assisting with flow cytometry. This study was funded by European Union European Research Council Advanced (grant no. 101055341-TROJAN-Cell, I.A.); Deutsche Forschungsgemeinschaft (Project-ID 259373024 – TRR 167, I.A.), Israel Science Foundation (ISF; grant no. 1944/22, I.A.), the ISF Israel Precision Medicine Program (607/20, I.A.), Human Frontier Science Program (long-term postdoctoral fellow LT 000230/2019, B.K.K.), European Molecular Biology Organization (postdoctoral fellowship, ALTF 112-2022, A.G.), Villum Fonden (Young Investigator award project no. 00025300, F.R.) and by the Swiss National Science Foundation (grant no. 310030_184813 R.H.W.).
Author information
Authors and Affiliations
Contributions
B.K.K. and I.A. conceived the project and designed the experiments. B.K.K. performed the experiments. A.G. and C.G. contributed to experimental and project design. L.G. contributed to experimental and project design and assisted with FACS experiments. A.M.B., Y.K., S.L.D., M.Z., O.B. and S.S.-L., assisted with experiments. A.G.-S., S.D., V.Y., J.F., S.W., P.P.M. and B.R. assisted with human kidney specimens. S.H. and C.G. contributed to the human experimental part. A.G., E.D. and K.X. analyzed the data. R.A. and S.-Y.W. contributed to the analysis. A.M.B., S.S.-L. and O.M.L. performed mRNA-FISH experiments and image analysis. B.L. and A.G. assisted with image analysis. F.R., E.W., J.T.P., M.S. and T.S.K. assisted with analysis. H.K.-S., F.S., T.S.P. and M.K. assisted the 10x multiome experiment. B.K.K., A.G. and I.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Katalin Susztak, Tammie Bishop and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Anna Maria Ranzoni, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 QC of single cell data.
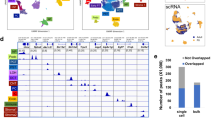
a, b, QC for mouse renal MARS-seq showing number of reads and genes per plate, respectively. Box plots presents the third quartile (top of the box), median (centre lines) and first quartile (bottom of the box) of measurements. The whiskers represent 1.5 times the interquartile range from the top or bottom of the box. From n = 22 independent experiments, kidneys from n = 52 mice (33 males, 19 females), cells were sorted into 384 well plates. Nu,ber of QC cells per plate are as follows: AB9040 = 103, AB9041 = 122, AB9042 = 103, AB9043 = 107, AB9044 = 109, AB9045 = 124, AB9046 = 180, AB9047 = 177, AB9202 = 132, AB9203 = 12, AB9204 = 101, AB9205 = 28, AB9206 = 11, AB9207 = 16, AB9208 = 19, AB9209 = 182, AB9210 = 180, AB9211 = 203, AB9212 = 168, AB9213 = 211, AB9214 = 253, AB9215 = 249, AB9216 = 196, AB9217 = 198, AB9218 = 214, AB9219 = 182, AB9220 = 197, AB9221 = 174, AB9316 = 118, AB9317 = 116, AB9453 = 365, AB9454 = 298, AB9455 = 361, AB9456 = 361, AB9464 = 176, AB10289 = 12, AB10290 = 21, AB10291 = 91, AB10292 = 34, AB10293 = 28, AB10344 = 279, AB10345 = 299, AB10346 = 263, AB10347 = 259, AB10448 = 127, AB10449 = 35, AB10450 = 103, AB10451 = 327, AB10452 = 321, AB10453 = 342, AB10454 = 365, AB10455 = 361, AB10460 = 64, AB10461 = 291, AB10765 = 339, AB10766 = 318, AB10767 = 328, AB10768 = 277, AB10769 = 319, AB10770 = 297, AB10771 = 329, AB10772 = 301, AB10926 = 369, AB10927 = 362, AB10928 = 361, AB10929 = 357, AB11060 = 360, AB11061 = 355, AB11062 = 365, AB11063 = 364, AB11064 = 352, AB11065 = 330, AB11066 = 360, AB11306 = 356, AB11307 = 344, AB11308 = 335, AB11309 = 329, AB11310 = 320, AB11311 = 324, AB11312 = 353, AB11313 = 333, AB11409 = 16, AB11410 = 11, AB11411 = 26, AB11412 = 20, AB11413 = 43, AB11414 = 17, AB11415 = 67, AB11416 = 137, AB11630 = 295, AB11631 = 297, AB11632 = 326, AB11633 = 318, AB11634 = 285, AB11635 = 241, AB11636 = 293, AB11637 = 282, AB11638 = 279, AB11639 = 278, AB11640 = 269, AB11641 = 241, AB12023 = 260, AB12024 = 234, AB12025 = 333, AB12026 = 268, AB12027 = 185, AB12028 = 164, AB12029 = 335, AB12030 = 306, AB12031 = 315, AB12032 = 150, AB12033 = 50, AB12034 = 55, AB12035 = 9, AB12890 = 299, AB12891 = 292, AB12892 = 320, AB12893 = 311, AB12894 = 338, AB12895 = 336, AB13561 = 246, AB13714 = 323, AB13715 = 315, AB13716 = 288, AB13717 = 280, AB13718 = 327, AB13719 = 323, AB13720 = 260, AB13721 = 256, AB13722 = 257, AB13723 = 307, AB14027 = 322, AB14028 = 289, AB14029 = 345, AB14030 = 325, AB14031 = 333, AB14032 = 335, AB14033 = 302, AB14034 = 328, AB14035 = 342, AB14036 = 329, AB14037 = 338, AB14038 = 334, AB14039 = 300, AB14040 = 287, AB14041 = 204, AB14042 = 231, AB14043 = 177, AB14044 = 203. c, QC for 10x multiome mRNA showing number of reads, genes per plate and percentage mitochondria. N = 3,861 cells pooled from 5 mice (3 males, 2 females) exposed to hypoxia 0.1% CO, 4 hours. Each cells has a mean read of 14,537. d, QC for mouse renal 10x multiome ATAC data showing number of reads and genes per plate. N = 3,306 cells pooled from 5 mice (3 males, 2 females) exposed to hypoxia 0.1% CO, 4 hours, in one experiment. Per cell we obtained 10,666 ATAC median high-quality fragments. e, QC for human renal RNA data derived from 10x scRNAseq showing number of reads, genes, and percentage mitochondria per cell, per patient. Box plots presents the third quartile (top of the box), median (centre lines) and first quartile (bottom of the box) of measurements. The whiskers represent 1.5 times the interquartile range from the top or bottom of the box. N = 3 independent experiments, each experiment using kidney from different individual n = 3 biologically independent samples: human sample 1 = 8185 cells, human sample 2 = 3707 cells, human sample 3 = 6358 cells.
Extended Data Fig. 2 Identification and validation of Norn cell markers.
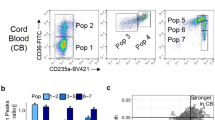
a, Label transfer from Park et al.43 to MARS-seq meta-cell clusters. Each coloured circle denotes a meta-cell. Colour scale denotes z-score of each gene signature across all meta-cells. Collecting duct i., collecting duct intercalated; collecting duct t., collecting duct transient; collecting duct p., collecting duct principal; Distal conv. Tub., distal convoluted tubule. b, Bubble plot depicting enrichment of selected marker genes used for cell labels. Coloured circles denote expression intensity (log2 fold change over the mean). Size of circle represents the percentage of cells expressing the gene within cell type. c, Plot depicts quantification of double positive cells in mRNA FISH imaging of hypoxic (0.1%, 4 hours) kidney sections. N = 1 biological specimen (female), n = 3 independent experiments. The mean is shown. Quantification of cells are shown on top of the plot. Y-axis represents the percentage of Epo expressing cells that are double positive for tested Norn cell marker expression (x-axis). X-axis is showing Norn cell markers tested. d, Schematic of experimental procedure and representative gating strategy using CD73 to enrich for Norn cells in wildtype C57BL/6J mice. Linage depletion using CD31 (endothelial), CD326 (epithelial), and CD45 (immune) markers, and enrichment of CD73 positive cells.
Extended Data Fig. 3 Extended ATAC analysis.
a, UMAP showing cell type clusters based on 10x multiome RNA using the Seurat tool71. Cell label transfer based on supplementary table 1. b, UMAP showing cell type clusters based on 10x multiome ATAC using Seurat tool. c, Heatmap representing sub clustering of selected ATAC-seq regions, and their associated genes.
Extended Data Fig. 4 Transcription-factor motif analysis.
Transcription-factor motif analysis showing enrichment of motifs in accessible regions n = 5,844 peaks Norn specific ATAC-enhancer peaks vs n = 360380 total background sequences. P-value is calculated using cumulative binomial distributions.
Extended Data Fig. 5 SCENIC analysis of Norn cells identifies Tcf21 regulome.
a, Heatmap representing TF regulome enrichment from MARS-seq dataset derived from normoxic kidneys. b, Heatmap representing TF regulome enrichment in our MARS-seq dataset derived from hypoxic kidneys. c, Tcf21 regulome: List of downstream targets of Tcf21.
Extended Data Fig. 6 Identification of Norn cells across development and species.
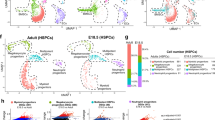
a, Prediction of identified renal cell types, including Norn cells, in adult mouse kidney MARS-seq data (as shown in Fig. 4a). Heatmap showing sum log2 of normalised UMIs. b–g Heatmaps representing predicted cell types, with red box highlighting Norn-like cells: (b) Norn enriched MARS-seq renal dataset (7.4% Norn cells), (c) C57BL/6J adult mice renal scRNA-seq44 (0.1% Norn cells), (d) E18.5 foetal kidney scRNA-seq data45 (7.6% Norn cells, n = 3 individuals), (e) E18.5 Foxd1 enriched foetal kidney scRNA seq data46 (19.9% Norn cells), (f) adult human kidney scRNA seq data (3% Norn cells, n = 12 individuals)42, and (g) foetal week 18 human kidney scRNA seq data (18.8% Norn cells, n = 1 individual)47.
Extended Data Fig. 7 Conservation of Norn cell markers across development and species.
a–e: Scatter plot depicting global log2 size-normalised gene expression in Norn cells versus all other cell types. Green circles represent genes shown in ‘selected’ bar plot. Bar plots showing log2 fold change of Norn cells versus all other cell types in selected genes, receptors, and TFs. a, C57BL/6J adult mice renal scRNA-seq44. b, E18.5 unenriched mouse foetal kidney scRNA-seq data45. c, E18.5 Foxd1 enriched foetal kidney scRNA seq data46. d, Adult human unenriched kidney scRNA seq data42. e, Foetal week 18 human unenriched kidney scRNA seq data (18.8% Norn cells)47. f, Plot showing quantification of double positive cells in mRNA FISH imaging from hypoxic human kidney. N = 3 independent experiments from n = 1 biological specimen (female). The mean is shown. Y-axis is showing percentage of EPO expressing cells that are double positive for tested Norn cell marker expression (x-axis). X-axis is presenting Norn cell markers tested.
Supplementary information
Supplementary Table
1. Gene markers for renal cell types. 2. Top 50 genes for renal cell types. 3. Differential expression between Norn cells versus stromal cells. 4. Differential expression in Norn cells in normoxia versus hypoxia. 5. Cell type-specific ATAC peaks. 6. qPCR analysis. 7. RNA in situ hybridization quantification.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kragesteen, B.K., Giladi, A., David, E. et al. The transcriptional and regulatory identity of erythropoietin producing cells. Nat Med 29, 1191–1200 (2023). https://doi.org/10.1038/s41591-023-02314-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-023-02314-7
This article is cited by
-
Erythrokinetic mechanism(s) causing the “late anemia” of hemolytic disease of the fetus and newborn
Journal of Perinatology (2024)
-
Mechanisms controlling cellular and systemic iron homeostasis
Nature Reviews Molecular Cell Biology (2024)
-
Unmasking the elusive erythropoietin-producing ‘Norn’ cell
Nature Medicine (2023)
-
Identification of erythropoietin-producing cells
Nature Reviews Nephrology (2023)