Abstract
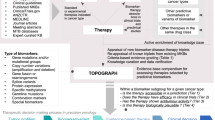
With the increasing use of genomic profiling for diagnosis and therapy guidance in many tumor types, precision oncology is rapidly reshaping cancer care. However, the current trajectory of drug development in oncology results in a paradox: if patients cannot access advanced diagnostics, we may be developing drugs that will reach few patients. In this Perspective, we outline the major challenges to the implementation of precision oncology and discuss critical steps toward resolving these, including facilitation of equal access to genomics tests, ensuring that clinical studies provide robust evidence for new drugs and technologies, enabling physicians to interpret genomics data, and empowering patients toward shared decision-making. A multi-stakeholder approach to evidence generation, value assessment, and healthcare delivery is necessary to translate advances in precision oncology into benefits for patients with cancer globally.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Yates, L. R. et al. The European Society for Medical Oncology (ESMO) Precision Medicine Glossary. Ann. Oncol. 29, 30–35 (2018).
Malone, E. R., Oliva, M., Sabatini, P. J. B., Stockley, T. L. & Siu, L. L. Molecular profiling for precision cancer therapies. Genome Med 12, 8 (2020).
Adashek, J. J., Subbiah, V. & Kurzrock, R. From tissue-agnostic to n-of-one therapies: (r)evolution of the precision paradigm. Trends Cancer 7, 15–28 (2021).
Brown, N. A. & Elenitoba-Johnson, K. S. Enabling precision oncology through precision diagnostics. Annu Rev. Pathol. 15, 97–121 (2020).
Frampton, G. M. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat. Biotechnol. 31, 1023–1031 (2013).
Gondos, A. et al. Genomic testing among patients (pts) with newly diagnosed advanced non-small cell lung cancer (aNSCLC) in the United States: a contemporary clinical practice patterns study. J. Clin. Oncol. 38, Abstract 9592 (2020).
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Non-Small Cell Lung Cancer. Version 5.2021 (2021).
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Prostate Cancer. Version 2.2021 (2021).
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Colon Cancer. Version 2.2021 (2021).
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Ovarian Cancer. Version 1.2021 (2021).
National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Breast Cancer. Version 4.2021 (2021).
Mosele, F. et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Ann. Oncol. 31, 1491–1505 (2020).
Singh, A. P. et al. Impact and diagnostic gaps of comprehensive genomic profiling in real-world clinical practice. Cancers 12, 1156 (2020).
Rodes Sanchez, M., Henderson, N. & Steuten, L. Bridging the Gap: Pathways for Regulatory and Health Technology Assessment of Histology Independent Therapies. Report No. 002290 (Office of Health Economics, 2020).
Thunnissen, E. et al. Lung cancer biomarker testing: perspective from Europe. Transl. Lung Cancer Res 9, 887–897 (2020).
Ettinger, D. S. et al. Non-Small Cell Lung Cancer, Version 5.2017, NCCN Clinical Practice Guidelines in Oncology. J. Natl Compr. Canc Netw. 15, 504–535 (2017).
. Gill, J., Fontrier, A.-M., Miracolo, A. & Kanavos, P. Access to Personalised Oncology in Europe (The London School of Economics and Political Science, 2020).
European Commission. Europe’s Beating Cancer Plan. https://ec.europa.eu/info/law/better-regulation/have-your-say/initiatives/12154-Europe%E2%80%99s-Beating-Cancer-Plan_en (2021).
National Institute for Health (NIH). All of Us Research Program. https://allofus.nih.gov/ (2018).
National Cancer Institute. Cancer Moonshot. https://www.cancer.gov/research/key-initiatives/moonshot-cancer-initiative (2016).
Thavaneswaran, S. et al. Cancer Molecular Screening and Therapeutics (MoST): a framework for multiple, parallel signal-seeking studies of targeted therapies for rare and neglected cancers. Med J. Aust. 209, 354–355 (2018).
Ebi, H. & Bando, H. Precision oncology and the universal health coverage system in Japan. JCO Precis Oncol., Po. 19, 00291 (2019).
Korea University. KU-MAGIC. https://kumagic.korea.edu/kumagic1/index.do (2015).
Faulkner, E. et al. Being precise about precision medicine: what should value frameworks incorporate to address precision medicine? A report of the personalized precision medicine special interest group. Value Health 23, 529–539 (2020).
Hoxhaj, I. et al. A systematic review of the value assessment frameworks used within health technology assessment of omics technologies and their actual adoption from hta agencies. Int J. Environ. Res Public Health 17, 8001 (2020).
Pennell, N. A. et al. Economic impact of next-generation sequencing versus single-gene testing to detect genomic alterations in metastatic non–small-cell lung cancer using a decision analytic model. JCO Precis Oncol. 3, 1–9 (2019).
Chawla, A. et al. Estimated cost of anticancer therapy directed by comprehensive genomic profiling in a single-center study. JCO Precis Oncol. 18, 00074 (2018).
Presley, C. J. et al. Association of broad-based genomic sequencing with survival among patients with advanced non–small cell lung cancer in the community oncology setting. J. Am. Med. Assoc. 320, 469–477 (2018).
Steuten, L., Goulart, B., Meropol, N. J., Pritchard, D. & Ramsey, S. D. Cost effectiveness of multigene panel sequencing for patients with advanced non–small-cell lung cancer. JCO Clin. Cancer Inf. 3, 1–10 (2019).
Weymann, D., Pataky, R. & Regier, D. A. Economic evaluations of next-generation precision oncology: a critical review. JCO Precis. Oncol. 2, 1–23 (2018).
Tan, O., Shrestha, R., Cunich, M. & Schofield, D. J. Application of next-generation sequencing to improve cancer management: a review of the clinical effectiveness and cost-effectiveness. Clin. Genet 93, 533–544 (2018).
McCafferty, J. et al. A systematic analysis of off-label drug use in real-world data (RWD) across more than 145,000 cancer patients. J. Clin. Oncol. 37, Abstract e18031 (2019).
Weda, M. et al. Study on Off-label Use of Medicinal Products in the European Union (European Commission, 2017).
Verbaanderd, C., Rooman, I., Meheus, L. & Huys, I. On-label or off-label? Overcoming regulatory and financial barriers to bring repurposed medicines to cancer patients. Front Pharm. 10, 1664 (2020).
Janiaud, P., Serghiou, S. & Ioannidis, J. P. A. New clinical trial designs in the era of precision medicine: an overview of definitions, strengths, weaknesses, and current use in oncology. Cancer Treat. Rev. 73, 20–30 (2019).
van der Velden, D. L. et al. The Drug Rediscovery Protocol facilitates the expanded use of existing anticancer drugs. Nature 574, 127–131 (2019).
Sicklick, J. K. et al. Molecular profiling of cancer patients enables personalized combination therapy: the I-PREDICT study. Nat. Med. 25, 744–750 (2019).
Cave, A., Kurz, X. & Arlett, P. Real-world data for regulatory decision making: challenges and possible solutions for Europe. Clin. Pharmacol. Ther. 106, 36–39 (2019).
Goossens-Laan, C. A., Kil, P. J., Bosch, J. L. & De Vries, J. Patient-reported outcomes for patients undergoing radical cystectomy: a prospective case–control study. Support Care Cancer 22, 189–200 (2014).
Booth, C. M., Karim, S. & Mackillop, W. J. Real-world data: towards achieving the achievable in cancer care. Nat. Rev. Clin. Oncol. 16, 312–325 (2019).
Siu, L. L. et al. Facilitating a culture of responsible and effective sharing of cancer genome data. Nat. Med. 22, 464–471 (2016).
Deverka, P. A., Douglas, M. P. & Phillips, K. A. Use of real-world evidence in us payer coverage decision-making for next-generation sequencing-based tests: challenges, opportunities, and potential solutions. Value Health 23, 540–550 (2020).
Feinberg, B. A. et al. Use of real-world evidence to support FDA approval of oncology drugs. Value Health 23, 1358–1365 (2020).
Oehrlein, E. M. et al. Peer-reviewed journal editors’ views on real-world evidence. Int. J. Technol. Assess. Health Care 34, 111–119 (2018).
Fatumo, S. et al. A roadmap to increase diversity in genomic studies. Nat. Med. 28, 243–250 (2022).
Nagahashi, M. et al. Formalin-fixed paraffin-embedded sample conditions for deep next generation sequencing. J. Surg. Res. 220, 125–132 (2017).
van de Haar, J. et al. Limited evolution of the actionable metastatic cancer genome under therapeutic pressure. Nat. Med. 27, 1553–1563 (2021).
Rieke, D. T. et al. Comparison of treatment recommendations by molecular tumor boards worldwide. JCO Precis Oncol. 2, 1–14 (2018).
Kato, S. et al. Real-world data from a molecular tumor board demonstrates improved outcomes with a precision n-of-one strategy. Nat. Commun. 11, 4965 (2020).
van de Haar, J., Hoes, L. & Voest, E. Advancing molecular tumour boards: highly needed to maximise the impact of precision medicine. ESMO Open 4, e000516(2019).
van der Velden, D. L. et al. Molecular tumor boards: current practice and future needs. Ann. Oncol. 28, 3070–3075 (2017).
Moore, D. A. et al. Prospective analysis of 895 patients on a UK Genomics Review Board. ESMO Open 4, e000469 (2019).
Priestley, P. et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 575, 210–216 (2019).
Food and Drug Administration. List of cleared or approved companion diagnostic devices (in vitro and imaging tools). https://www.fda.gov/medical-devices/in-vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-in-vitro-and-imaging-tools (2021).
Brannon, A. R. et al. Enhanced specificity of clinical high-sensitivity tumor mutation profiling in cell-free DNA via paired normal sequencing using MSK-ACCESS. Nat. Commun. 12, 3770 (2021).
Leighl, N. B. et al. Clinical utility of comprehensive cell-free DNA analysis to identify genomic biomarkers in patients with newly diagnosed metastatic non–small cell lung cancer Clin. Cancer Res. 25, 4691–4700 (2019).
Woodhouse, R. et al. Clinical and analytical validation of FoundationOne Liquid CDx, a novel 324-Gene cfDNA-based comprehensive genomic profiling assay for cancers of solid tumor origin. PLoS ONE 15, e0237802 (2020).
de Moor, J. S., Gray, S. W., Mitchell, S. A., Klabunde, C. N. & Freedman, A. N. Oncologist confidence in genomic testing and implications for using multimarker tumor panel tests in practice. JCO Precis Oncol. 19, 00338 (2020).
Horgan, D. et al. Bringing greater accuracy to europe’s healthcare systems: the unexploited potential of biomarker testing in oncology. Biomed. Hub. 5, 182–223 (2020).
Dittrich, C. et al. ESMO / ASCO Recommendations for a Global Curriculum in Medical Oncology Edition 2016. ESMO Open 1, e000097 (2016).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis Oncol. 2017, PO.17.00011 (2017).
Tamborero, D. et al. Support systems to guide clinical decision-making in precision oncology: The Cancer Core Europe Molecular Tumor Board Portal. Nat. Med. 26, 992–994 (2020).
Mateo, J. et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO Scale for Clinical Actionability of molecular Targets (ESCAT). Ann. Oncol. 29, 1895–1902 (2018).
Sukhai, M. A. et al. A classification system for clinical relevance of somatic variants identified in molecular profiling of cancer. Genet. Med. 18, 128–136 (2016).
Condorelli, R. et al. Genomic alterations in breast cancer: Level of evidence for actionability according to ESMO Scale for Clinical Actionability of molecular Targets (ESCAT). Ann. Oncol. 30, 365–373 (2019).
Moscow, J. A., Fojo, T. & Schilsky, R. L. The evidence framework for precision cancer medicine. Nat. Rev. Clin. Oncol. 15, 183–192 (2018).
Luchini, C., Lawlor, R. T., Milella, M. & Scarpa, A. Molecular tumor boards in clinical practice. Trends Cancer 6, 738–744 (2020).
Sharma, V. et al. Eye-tracking study to enhance usability of molecular diagnostics reports in cancer precision medicine. JCO Precis. Oncol. 2, 1–11 (2018).
Westphalen, B. C. et al. Conceptual framework for precision cancer medicine in Germany: consensus statement of the Deutsche Krebshilfe working group ‘Molecular Diagnostics and Therapy’. Eur. J. Cancer 135, 1–7 (2020).
Pishvaian, M. J. et al. A virtual molecular tumor board to improve efficiency and scalability of delivering precision oncology to physicians and their patients. JAMIA Open 2, 505–515 (2019).
Johnston, J. J. et al. Secondary variants in individuals undergoing exome sequencing: screening of 572 individuals identifies high-penetrance mutations in cancer-susceptibility genes. Am. J. Hum. Genet 91, 97–108 (2012).
Kamps, R. et al. Next-generation sequencing in oncology: genetic diagnosis, risk prediction and cancer classification. Int. J. Mol. Sci. 18, 308 (2017).
Acknowledgements
Funding support and role of the funder/sponsor: F. Hoffmann-La Roche funded third-party writing assistance for the initial draft of this manuscript, furnished by S. Salem at Health Interactions. The sponsor was not involved in discussions relating to content. All subsequent versions were written, reviewed, and submitted solely by the authors.
Author information
Authors and Affiliations
Contributions
J.M., L.S., and E.V. contributed to drafting of the manuscript. All authors contributed to critical revision of the manuscript for important intellectual content and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
J.M. has received grants to his institution (as principal investigator) from AstraZeneca and Pfizer Oncology; consulting fees from Monterosa, consulting fees for an advisory board from AstraZeneca, MSD, Clovis Oncology, F. Hoffmann-La Roche, Pfizer, and Janssen; payment or honoraria for lectures, presentations, speakers bureaus, manuscript writing, or educational events from AstraZeneca, Pfizer Oncology, F. Hoffmann-La Roche, Guardant Health, Astellas, and Janssen; support for attending meetings and/or travel from AstraZeneca and IPSEN; and drugs for preclinical testing in research from AstraZeneca. L.S. has received support for the present manuscript (for example, funding, provision of study materials, medical writing, article processing charges) to her institution from F. Hoffmann-La Roche, grants or contracts to her institution from the Personalized Medicine Coalition, and support for attending meetings and/or travel from the Personalized Medicine Coalition. P.A. has received personal fees from Boehringer Ingelheim, Marcogenics, Amcure, Synthon, Servier, G1 Therapeutics, F. Hoffmann-La Roche, Novartis, Deloitte, Radius, Menarini, Gilead, and Amgen; and travel grants from MSD, F. Hoffmann-La Roche, Pfizer, and Amgen. F.A. has received institutional research funding from F. Hoffmann-La Roche, Pfizer, Eli Lilly, Novartis, AstraZeneca, and Daiichi Sankyo. M.D. has received personal-speaker fees from F. Hoffmann-La Roche, Pfizer, and Eli Lilly. E.G. has received grants or contracts from Novartis, F. Hoffmann-La Roche, ThermoFisher, AstraZeneca, Taiho, and BeiGene; payment or honoraria for lectures, presentations, speakers’ bureaus, manuscript writing, or educational events from F. Hoffmann-La Roche/Genentech, Ellipses Pharma, Neomed Therapeutics 1, Boehringer Ingelheim, Janssen Global Services, Seagen, TFS, Alkermes, ThermoFisher, and BMS; and has participated on a data-safety monitory or advisory board for F. Hoffmann-La Roche/Genentech, Boehringer Ingelheim, Janssen Global Services, MabDiscovery, Anaveon, and ThermoFisher. J.G. has served a consulting or advisory role for Amgen, Alnylam, BMS, Bayer, BioMarin, Janssen, Novartis, Pfizer, F. Hoffmann-La Roche, Servier, Takeda, and UCB; and has received institutional research funding from EFPIA companies to IMI projects, and travel/accommodation expenses from Amgen, Alnylam, BMS, Bayer, BioMarin, Janssen, Novartis, Pfizer, F. Hoffmann-La Roche, Servier, Takeda, and UCB. D.H. has received honoraria and consulting fees from F. Hoffmann-La Roche, Thermofisher Scientific, Amgen, AstraZeneca, BMS, Boeringher Ingelheim, Eli Llilly, Janssen, and Pfizer, as well as from contract research organizations and not-for-profit entities, including the Canadian Agency for Drugs and Technologies in Health, the Institute of Health Economics, and The Ministry of Health of Ontario; and has received consultation fees from life-sciences companies with an interest in the adoption of advanced testing. I.M.-L. has received support for the present manuscript (for example, funding, provision of study materials, medical writing, article processing charges) from F. Hoffmann-La Roche; consulting fees from Novartis; and payment or honoraria for lectures, presentations, speakers’ bureaus, manuscript writing, or educational events from GSK, AstraZeneca, and BH. N.N. has received personal fees from MSD, Qiagen, Biocartis, Incyte, F. Hoffmann-La Roche, BMS, Merck, ThermoFisher, Boehringer Ingelheim, AstraZeneca, Sanofi, Eli Lilly, Bayer, ArcherDx, Illumina, and Amgen; and grants from Qiagen, Biocartis, Incyte, F. Hoffmann-La Roche, BMS, Merck, ThermoFisher, Boehringer Ingelheim, AstraZeneca, and Illumina. J.S.R.-F. has received consulting fees from Paige.AI, Repare Therapeutics, Goldman Sachs, and Eli Lilly; has participated on a data-safety monitoring or an advisory board for F. Hoffmann-La Roche, Roche Tissue Diagnostics, Genentech, Novartis, and InVicro; has a leadership or fiduciary role in Grupo Oncoclinicas (as a member of the board of directors); and has stock or stock options in Paige.AI, Repare Therapeutics, and Grupo Oncoclinicas. S.S. has received consulting fees (all were institutional contracts) from F. Hoffmann-La Roche, Novartis, AstraZeneca, Pfizer, Libbs, Merck, MSD, Eli Lilly, BMS, and Sanofi Aventis. D.M.T. reports grants, personal fees, and non-financial support from F. Hoffmann-La Roche, Pfizer, and Bayer; grants and non-financial support from AstraZeneca, Seattle Genetics, Amgen, Eli Lilly, and Eisai; personal fees from Omico; and non-financial support from Elevation Oncology, outside the submitted work. C.B.W. has received honoraria from Amgen, Bayer, Chugai, Celgene, Falk, GSK, MSD, Merck, Janssen, Ipsen, Roche, Servier, SIRTeX, Taiho; has served on advisory boards for Bayer, BMS, Celgene, Servier, Shire/Baxalta, Rafael Pharmaceuticals, RedHill, and Roche; and has received travel support from Bayer, Celgene, RedHill, Roche, Servier, and Taiho and research grants (institutional) from Roche outside of the submitted work. E.V. has received clinical study grants to his institution from Amgen, AstraZeneca, Pfizer, F. Hoffmann-La Roche, Clovis, GSK, Novartis, Bayer, Sanofi, BMS, MSD, and BI. The Netherlands Cancer Institute (NKI) has received a speaker’s fee from F. Hoffmann-La Roche. All authors received support in the form of third-party medical writing assistance for this manuscript, furnished by S. Salem of Health Interactions, from F. Hoffmann-La Roche, Basel, Switzerland.
Peer review
Peer review information
Nature Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. Karen O’Leary was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mateo, J., Steuten, L., Aftimos, P. et al. Delivering precision oncology to patients with cancer. Nat Med 28, 658–665 (2022). https://doi.org/10.1038/s41591-022-01717-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01717-2
This article is cited by
-
A multimodal graph neural network framework for cancer molecular subtype classification
BMC Bioinformatics (2024)
-
Advancing precision rheumatology through tissue and blood profiling
Nature Reviews Rheumatology (2024)
-
Machine learning-based radiomics analysis in predicting RAS mutational status using magnetic resonance imaging
La radiologia medica (2024)
-
Supporting the decision to perform molecular profiling for cancer patients based on routinely collected data through the use of machine learning
Clinical and Experimental Medicine (2024)
-
Molecular diagnostics tailoring personalized cancer therapy—an oncologist’s view
Virchows Archiv (2024)